Abstract
Amyotrophic lateral sclerosis (ALS) is a progressive, neurodegenerative disease of the lower and upper motor neurons with sporadic or hereditary occurrence. Age of onset, pattern of motor neuron degeneration and disease progression vary widely among individuals with ALS. Various cellular processes may drive ALS pathomechanisms, but a monogenic direct metabolic disturbance has not been causally linked to ALS. Here we show SPTLC1 variants that result in unrestrained sphingoid base synthesis cause a monogenic form of ALS. We identified four specific, dominantly acting SPTLC1 variants in seven families manifesting as childhood-onset ALS. These variants disrupt the normal homeostatic regulation of serine palmitoyltransferase (SPT) by ORMDL proteins, resulting in unregulated SPT activity and elevated levels of canonical SPT products. Notably, this is in contrast with SPTLC1 variants that shift SPT amino acid usage from serine to alanine, result in elevated levels of deoxysphingolipids and manifest with the alternate phenotype of hereditary sensory and autonomic neuropathy. We custom designed small interfering RNAs that selectively target the SPTLC1 ALS allele for degradation, leave the normal allele intact and normalize sphingolipid levels in vitro. The role of primary metabolic disturbances in ALS has been elusive; this study defines excess sphingolipid biosynthesis as a fundamental metabolic mechanism for motor neuron disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All data and material generated and analyzed within the main figures and supplementary material of this study are available upon request from the authors. All requests for raw and analyzed data and materials related to this article will be reviewed by the respective institution to verify whether the request is subject to any intellectual property or confidentiality obligations. Some patient-related data, including genetic sequencing data, not included in the paper or its supplements were generated as part of clinical care and may be subject to patient confidentiality. Any data and materials that can be shared will be released via a material transfer agreement. The following databases were searched for the presence of variants reported in this paper: the Genome aggregation database (https://gnomad.broadinstitute.org), the Decipher database (https://www.deciphergenomics.org), the Leiden database (https://www.lovd.nl) and the TopMed database (https://bravo.sph.umich.edu/freeze8/hg38/).
References
Al-Chalabi, A. & Hardiman, O. The epidemiology of ALS: a conspiracy of genes, environment and time. Nat. Rev. Neurol. 9, 617–628 (2013).
Swinnen, B. & Robberecht, W. The phenotypic variability of amyotrophic lateral sclerosis. Nat. Rev. Neurol. 10, 661–670 (2014).
Brown, R. H. & Al-Chalabi, A. Amyotrophic lateral sclerosis. N. Engl. J. Med. 377, 162–172 (2017).
Lee, Y. et al. Oligodendroglia metabolically support axons and contribute to neurodegeneration. Nature 487, 443–448 (2012).
Bannwarth, S. et al. A mitochondrial origin for frontotemporal dementia and amyotrophic lateral sclerosis through CHCHD10 involvement. Brain 137, 2329–2345 (2014).
Breslow, D. K. Sphingolipid homeostasis in the endoplasmic reticulum and beyond. Cold Spring Harb. Perspect. Biol. 5, a013326 (2013).
Harrison, P. J., Dunn, T. M. & Campopiano, D. J. Sphingolipid biosynthesis in man and microbes. Nat. Prod. Rep. 35, 921–954 (2018).
Wang, G. & Bieberich, E. Sphingolipids in neurodegeneration (with focus on ceramide and S1P). Adv. Biol. Regul. 70, 51–64 (2018).
Hojjati, M. R., Li, Z. & Jiang, X. C. Serine palmitoyl-CoA transferase (SPT) deficiency and sphingolipid levels in mice. Biochim. Biophys. Acta 1737, 44–51 (2005).
Zhao, L. et al. Elevation of 20-carbon long chain bases due to a mutation in serine palmitoyltransferase small subunit b results in neurodegeneration. Proc. Natl Acad. Sci. USA 112, 12962–12967 (2015).
Bejaoui, K. et al. SPTLC1 is mutated in hereditary sensory neuropathy, type 1. Nat. Genet. 27, 261–262 (2001).
Gantner, M. L. et al. Serine and lipid metabolism in macular disease and peripheral neuropathy. N. Engl. J. Med. 381, 1422–1433 (2019).
Gable, K. et al. A disease-causing mutation in the active site of serine palmitoyltransferase causes catalytic promiscuity. J. Biol. Chem. 285, 22846–22852 (2010).
Penno, A. et al. Hereditary sensory neuropathy type 1 is caused by the accumulation of two neurotoxic sphingolipids. J. Biol. Chem. 285, 11178–11187 (2010).
Brooks, B. R., Miller, R. G., Swash, M. Munsat, T. L. & World Federation of Neurology Research Group on Motor Neuron Diseases. El Escorial revisited: revised criteria for the diagnosis of amyotrophic lateral sclerosis. Amyotroph. Lateral Scler. Other Motor Neuron Disord. 1, 293–299 (2000).
Karczewski, K. J. et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581, 434–443 (2020).
Xiong, H. Y. et al. RNA splicing. The human splicing code reveals new insights into the genetic determinants of disease. Science 347, 1254806 (2015).
Han, G. et al. The topology of the Lcb1p subunit of yeast serine palmitoyltransferase. J. Biol. Chem. 279, 53707–53716 (2004).
Fernandopulle, M. S. et al. Transcription factor-mediated differentiation of human iPSCs into neurons. Curr. Protoc. Cell Biol. 79, e51 (2018).
Fridman, V. et al. Randomized trial of l-serine in patients with hereditary sensory and autonomic neuropathy type 1. Neurology 92, e359–e370 (2019).
Garofalo, K. et al. Oral L-serine supplementation reduces production of neurotoxic deoxysphingolipids in mice and humans with hereditary sensory autonomic neuropathy type 1. J. Clin. Invest. 121, 4735–4745 (2011).
Mandon, E. C., van Echten, G., Birk, R., Schmidt, R. R. & Sandhoff, K. Sphingolipid biosynthesis in cultured neurons. Down-regulation of serine palmitoyltransferase by sphingoid bases. Eur. J. Biochem. 198, 667–674 (1991).
Siow, D. L. & Wattenberg, B. W. Mammalian ORMDL proteins mediate the feedback response in ceramide biosynthesis. J. Biol. Chem. 287, 40198–40204 (2012).
Davis, D. L., Gable, K., Suemitsu, J., Dunn, T. M. & Wattenberg, B. W. The ORMDL/Orm-serine palmitoyltransferase (SPT) complex is directly regulated by ceramide: reconstitution of SPT regulation in isolated membranes. J. Biol. Chem. 294, 5146–5156 (2019).
Hjelmqvist, L. et al. ORMDL proteins are a conserved new family of endoplasmic reticulum membrane proteins. Genome Biol. 3, RESEARCH0027 (2002).
Han, G. et al. The ORMs interact with transmembrane domain 1 of Lcb1 and regulate serine palmitoyltransferase oligomerization, activity and localization. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 1864, 245–259 (2019).
Clarke, B. A. et al. The Ormdl genes regulate the sphingolipid synthesis pathway to ensure proper myelination and neurologic function in mice. eLife 8, e51067 (2019).
Li, S., Xie, T., Liu, P., Wang, L. & Gong, X. Structural insights into the assembly and substrate selectivity of human SPT-ORMDL3 complex. Nat. Struct. Mol. Biol. 28, 249–257 (2021).
Wang, Y. et al. Structural insights into the regulation of human serine palmitoyltransferase complexes. Nat. Struct. Mol. Biol. 28, 240–248 (2021).
Blackstone, C. Hereditary spastic paraplegia. Handb. Clin. Neurol. 148, 633–652 (2018).
Connolly, O. et al. A systematic review of genotype phenotype correlation across cohorts having causal mutations of different genes in ALS. J. Pers. Med. 10, 58 (2020).
Nicolas, G. & Veltman, J. A. The role of de novo mutations in adult-onset neurodegenerative disorders. Acta Neuropathol. 137, 183–207 (2019).
Cervenakova, L. et al. Progressive muscular atrophy variant of familial amyotrophic lateral sclerosis (PMA/ALS). J. Neurol. Sci. 177, 124–130 (2000).
Cudkowicz, M. E., McKenna-Yasek, D., Chen, C., Hedley-Whyte, E. T. & Brown, R. H. Jr. Limited corticospinal tract involvement in amyotrophic lateral sclerosis subjects with the A4V mutation in the copper/zinc superoxide dismutase gene. Ann. Neurol. 43, 703–710 (1998).
Auer-Grumbach, M. et al. Mutations at Ser331 in the HSN type I gene SPTLC1 are associated with a distinct syndromic phenotype. Eur. J. Med. Genet. 56, 266–269 (2013).
Suh, B. C. et al. Early-onset severe hereditary sensory and autonomic neuropathy type 1 with S331F SPTLC1 mutation. Mol. Med. Rep. 9, 481–486 (2014).
Rotthier, A. et al. Genes for hereditary sensory and autonomic neuropathies: a genotype–phenotype correlation. Brain 132, 2699–2711 (2009).
Bode, H. et al. HSAN1 mutations in serine palmitoyltransferase reveal a close structure–function–phenotype relationship. Hum. Mol. Genet. 25, 853–865 (2016).
Suriyanarayanan, S. et al. A novel variant (Asn177Asp) in SPTLC2 causing hereditary sensory autonomic neuropathy type 1C. Neuromolecular Med. 21, 182–191 (2019).
Harmon, J. M. et al. Topological and functional characterization of the ssSPTs, small activating subunits of serine palmitoyltransferase. J. Biol. Chem. 288, 10144–10153 (2013).
Blasco, H. et al. Lipidomics reveals cerebrospinal-fluid signatures of ALS. Sci. Rep. 7, 17652 (2017).
Cutler, R. G., Pedersen, W. A., Camandola, S., Rothstein, J. D. & Mattson, M. P. Evidence that accumulation of ceramides and cholesterol esters mediates oxidative stress-induced death of motor neurons in amyotrophic lateral sclerosis. Ann. Neurol. 52, 448–457 (2002).
Dodge, J. C. et al. Glycosphingolipids are modulators of disease pathogenesis in amyotrophic lateral sclerosis. Proc. Natl Acad. Sci. USA 112, 8100–8105 (2015).
Henriques, A. et al. Sphingolipid metabolism is dysregulated at transcriptomic and metabolic levels in the spinal cord of an animal model of amyotrophic lateral sclerosis. Front. Mol. Neurosci. 10, 433 (2017).
Alterman, J. F. et al. A divalent siRNA chemical scaffold for potent and sustained modulation of gene expression throughout the central nervous system. Nat. Biotechnol. 37, 884–894 (2019).
Kim, J. et al. Patient-customized oligonucleotide therapy for a rare genetic disease. N. Engl. J. Med. 381, 1644–1652 (2019).
Pacold, M. E. et al. Corrigendum: a PHGDH inhibitor reveals coordination of serine synthesis and one-carbon unit fate. Nat. Chem. Biol. 12, 656 (2016).
Lowther, J. et al. Inhibition of the PLP-dependent enzyme serine palmitoyltransferase by cycloserine: evidence for a novel decarboxylative mechanism of inactivation. Mol. Biosyst. 6, 1682–1693 (2010).
Carvalho, M. D. & Swash, M. Awaji diagnostic algorithm increases sensitivity of El Escorial criteria for ALS diagnosis. Amyotroph. Lateral Scler. 10, 53–57 (2009).
Han, G. et al. Identification of small subunits of mammalian serine palmitoyltransferase that confer distinct acyl-CoA substrate specificities. Proc. Natl Acad. Sci. USA 106, 8186–8191 (2009).
Gupta, S. D. et al. Expression of the ORMDLS, modulators of serine palmitoyltransferase, is regulated by sphingolipids in mammalian cells. J. Biol. Chem. 290, 90–98 (2015).
Mohassel, P. et al. Dominant collagen XII mutations cause a distal myopathy. Ann. Clin. Transl. Neurol. 6, 1980–1988 (2019).
Miyaoka, Y. et al. Isolation of single-base genome-edited human iPS cells without antibiotic selection. Nat. Methods 11, 291–293 (2014).
Lone, M. A. et al. Subunit composition of the mammalian serine-palmitoyltransferase defines the spectrum of straight and methyl-branched long-chain bases. Proc. Natl Acad. Sci. USA 117, 15591–15598 (2020).
Merrill, A. H. Jr., Sullards, M. C., Allegood, J. C., Kelly, S. & Wang, E. Sphingolipidomics: high-throughput, structure-specific, and quantitative analysis of sphingolipids by liquid chromatography tandem mass spectrometry. Methods 36, 207–224 (2005).
Acknowledgements
We thank the patients and their families for participating in our research study, C. Mendoza (NINDS/NNDCS) and G. Averion (NINDS/NNDCS) for their help in supporting the clinic, F. Eichler (Massachusetts General Hospital) for helpful discussions, and B. Smith (NINDS) and A. Nath (NINDS) for providing control serum samples. We also thank the NIH Intramural Sequencing Center for performing exome sequencing; K. Linask and J. Zhou from the NHLBI iPSC core for generating the isogenic mutant iPSC lines; and S. Lara and S. J. H. Tao Cheng in the NINDS electron microscopy facility. Work in C.G.B.’s laboratory is supported by intramural funds of NINDS/NIH. Work in T.M.D.’s laboratory was supported by CDMRP grant W81XWH-20-1-0219. C.V.L is supported by a career development grant from NIH/NINDS (K08 NS10762). T.H. is supported by Swiss National Science Foundation grant 31003A_179371. M.A.L is supported by Swiss Foundation for Research on Muscle Diseases (FSRMM). R.H.B. is funded by NIH/NINDS (R01 NS072446) and the Deater foundation. Sequencing and analysis provided by the Broad Institute of MIT and Harvard Center for Mendelian Genomics (Broad CMG) was funded by the National Human Genome Research Institute, the National Eye Institute and the National Heart, Lung and Blood Institute (grant UM1 HG008900) and, in part, by the National Human Genome Research Institute (grant R01 HG009141).
Author information
Authors and Affiliations
Contributions
P.M. performed laboratory experiments, evaluated patients, analyzed data and drafted the manuscript. S.D., A.R.F., J.A.M.S., A.L.M., S.B.N., D.S., F.K., A.I., G.L., A.G., H.K., C.F., C.V.L., A.T., L.B., S.S., A.Z., H.P., E.M., F.P.T., C.G.K., M.T.C., T.B., V.S., A.M.C., U.S., A.R., R.H.B., M.T., K.R.C., A.H., C.-H.L., Z.P. and C.E.L.P., provided and interpreted clinical and genetic information, crucial samples and material and/or analyzed data. M.N., M.A.L., K.G., S.D.G., Y.H., C.G., A.R.N., N.P. and T.H. performed laboratory experiments and/or analyzed data. T.M.D. and C.G.B. designed the overall study, interpreted data and drafted and revised the manuscript. All authors reviewed and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Medicine thanks Merit Cudkowicz, Ammar Al-Chalabi and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Jerome Staal was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Sphingolipid biosynthesis pathway.
Serine palmitoyltransferase (SPT) catalyses the first and rate-limiting step in sphingolipid biosynthesis. The most abundant acyl-CoA used by SPT is palmitoyl-CoA, which after condensation with L-serine results in an 18-carbon long-chain base, 3-dehydrosphinganine. 3-dehydrosphinganine is rapidly reduced by 3-dehydrosphinganine reductase (KDSR) to form dihydrosphingosine (sphinganine). Sphinganine is acylated by a variety of ceramide synthases (CerS) to form dihydroceramides, which can be converted to ceramides by sphingolipid desaturase (DEGS). Many complex sphingolipids can be synthesized from a ceramide backbone. However, the only true exit from the pathway depends on activity of sphingosine kinase (SK) and sphingosine-1-phosphate aldolase (SGPL) to form phosphoethanolamine. SGPP = sphingosine-1-phosphate phosphatase.
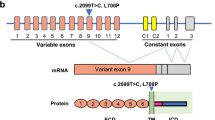
Extended Data Fig. 2 SPTLC1 variant associated with amyotrophic lateral sclerosis and their effects on splicing.
a, Gel electrophoresis of PCR amplification of cDNA obtained from patient fibroblast cultures or whole blood. Fibroblasts were not available for c.58G>T patient. PCR primers were in exon 1 and 6 of SPTLC1 (NM_006415.4). SPTLC1 c.58G>T variant resulted in two bands with an apparent difference of ~100 bp. All other variants and controls resulted in a single band. b, Sequencing of the gel purified PCR product showed that the c.58G>T variant resulted in complete skipping of exon 2 (lower band). Neither the full-length product (upper band) nor the internally deleted transcript (lower band) contained the c.58G>T variant. However, it is possible that the missense variant exists in very small amounts and escaped amplification by PCR and sequencing. All other variants were confirmed and do not affect splicing.
Extended Data Fig. 3 Amyotrophic lateral sclerosis associated variant SPTLC1 cellular localization.
a, Immunostaining for SPTLC1 in patient derived fibroblast cultures shows normal subcellular localization in these cells, with an apparent distribution in the endoplasmic reticulum. Immunostaining for TDP43 also appears normal. Scale bar=40 μm b, Immunostaining for SPTLC1 in iPSC-derived human motor neurons with p.F40_S41 variant do not show any changes in its subcellular localization. Scale bar=20 μm c, Western blot of SPTLC1 in iPSC-derived human motor neurons confirms comparable expression levels in p.F40_S41 line and its isogenic control. Two independent differentiation experiments followed by western blotting were performed with similar results.
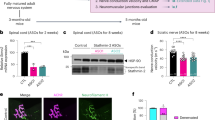
Extended Data Fig. 4 Induced pluripotent stem cell-derived human motor neurons (iPSC-hMN) with SPTLC1 ALS variants.
a, After differentiation by addition of doxycycline, SPTLC1 mutant iPSC-hMNs, one with heterozygous SPTLC1 p.F40_S41del variant and the other with compound heterozygous variants SPTLC1 p.F40_S41del; E2del (complete deletion of exon 2) and their isogenic control assume neuronal morphology. They express choline acetyltransferase (ChAT), Hb9 (a motor neuron specific transcription factor) as well as Tuj1 (neuronal microtubule marker). Scale bar = 20 μm b, Electron microscopy of iPSC-hMNs shows normal neuronal cell body morphology, nuclear heterochromatin, and organelles. In the processes (right), microtubules and occasional mitochondria and vesicles are noted. Scale bar left = 1 μm, middle and right = 200 nm. c, Sphingolipidomic analysis of de novo sphingolipid synthesis using isotope labelling (with 3,3-D2 L-serine) depicted in the form a heatmap with each row representing a sphingolipid species. Row Z-scores were calculated and depicted in colour. Canonical sphingolipids ceramides and glucosyl ceramides (glucCer) are increased in p.F40_S41del iPSC-hMNs compared to their isogenic control and increase further in the double mutant iPSC-hMNs. The bar graphs show the increase in total 2+ long-chain bases (LCB), ceramides (Cer), and glucosyl ceramides (GlucCer).
Extended Data Fig. 5 Allele specific knockdown of variant SPTLC1 mRNA.
a, The two allele-specific siRNAs targeting the L39del and F40_S41del alleles do not reduce SPTLC1 mRNA levels in control cells at concentrations as high as 100nM. Error bars are SEM. Four replicates were performed. b, Sphingolipidomic analysis of patient derived fibroblasts with SPTLC1 ALS variants (F40_S41del and two patient lines with L39del) after treatment with 10 nM allele-specific siRNA targeting each corresponding variant were analysed and compared to control fibroblasts. Following the knockdown, the cells were labeled with D3,15N-L-serine and D4-L-alanine. The de novo synthesized +3 ceramide and sphingomyelin levels show a reduction in patient cells when treated with variant-specific siRNAs. Error bars are SEM. Three replicates were performed.
Supplementary information
Supplementary Information
Supplementary Tables 1–6.
Supplementary Video 1
Clinical findings on bedside neurologic evaluation of patients with SPTLC1-associated amyotrophic lateral sclerosis. Patient 1 (p.Y23F) and patient 3 (p.F40_S41del) are depicted, highlighting tongue fasciculations, severe upper extremity weakness and ankle clonus.
Supplementary Video 2
Skeletal muscle ultrasound in a patient with SPTLC1-associated amyotrophic lateral sclerosis. Ultrasound of patient 1 (p.Y23F) shows diffuse fasciculations. In addition, skeletal muscles appear atrophied and show increased echogenicity with a mixed granular and streaking pattern.
Rights and permissions
About this article
Cite this article
Mohassel, P., Donkervoort, S., Lone, M.A. et al. Childhood amyotrophic lateral sclerosis caused by excess sphingolipid synthesis. Nat Med 27, 1197–1204 (2021). https://doi.org/10.1038/s41591-021-01346-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-021-01346-1
This article is cited by
-
Possibilities and limitations of antisense oligonucleotide therapies for the treatment of monogenic disorders
Communications Medicine (2024)
-
Disruption of MAM integrity in mutant FUS oligodendroglial progenitors from hiPSCs
Acta Neuropathologica (2024)
-
Diagnostic model based on key autophagy-related genes in intervertebral disc degeneration
BMC Musculoskeletal Disorders (2023)
-
Mutation screening of SPTLC1 and SPTLC2 in amyotrophic lateral sclerosis
Human Genomics (2023)
-
Structure of the ceramide-bound SPOTS complex
Nature Communications (2023)