Abstract
Aside from PD-L1 expression, biomarkers of response to immune checkpoint inhibitors (ICIs) in non-small-cell lung cancer (NSCLC) are needed. In a previous retrospective analysis, we documented that fecal Akkermansia muciniphila (Akk) was associated with clinical benefit of ICI in patients with NSCLC or kidney cancer. In the current study, we performed shotgun-metagenomics-based microbiome profiling in a large cohort of patients with advanced NSCLC (n = 338) treated with first- or second-line ICIs to prospectively validate the predictive value of fecal Akk. Baseline stool Akk was associated with increased objective response rates and overall survival in multivariate analyses, independent of PD-L1 expression, antibiotics, and performance status. Intestinal Akk was accompanied by a richer commensalism, including Eubacterium hallii and Bifidobacterium adolescentis, and a more inflamed tumor microenvironment in a subset of patients. However, antibiotic use (20% of cases) coincided with a relative dominance of Akk above 4.8% accompanied with the genus Clostridium, both associated with resistance to ICI. Our study shows significant differences in relative abundance of Akk that may represent potential biomarkers to refine patient stratification in future studies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The data generated or analyzed during this study are included within the paper, its Supplementary Information files and public repositories. Detailed information on the cohort is available in Supplementary Table 8 raw metagenomic sequences are available in the SRA under the Bioproject accession PRJNA751792 (https://www.ncbi.nlm.nih.gov/bioproject/PRJNA751792) and PRJNA782662 (https://www.ncbi.nlm.nih.gov/bioproject/PRJNA782662), and raw RNA sequencing are available in the SRA under accession the NCBI accession GSE182328 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE182328). All BioSample (PATIENTS_Metadata.csv) are also provided as supplementary information. Source data are provided with this paper.
Code availability
No unique software or computational code was created for this study. Code detailing implementation of established tools/pipelines are described in details in the Method section and available upon request to the corresponding author.
References
Herbst, R. S. et al. Pembrolizumab versus docetaxel for previously treated, PD-L1-positive, advanced non-small-cell lung cancer (KEYNOTE-010): a randomised controlled trial. Lancet 387, 1540–1550 (2016).
Brahmer, J. et al. Nivolumab versus docetaxel in advanced squamous-cell non–small-cell lung cancer. N. Engl. J. Med. 373, 123–135 (2015).
Borghaei, H. et al. Nivolumab versus docetaxel in advanced nonsquamous non–small-cell lung cancer. N. Engl. J. Med. 373, 1627–1639 (2015).
Gandhi, L. et al. Pembrolizumab plus chemotherapy in metastatic non-small-cell lung cancer. N. Engl. J. Med. 378, 2078–2092 (2018).
Paz-Ares, L. et al. Pembrolizumab plus chemotherapy for squamous non–small-cell lung cancer. N. Engl. J. Med. 379, 2040–2051 (2018).
Reck, M. et al. Pembrolizumab versus chemotherapy for pd-l1–positive non–small-cell lung cancer. N. Engl. J. Med. 375, 1823–1833 (2016).
Gadgeel, S. et al. Updated analysis from KEYNOTE-189: pembrolizumab or placebo plus pemetrexed and platinum for previously untreated metastatic nonsquamous non-small-cell lung cancer. J. Clin. Oncol. 38, 1505–1517 (2020).
Ferrara, R. et al. Hyperprogressive disease in patients with advanced non-small cell lung cancer treated with PD-1/PD-L1 inhibitors or with single-agent chemotherapy. JAMA Oncol. 4, 1543–1552 (2018).
Riaz, N. et al. Tumor and microenvironment evolution during immunotherapy with nivolumab. Cell 171, 934–949.e16 (2017).
Rizvi, N. A. et al. Cancer immunology. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 348, 124–128 (2015).
Spranger, S., Bao, R. & Gajewski, T. F. Melanoma-intrinsic β-catenin signalling prevents anti-tumour immunity. Nature 523, 231–235 (2015).
Smyth, M. J., Ngiow, S. F., Ribas, A. & Teng, M. W. L. Combination cancer immunotherapies tailored to the tumour microenvironment. Nat. Rev. Clin. Oncol. 13, 143–158 (2016).
Koyama, S. et al. Adaptive resistance to therapeutic PD-1 blockade is associated with upregulation of alternative immune checkpoints. Nat. Commun. 7, 10501 (2016).
Young, A. et al. Co-inhibition of CD73 and A2AR adenosine signaling improves anti-tumor immune responses. Cancer Cell 30, 391–403 (2016).
Chen, D. S. & Mellman, I. Elements of cancer immunity and the cancer–immune set point. Nature 541, 321–330 (2017).
Derosa, L. et al. Microbiota-centered interventions: the next breakthrough in immuno-oncology? Cancer Discov. 11, 2396–2412 (2021).
Routy, B. et al. Gut microbiome influences efficacy of PD-1-based immunotherapy against epithelial tumors. Science 359, 91–97 (2018).
Vétizou, M. et al. Anticancer immunotherapy by CTLA-4 blockade relies on the gut microbiota. Science 350, 1079–1084 (2015).
Gopalakrishnan, V. et al. Gut microbiome modulates response to anti-PD-1 immunotherapy in melanoma patients. Science 359, 97–103 (2018).
Derosa, L. et al. Negative association of antibiotics on clinical activity of immune checkpoint inhibitors in patients with advanced renal cell and non-small-cell lung cancer. Ann. Oncol. 29, 1437–1444 (2018).
Derosa, L. & Zitvogel, L. Antibiotics impair immunotherapy for urothelial cancer. Nat. Rev. Urol. 17, 605–606 (2020).
Matson, V. et al. The commensal microbiome is associated with anti-PD-1 efficacy in metastatic melanoma patients. Science 359, 104–108 (2018).
Jin, Y. et al. The diversity of gut microbiome is associated with favorable responses to anti-programmed death 1 immunotherapy in Chinese patients with NSCLC. J. Thorac. Oncol. 14, 1378–1389 (2019).
Daisley, B. A. et al. Abiraterone acetate preferentially enriches for the gut commensal Akkermansia muciniphila in castrate-resistant prostate cancer patients. Nat. Commun. 11, 4822 (2020).
Santoro, A. et al. Gut microbiota changes in the extreme decades of human life: a focus on centenarians. Cell. Mol. Life Sci. 75, 129–148 (2018).
Zhou, Q. et al. Gut bacteria Akkermansia is associated with reduced risk of obesity: evidence from the American Gut Project. Nutr. Metab. 17, 90 (2020).
Blacher, E. et al. Potential roles of gut microbiome and metabolites in modulating ALS in mice. Nature 572, 474–480 (2019).
Bárcena, C. et al. Healthspan and lifespan extension by fecal microbiota transplantation into progeroid mice. Nat. Med. 25, 1234–1242 (2019).
Romano, S. et al. Meta-analysis of the Parkinson’s disease gut microbiome suggests alterations linked to intestinal inflammation. NPJ Parkinsons Dis. 7, 1–13 (2021).
Shono, Y. et al. Increased GVHD-related mortality with broad-spectrum antibiotic use after allogeneic hematopoietic stem cell transplantation in human patients and mice. Sci. Transl. Med. 8, 339ra71 (2016).
Desai, M. S. et al. A dietary fiber-deprived gut microbiota degrades the colonic mucus barrier and enhances pathogen susceptibility. Cell 167, 1339–1353.e21 (2016).
Karcher, N. et al. Genomic diversity and ecology of human-associated Akkermansia species in the gut microbiome revealed by extensive metagenomic assembly. Genome Biol. 22, 209 (2021).
Le Chatelier, E. et al. Richness of human gut microbiome correlates with metabolic markers. Nature 500, 541–546 (2013).
Routy, B. et al. The gut microbiota influences anticancer immunosurveillance and general health. Nat. Rev. Clin. Oncol. 15, 382–396 (2018).
Derosa, L. et al. Gut bacteria composition drives primary resistance to cancer immunotherapy in renal cell carcinoma patients. Eur. Urol. 78, 195–206 (2020).
Meng, X. et al. Immune microenvironment differences between squamous and non-squamous non-small-cell lung cancer and their influence on the prognosis. Clin. Lung Cancer 20, 48–58 (2019).
Hwang, S. et al. Immune gene signatures for predicting durable clinical benefit of anti-PD-1 immunotherapy in patients with non-small cell lung cancer. Sci. Rep. 10, 643 (2020).
Nakajima, K. et al. IAP inhibitor, Embelin increases VCAM-1 levels on the endothelium, producing lymphocytic infiltration and antitumor immunity. Oncoimmunology 9, 1838812 (2020).
Hakozaki, T. et al. The gut microbiome associates with immune checkpoint inhibition outcomes in patients with advanced non-small cell lung cancer. Cancer Immunol. Res. 8, 1243–1250 (2020).
Tsay, J.-C. J. et al. Lower airway dysbiosis affects lung cancer progression. Cancer Discov. 11, 293–307 (2020). https://doi.org/10.1158/2159-8290.CD-20-0263
Benevides, L. et al. New insights into the diversity of the genus Faecalibacterium. Front. Microbiol. 8, 1790 (2017).
Lee, S.-H. et al. Bifidobacterium bifidum strains synergize with immune checkpoint inhibitors to reduce tumour burden in mice. Nat. Microbiol. 6, 277–288 (2021).
Elkrief, A., Derosa, L., Kroemer, G., Zitvogel, L. & Routy, B. The negative impact of antibiotics on outcomes in cancer patients treated with immunotherapy: a new independent prognostic factor? Ann. Oncol. 30, 1572–1579 (2019).
Seo, S. W., Kim, D., Szubin, R. & Palsson, B. O. Genome-wide reconstruction of OxyR and SoxRS Transcriptional regulatory networks under oxidative stress in Escherichia coli K-12 MG1655. Cell Rep. 12, 1289–1299 (2015).
Atarashi, K. et al. Induction of colonic regulatory T cells by indigenous Clostridium species. Science 331, 337–341 (2011).
Ruusunen, A., Rocks, T., Jacka, F. & Loughman, A. The gut microbiome in anorexia nervosa: relevance for nutritional rehabilitation. Psychopharmacol. 236, 1545–1558 (2019).
van der Lugt, B. et al. Akkermansia muciniphila ameliorates the age-related decline in colonic mucus thickness and attenuates immune activation in accelerated aging Ercc1−/Δ7 mice. Immun. Ageing 16, 6 (2019).
Depommier, C. et al. Supplementation with Akkermansia muciniphila in overweight and obese human volunteers: a proof-of-concept exploratory study. Nat. Med. 25, 1096–1103 (2019).
Ouyang, J. et al. The bacterium Akkermansia muciniphila: a sentinel for gut permeability and its relevance to HIV-related inflammation. Front. Immunol. 11, 645 (2020).
Huck, O. et al. Akkermansia muciniphila reduces Porphyromonas gingivalis-induced inflammation and periodontal bone destruction. J. Clin. Periodontol. 47, 202–212 (2020).
Wu, W. et al. Protective effect of Akkermansia muciniphila against immune-mediated liver injury in a mouse model. Front. Microbiol. 8, 1804 (2017).
Baruch, E. N. et al. Fecal microbiota transplant promotes response in immunotherapy-refractory melanoma patients. Science 371, 602–609 (2020).
Shi, L. et al. Combining IL-2-based immunotherapy with commensal probiotics produces enhanced antitumor immune response and tumor clearance. J. Immunother. Cancer 8, e000973 (2020).
Cani, P. D. & de Vos, W. M. Next-generation beneficial microbes: the case of Akkermansia muciniphila. Front. Microbiol. 8, 1765 (2017).
Eisenhauer, E. A. et al. New response evaluation criteria in solid tumours: revised RECIST guideline (version 1.1). Eur. J. Cancer 45, 228–247 (2009).
Li, J. et al. An integrated catalog of reference genes in the human gut microbiome. Nat. Biotechnol. 32, 834–841 (2014).
Nielsen, H. B. et al. Identification and assembly of genomes and genetic elements in complex metagenomic samples without using reference genomes. Nat. Biotechnol. 32, 822–828 (2014).
Beghini, F. et al. Integrating taxonomic, functional, and strain-level profiling of diverse microbial communities with bioBakery 3. Elife 10, e65088 (2021).
Fumet, J.-D. et al. Prognostic and predictive role of CD8 and PD-L1 determination in lung tumor tissue of patients under anti-PD-1 therapy. Br. J. Cancer 119, 950–960 (2018).
Bray, N. L., Pimentel, H., Melsted, P. & Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 34, 525–527 (2016).
Segata, N. et al. Metagenomic biomarker discovery and explanation. Genome Biol. 12, R60 (2011).
Lin, H. & Peddada, S. D. Analysis of compositions of microbiomes with bias correction. Nat. Commun. 11, 3514 (2020).
Mallick, R. et al. Subduction initiation and the rise of the Shillong Plateau. Earth Planet. Sci. Lett. 543, 116351 (2020).
Acknowledgements
L. Z. and B. B. were funded by the RHU Torino Lumière (ANR-16-RHUS-0008), while L. Z. was supported by H2020 ONCOBIOME and ANR, French–German Ileobiome, 19-CE15-0029-01. L. Z. and G. K. received a donation from Seerave Foundation. L. Z., L. D., and E. D. were supported by the SIRIC Stratified Oncology Cell DNA Repair and Tumor Immune Elimination (SOCRATE). L. D. and L. Z. were supported by SIGN’IT 2020 ARC foundation. This work was also supported by the Prism project funded by the Agence Nationale de la Recherche under grant number ANR-18-IBHU-0002. L. Z. and G. K. were supported by the Ligue contre le Cancer (équipe labelisée); ANR Projets blancs; ANR under the frame of E-Rare-2, the ERA-Net for Research on Rare Diseases; Association Pour La Recherche Sur Le Cancer (ARC); Bristol-Myers Squibb (BMS) (International Immuno-Oncology Network), Cancéropôle Ile-de-France; Chancellerie des Universités de Paris (Legs Poix), Fondation pour la Recherche Médicale (FRM); a donation by Elior; the European Commission (ArtForce); the European Research Council (ERC); Fondation Carrefour; Institut National du Cancer (INCa); Inserm (HTE); Institut Universitaire de France; LeDucq Foundation; the LabEx Immuno-Oncology; FHU CARE, Dassault, and Badinter Philantropia, and the Paris Alliance of Cancer Research Institutes (PACRI). B. R. is supported by the Canadian Institutes of Health Research (CIHR), Institut du cancer de Montreal and the Prefontaine family. A. C. is supported by the CPRIT Research Training Program (RP170067). N. S. was supported by the European Research Council (ERC-STG project MetaPG-716575); MIUR ‘Futuro in Ricerca’ (grant No. RBFR13EWWI_001); the European H2020 program (ONCOBIOME-825410 project and MASTER-818368 project); the National Cancer Institute of the National Institutes of Health (1U01CA230551); by the Premio Internazionale Lombardia e Ricerca 2019 to G. K. A. M. has been or is currently an investigator in clinical trials sponsored by BMS, MSD, GlaxoSmithKline (GSK)/Tesaro, Janssen, Roche/Genentech, Pfizer, AstraZeneca, Amgen. A. M. has been or is currently a member of Clinical Trial Scientific Steering Committee for AstraZeneca and GSK. A. M. has been or is currently a member of the scientific advisory board of the following companies: Merck Serono, Novartis, BMS, Symphogen, Amgen, Tesaro/GSK, Pfizer, AstraZeneca/Medimmune, Servier, and Sanofi. A. M. has provided scientific and medical consulting to the following companies: Roche, Sanofi. A. M. is a member of the data safety and monitoring board for trial NCT02423863 (TLR3 agonist; Oncovir). A. M. has received research funding and/or drug supply for preclinical and clinical research projects from: BMS, Boehringer Ingelheim, Idera, MSD, Fondation MSD Avenir, and SIRIC (INCa-DGOS-Inserm_12551).
Author information
Authors and Affiliations
Contributions
L. Z., L. D., B. R., and B. B. conceived the study. L. D. and B. R. orchestrated the ONCOBIOTICS study. B. B. coordinated the clinical inclusions. L. D., G. Z., S. F., J. M., C. A.-V., D. M.-S., F. Goldwasser, A. S., H. P., F. Ghiringhelli, F. B., D. P., S. M., N. B., B. B., and J. C. S. enrolled patients and managed patient stool collection. L. D., C. A. C. S., M. B., S. T., and B. R. coordinated the ONCOBIOTICS network for stool traceability and data collection and management. F. B.-D. recovered the Tumor Mutational Burden. F. C., A. B., and N. S. worked on the new markers for Akk in MetaPhlAn 3. R. B. and S. C. worked on RNA-seq. A. M. T., V. I., and N. S. interpreted the MG analyses and performed independent statistical analyses. C. R. O. Absilon planned the data management and kept the electronic files. D. D. double checked the statistical analyses. L. D., A. D., B. R., and M. M. designed the figures. R. D. and L. D. managed mice experiments. E. L. C., L. A., N. P., N. G., H. R., managed the sequencing of stool samples G. K. participated in data interpretation and edited the paper. All authors participated in paper editing.
Corresponding author
Ethics declarations
Competing interests
L. Z. received research contract from Kaleido and Innovate Pharma and Pilege. L. D. has consulted for and had an advisory role for BMS and Sanofi and was supported by Philantropia Fondation Gustave Roussy. G. Z. received a research grant from Fondation Roche, received fees from Roche, MSD, BMS, and Astra-Zeneca, and is consultant for Da Volterra and Inventiva. L.Z and R.D. are the scientific cofounders of Everimmune. E. D. reports grants and personal fees from Roche Genentech, grants from Boehringer, grants from AstraZeneca, grants and personal fees from Merck Serono, grants from BMS, and grants from MSD. P. D. has had consulting and advisory roles for AstraZeneca, BMS, Boehringer Ingelheim, Celgene, Daiichi Sankyo, Eli Lilly, Merck, Novartis, Pfizer, prIME Oncology, Peer CME, Roche, and Samsung, as well as honoraria from AstraZeneca, BMS, Boehringer Ingelheim, Celgene, Eli Lilly, Merck, Novartis, Pfizer, prIME Oncology, Peer CME, Roche, and Samsung. P. D. ran clinical trials as principal or co-investigator for AstraZeneca, BMS, Boehringer Ingelheim, Eli Lilly, Merck, Novartis, Pfizer, Roche, Medimmun, Sanofi-Aventis, Taiho Pharma, Novocure, and Daiichi Sankyo, and has received travel or accommodation expenses from AstraZeneca, Roche, Novartis, prIME Oncology, and Pfizer. F. G. received honoraria from Amgen, Sanofi, Merk Serono, MSD, BMS, and AstraZeneca, had a consultancy or advisory role for Roche and Enterome, and received direct research fundings from Roche, Enterome, AstraZeneca, and Servier and traveling support from Servier, Amgen, and Roche. J. C. S. in the last 2 years has received consultancy fees from Relay Therapeutics and Gritstone and holds shares of Hookipa, Gritstone, AstraZeneca, and Daiichi Sankyo, and was a full time employee for AstraZeneca in 2017–2019. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Medicine thanks Christian Jobin, Anna Heintz-Buschart, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Javier Carmona was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Consort diagram describing the stool collection in the whole NSCLC ONCOBIOTICS cohort.
Consortium diagram for patients enrollment in the ONCOBIOTICS study (n=493) according to stool availability, presence of Akkermansia muciniphila (Akk), and tumor PD-L1 expression levels. V1; baseline fecal sample. V2; before the second injection of the immune checkpoint inhibitor; MGS: Metagenomic Sequencing. Akk+: detection of Akk; Akk-: no detection of Akk by shotgun metagaenomics sequencing (MGS) analysis at diagnosis.
Extended Data Fig. 2 MetaOMineR-based analysis of the association between stool A. muciniphila (Akk) and clinical benefit to ICI in patients.
A-B. Correlations between stool prevalence of Akk (MetaOMineR pipeline) and ORR (A) or OS (B) in 1+2L NSCLC patients (N=338, A-B) based on MGS identification of Akk in the MetaOMineR algorithm (INRAE). Chi-square test (A) and Cox regression analysis for median overall survival (OS) depicted in Kaplan Meier curves according to detectable or undetectable Akk (Akk+ or Akk) analyzed in 2 groups (B, left panel) or segregated in 3 groups (Akk-, Akklow and Akkhigh) (B, right panel). Chi-square test P-values are two-sided, with no adjustments made for multiple comparisons (A). The Akk status was compared using the stratified log-rank test. P-values are one-sided with no adjustment (B). C. Experimental setting of avatar mice. FMT of NSCLC patients (Supplementary Table S3) segregated according to the presence or absence of Akk into MCA-205 tumor bearing C57BL/6 mice. Treatments are indicated by arrows (ATB, FMT, anti-PD-1 (ICI) mAbs, or isotype control mAbs (Iso)). D. MCA-205 tumor growth kinetics in each group of FMT according to the prevalence of Akk. in isotype Ctl versus anti-PD-1 mAbs treated mice. Data are presented as mean values +/-SEM of tumor sizes within 6 animals/group. Concatenation of at least n=8 experiments (using a different stool of NSCLC patient) containing 6 mice/group. Tumor sizes according to FMT Akk- (D, left panel) versus Akk+ (D, right panel) are depicted, each dot representing one mouse. Statistics were mixed-effect modeling with specific software ((https://kroemerlab.shinyapps.io/TumGrowth/) for longitudinal tumor growth analysis. P-value are indicated. E. Percentages of responding mice (tumor reduction of>25 % compared with means of controls in the anti-PD-1 mAbs -treated group) and patients (ORR) in each category of stools used for FMT (derived from patients in Supplementary Table S3). CR; complete response. PR; partial response, SD; stable disease, PD; progressive disease.
Extended Data Fig. 3 Metagenomic species characterizing Akk+ stools and patient survival.
Shannon diversity index representing stool alpha diversity in Akk+ and Akk- groups of fecal specimens (N=338) (A, upper panel). Beta-diversity measured by Bray-Curtis Index represented by Principal Coordinates analysis (PCoA) between Akk+ versus Akk- groups in the whole cohort of 1+2L (A, lower panel). p-values were calculated using PERMANOVA with 999 permutations. The lower and the upper hinges of boxplots corresponds to the 25th and 75th percentiles, respectively. The midline is the median. The upper and lower whiskers extend from the hinges to the largest (or smallest) value no further than ×1.5 interquartile range from the hinge, defined as the distance between the 25th and 75th percentiles. P-values were calculated testing the null hypothesis and using a two-sided test. Exact p-value: 3.84573e-05. B-C. Differential abundance of metagenomic species measured by linear discriminant analysis of effect size (LEfSe) according to the presence of A. muciniphila (Akk) (B) and the OS at 12 months (C) within Akk+ group (C, left panel) and Akk- group (C, right panel). LDA; Linear discriminant analysis. OS: overall survival. P-values were calculated using a two-sided nonparametric factorial Kruskal-Wallis (KW) sum-rank test. # Multivariate analysis (ANCOM-BC/Maaslin2) with a false discovery rate (FDR) adjusted p-value <0.2.
Extended Data Fig. 4 Compositional taxonomic differences in stools of NSCLC patients segregated according to Akk relative abundance.
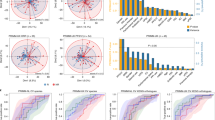
A. Alpha diversity according to Akk relative abundance segregated in 3 groups Akk-: undetectable Akk, Akklow: A. muciniphila relative abundance between 0.035-4.799% (<77th percentile of positive samples), and Akkhigh: 4.799% (> 77th percentile) (N=338). The lower and upper hinges of boxplots correspond to the 25th and 75th percentiles, respectively. The midline is the median. The upper and lower whiskers extend from the hinges to the largest (or smallest) value no further than ×1.5 interquartile range from the hinge, defined as the distance between the 25th and 75th percentiles. P-values were calculated using a two-sided nonparametric Wilcoxon sum-rank test. B-C. Beta-diversity using PCoA between Akk- and Akklow (B) and between Akklow and Akkhigh (C) p-values were calculated using PERMANOVA with 999 permutations. The PERMANOVA test compares groups of objects and tests the null hypothesis that the centroids and dispersion of the groups are equivalent. The P-value is calculated by comparing the actual F test to that gained from (in this case 999) random permutations of the objects between the groups. If p <0.05, the null hypothesis is disregarded and we conclude that the centroids and dispersion between the groups are not equivalent. D-E. Variable importance plot (VIP) discriminant analysis of taxonomic stool composition according to Akk relative abundance, between Akk- versus Akklow (D) and Akklow versus Akkhigh (E). Differences in bacterial prevalence and abundance in fold ratios are indicated in these VIP plots. VIP: Variable importance plot. * p <0.05,** p <0.01, *** p <0.001. P-values were calculated using a two-sided nonparametric Wilcoxon sum-rank test. # Multivariate analysis (ANCOM-BC/Maaslin2) with a false discovery rate (FDR) adjusted p-value <0.2.
Extended Data Fig. 5 Interaction between ATB and A.muciniphila on survival and microbiome composition.
A. Kaplan-Meier curve and Cox regression analysis of overall survival in the n=338 patients according to detectable versus undetectable Akk (Akk+ and Akk-) and ATB use (noATB: no exposure to ATB, ATB: antibiotics exposure within 2 months prior to ICI initiation). The Akk status and ATB use were compared using the stratified log-rank test. P-values are one-sided with no adjustment. B. Shannon diversity index representing stool alpha diversity in Akk+ and Akk- groups of fecal specimen from patients exposed or not to ATB (N=338). The lower and upper hinges of boxplots correspond to the 25th and 75th percentiles, respectively. The midline is the median. The upper and lower whiskers extend from the hinges to the largest (or smallest) value no further than ×1.5 interquartile range from the hinge, defined as the distance between the 25th and 75th percentiles. P-values were calculated using a two-sided nonparametric Wilcoxon sum-rank test. C. Box Plots representing the relative abundance (mean+/-SEM) of Akk according to overall survival at 12 months and exposure or not to ATB in n=338 patients. The lower and upper hinges of boxplots correspond to the 25th and 75th percentiles, respectively. The midline is the median. The upper and lower whiskers extend from the hinges to the largest (or smallest) value no further than ×1.5 interquartile range from the hinge, defined as the distance between the 25th and 75th percentiles. The test used was Kruskal-Wallis, two-sided, 5% level of significance. No adjustments were made for multiple comparisons. D. Heatmap showing differentially abundant species identified in stools with detectable Akk (Akk+) in patients exposed to (D) or not exposed to (E) ATB within 2 months prior to ICI initiation. Species were identified using a non-parametric Kruskall-Wallis test comparing 4 groups made up of 2 variables: Akkermansia muciniphila presence/absence and antibiotic use. The figure shows species’ abundances across samples whose False Discovery Rate (FDR) was <0.2 in the KW test and whose Wilcoxon Rank Sum Test p-value was <0.05 when comparing the highlighted group to the rest.
Extended Data Fig. 6 Akkp2261 modulated the murine microbiome composition, rescuing responsiveness to PD-1 blockade.
A. Experimental setting. After 3 days of ATB, FMT was performed in mice by oral gavage using patient stools classified according to Akk (Akk+ and Akk-). 14 days later, MCA-205 tumors were i.d inoculated, and mice were treated with anti-PD-1 or iso-control mAbs 4 times every 3 days concomitantly with oral supplementation of Akkp2261 four times every 3 days. B-D. Mean MCA-205 tumor sizes+/-SEM are depicted at day 12 after 4 therapeutic injections of anti-PD-1 mAbs, in each FMT groups (Akk+ and Akk-) supplemented or not with Akkp2261 as well as in animals reared in SPF conditions (FMT-). Concatenation of>25 experiments using n=53 mice in Iso group, n=51 in Iso FMT+ group, n=56 in anti-PD-1 and anti-PD-1 FMT+ groups. Each experiment comprising 6 mice/group and was performed at least 2 times for each FMT (Supplementary Table S6) (B). Tumor sizes according to FMT Akk- (C left, n=72/group; C right, n=49 in Iso group and n=48 in other groups) versus Akk+ (D left, n=6/group, D right, n=12 in Iso and anti-PD-1 groups, n=14 in anti-PD-1 with Akkp2261) are depicted, each dot representing one mouse. Statistics were mixed-effect modeling with specific software ((https://kroemerlab.shinyapps.io/TumGrowth/) for longitudinal tumor growth analysis (D) and Mann–Whitney U-test (B-C) to compare two independent groups (after Kruskal–Wallis test was implemented using Dunn’s test for multiple groups). ns=not significant. E. Clustermap of ratios of Akkp2261-related tumor reduction at day 12-15 following PD-1 mAbs in FMT normalized onto ratios obtained in SPF mice. The relative tumor size reduction follows a blue color code (the darker the greater; R, Responders)). 29 FMT were performed according to A. N=29-30 mice/group in total. Each experiment contained 6 mice/group and was performed 2-3 times for each tumor model (E, left panel). 16S rRNA sequencing of gene amplicons of stools harvested in recipient avatar tumor bearers at day 12 post-4 injections of anti-PD-1 Abs and 4 oral gavages with Akkp2261 divided into green (R) and red (NR) groups. VIP plot repartition of discriminant metagenomic species segregating groups of mice that responded to oral Akkp2261 (R, green bars) or not (NR, red bars). (E, right panel). Asterisks represent significant Mann-Whitney U test without FDR at 10%. * p <0.05, ** p <0.01, *** p <0.001. P-values were calculated using a two-sided nonparametric Wilcoxon sum-rank test. Adjustments for multiple comparisons were not made.
Supplementary information
Supplementary Information
Supplementary Tables 1–7
Supplementary Table 8
Detailed information on the cohort
Supplementary Data
Patients’ metadata
Source data
Source Data Fig. 1
Statistical source data for Fig. 1f–h
Source Data Fig. 2
Statistical source data for Fig. 2a,b
Source Data Fig. 3
Statistical source data for Fig. 3c,e,g
Source Data Extended Data Fig. 2
Statistical source data for Extended Data Fig. 2c–e
Source Data Extended Data Fig. 3
Statistical source data for Extended Data Fig. 3
Source Data Extended Data Fig. 4
Statistical source data for Extemded Data Fig. 4
Source Data Extended Data Fig. 5
Statistical source data for Extended Data Fig. 5d,e
Source Data Extended Data Fig. 6
Statistical source data for Extended Data Fig. 6a–d
Source Data Extended Data Fig. 6
DataFrame for Extended Dta Fig. 6e
Source Data Extended Data Fig. 6
Metadata for Extended Data Fig. 6e
Rights and permissions
About this article
Cite this article
Derosa, L., Routy, B., Thomas, A.M. et al. Intestinal Akkermansia muciniphila predicts clinical response to PD-1 blockade in patients with advanced non-small-cell lung cancer. Nat Med 28, 315–324 (2022). https://doi.org/10.1038/s41591-021-01655-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-021-01655-5
This article is cited by
-
Univariable and multivariable Mendelian randomization study identified the key role of gut microbiota in immunotherapeutic toxicity
European Journal of Medical Research (2024)
-
mEnrich-seq: methylation-guided enrichment sequencing of bacterial taxa of interest from microbiome
Nature Methods (2024)
-
Data-driven prediction of colonization outcomes for complex microbial communities
Nature Communications (2024)
-
A gut microbial signature for combination immune checkpoint blockade across cancer types
Nature Medicine (2024)
-
Aging induces changes in cancer formation and microbial content in a murine model of bladder cancer
GeroScience (2024)