Abstract
The first severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection in South Africa was identified on 5 March 2020, and by 26 March the country was in full lockdown (Oxford stringency index of 90)1. Despite the early response, by November 2020, over 785,000 people in South Africa were infected, which accounted for approximately 50% of all known African infections2. In this study, we analyzed 1,365 near whole genomes and report the identification of 16 new lineages of SARS-CoV-2 isolated between 6 March and 26 August 2020. Most of these lineages have unique mutations that have not been identified elsewhere. We also show that three lineages (B.1.1.54, B.1.1.56 and C.1) spread widely in South Africa during the first wave, comprising ~42% of all infections in the country at the time. The newly identified C lineage of SARS-CoV-2, C.1, which has 16 nucleotide mutations as compared with the original Wuhan sequence, including one amino acid change on the spike protein, D614G (ref. 3), was the most geographically widespread lineage in South Africa by the end of August 2020. An early South African-specific lineage, B.1.106, which was identified in April 2020 (ref. 4), became extinct after nosocomial outbreaks were controlled in KwaZulu-Natal Province. Our findings show that genomic surveillance can be implemented on a large scale in Africa to identify new lineages and inform measures to control the spread of SARS-CoV-2. Such genomic surveillance presented in this study has been shown to be crucial in the identification of the 501Y.V2 variant in South Africa in December 2020 (ref. 5).
Similar content being viewed by others
Main
SARS-CoV-2 is a novel betacoronavirus, first detected in China in December 2019 (refs. 6,7). Since then, Coronavirus Disease 2019 (COVID-19) has developed into a global pandemic, resulting in several waves of epidemics, infecting over 85 million people and causing over 1.9 millions deaths globally by January 9, 2021. Lockdown and travel restriction measures have varied from country to country, which has dictated the profile of local outbreaks. Through the sharing of SARS-CoV-2 sequences during this pandemic, including from one of the first cases in Wuhan, China (MN908947.3)7, genomic epidemiology investigations are playing a major role in characterizing and understanding this emerging virus8,9,10,11,12,13. SARS-CoV-2 has typically been classified into two main phylogenetic lineages: lineage A and lineage B. Although both lineages originated in China, lineage A spread from Asia to the rest of the world, whereas lineage B predominantly spread from Europe14.
The COVID-19 epidemic in South Africa is by far the largest in Africa, with more than 785,000 individuals infected and more than 20,000 deaths by end of November 2020. The first case of SARS-CoV-2 infection in South Africa was recorded in KwaZulu-Natal (KZN) Province on 5 March 2020, in a traveler returning from Italy15. Around mid-March, cases of community transmission were reported across the country. The profile of SARS-CoV-2 epidemiological progression in South Africa was largely influenced by the implementation of lockdown measures in the early phases of the epidemic and the subsequent easing of these measures. On 26 March 2020, the government-imposed nationwide lockdown included the prohibition of all gatherings, travel restrictions and closure of schools and non-essential businesses (Oxford stringency index of 90, or commonly known in South Africa as level 5 lockdown; Supplementary Table 1)16. Although the epidemic was growing, lockdown measures were progressively eased on 1 May 2020 (level 4), and on 1 June 2020 (level 3), to mitigate negative effects on the country’s economy. For example, by 1 June, interprovincial travel was allowed, and there was no curfew on the movement of people. Restrictions were further relaxed on 17 August (level 2), allowing restaurants and bars to open. More restrictions were lifted on October 1 (Fig. 1a) once the initial peak of new daily infections had passed, allowing students to return to university campuses and South Africa to return to normality. The epidemic in South Africa can generally be characterized by two phases: one dominated by travel-related early introductions, the other being the period of peak infections (Fig. 1a).
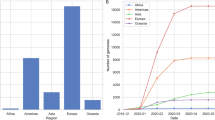
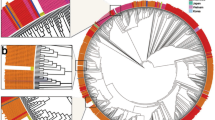
a, Epidemiological curve showing the progression of daily COVID-19 numbers in South Africa, changes in Re estimations (mean estimated median Re with upper and lower bounds of the 95% confidence interval shown), lockdown levels and the timing of genomic sampling in South Africa from the beginning of the epidemic to 15 September. b, Estimated numbers of introductions into South Africa colored by region of origin. c, Overall sampling of genomes in South Africa colored by whether the genomes are associated with introduction events (origins outside South Africa) or not (origins in South Africa). d, MCC tree of 7,213 global genomes including 1,365 South African sequences, indicating a period of early introductions and a period of peak infection separated by a period of emergence of new lineages. The three largest monophyletic lineage clusters in South Africa, along with the early B.1.106 South African lineage, are labeled.
We monitored the likelihood of SARS-CoV-2 transmission by estimating the effective reproduction number (Re), which provides a measure of the average number of secondary infections caused by an infected person17. Typically, a growing epidemic is characterized by Re >1, and Re <1 indicates a slowed progression. At the start of the epidemic, in mid-March 2020, we estimated the Re value to be >3, quickly falling after the start of lockdown to a value of <1 in late March 2020. A subsequent jump in the Re value to >1 in April 2020 was found to be concurrent with the timing of several localized outbreaks in the country, including nosocomial outbreaks18. The Re value again dropped to <1 at the beginning of August 2020, coinciding with a decrease in the daily number of positive cases recorded (Fig. 1a).
Genomic epidemiology is important to understand SARS-CoV-2 evolution and track the dynamics of transmission across the world8,9,10,11,12,13. By 15 September 2020, at the tail end of the first epidemiological peak in the country, we had produced 1,365 SARS-CoV-2 genomic sequences (>90% coverage; publicly shared on GISAID19) in our laboratories as part of the Network for Genomic Surveillance in South Africa (NGS-SA) consortium20. These genomes were sampled between 6 March and 26 August 2020 in eight of the nine provinces of South Africa and in all the districts of KZN Province (Extended Data Fig. 1) and represented consistent sampling from the beginning of the epidemic and corresponding to important events of the epidemiological progression (Fig. 1a).
We estimated maximum likelihood (ML) and molecular clock phylogenies for a dataset containing 7,213 global genomes, including 1,365 South African genomes, sampled from 24 December 2019 to 26 August 2020 (Fig. 1c). Time-measured phylogeographic analyses estimated at least 101 introductions into South Africa. The bulk of imported introductions happened before lockdown (26 March 2020) from Europe, when the epidemic was most quickly progressing (Fig. 1b). Although at least 67 introduction events are inferred to have occurred after lockdown, these represent only 5% of the genomes that were sampled after lockdown (Fig. 1c). In the early phases of the epidemic, before 1 April, 34 introductions were inferred from 35 genomes sampled (97.1%), which we call early introductions (Fig. 1b). The small number of apparent introductions after lockdown could be explained by more intensive genomic sampling at later stages, which likely revealed introduction events linked to previously undetected transmission chains.
The early introductions were mostly isolated cases with a few occurrences of small onward transmission clusters, by contrast with large transmission clusters during the peak infection phase (Fig. 1d). The period between these two phases was inferred to be characterized by localized transmission events, which saw the emergence and spread of new lineages, which were later amplified during the first peak of the epidemic. The South African genomes in this study were assigned to 42 different lineages based on the proposed dynamic nomenclature for SARS-CoV-2 lineages14. This included 16 South Africa-specific lineages, defined by cov-lineages.org as being lineages that are presently predominant in South Africa as of 15 September 2020 (ref. 21) (Extended Data Fig. 2). One of these has been assigned a novel SARS-CoV-2 main lineage classification, lineage C, the parent of which is lineage B.1.1.1.
Extensive SARS-CoV-2 genomic sampling, which has spanned the duration of the epidemic to date and was analyzed until the end of the first wave in this study, enabled for such lineage emergence to be observed, similar to the genomic investigation of SARS-CoV-2 in the United Kingdom22. During the first wave of the epidemic in South Africa, until 15 September 2020, a total of 42 detectable SARS-CoV-2 phylogenetic lineages were circulating in the country, with an average of around ten lineages circulating per epidemiological week, peaking at 24 in the weeks of highest infections. During the same time frame, more than 1,000 such transmission lineages were circulating in the United Kingdom23. We focused on the three largest monophyletic lineage clusters (C.1, B.1.1.54 and B.1.1.56) that spread in South Africa during lockdown and then grew into large transmission clusters during the peak infection phase of the epidemic (Fig. 1d).
B.1.1.54, B.1.1.56 and C.1 were the three largest monophyletic clusters of observed South African lineages that emerged and spread in the country after lockdown and into the first peak of the epidemic. They contain 320, 104 and 151 genomes, respectively, which represent 42.1% of the total genomes in this study (Extended Data Fig. 2), with a clear overrepresentation from mid-May to September 2020 (Fig. 2d). Genomes belonging to these lineages were sampled in five adjacent provinces of South Africa and in all 11 districts of KZN Province (Fig. 2b,c and Extended Data Fig. 3) and corresponded to time points spanning from 31 March 2020 to 26 August 2020 (Fig. 2b,c). We compared cycle threshold (Ct) scores, as approximations of viral loads, for genomes for which this was measured (n = 653) and found no significant difference between the Ct scores of sequences belonging to these three lineages and the others (Extended Data Fig. 4). This suggests that the fast spread of the lineages of interest is likely a result of localized outbreaks and expected transmission dynamics, rather than caused by any fitness advantage, with the caveat that samples with Ct scores measured might have been collected at different times during the course of infection, which could obscure lineage-associated differences.
a, Mapping the spread of the B.1.1.56 cluster (left) and the C.1 cluster (right) from phylogeographic reconstructions. Time scale is specified in decimal dates from 2020.2 (March 2020) to 2020.6 (July 2020). b, Sampling timeline and locations of genomes belonging to each lineage cluster in five provinces. c, Sampling timeline and locations of genomes belonging to each lineage cluster in all 11 districts of KZN Province. d, The progression of the proportions of genomes belonging to the main lineage clusters over time.
To provide details on the spatiotemporal diffusion of South African-specific lineages, we used a continuous phylogeographic model that maps the phylogenetic nodes to their inferred geographical origin locations (Fig. 2a). Bayesian Markov chain Monte Carlo (MCMC) analysis suggests that these lineages emerged between 15 February and 24 May 2020 (Extended Data Fig. 6). Our phylogeographic reconstruction suggests that lineage B.1.1.56 emerged in the city of Durban (eThekwini) around mid-March 2020 (95% highest posterior density (HPD) 2020-02-15 to 2020-03-30). It appears that, from June onwards, this lineage quickly disseminated to all of the districts in KZN Province. This occurred when the country moved from lockdown level 4 to 3, which allowed for increased movement of people and goods between districts. Lineage C.1 most likely emerged in early May 2020 (95% HPD 2020-04-24 to 2020-05-24) in the city of Johannesburg, located in Gauteng Province, from where it quickly spread to the adjacent North West Province, causing a large nosocomial outbreak24. Furthermore, the lineage spread through two independent events to the northern province of Limpopo and to northwestern KZN. From this location, the lineage further spread into all districts of KZN and to the adjacent Free State Province. Unfortunately, lineage B.1.1.54 showed poor temporal signaling (Extended Data Fig. 5), and, therefore, Bayesian spatiotemporal analyses could not be performed for this cluster. A closer look at the cluster (from the ML timetree) is, however, shown in Extended Data Fig. 6 and indicates that this lineage was first sampled in KZN and Gauteng and later spread in large numbers in the provinces of KZN, North West and Free State.
We analyzed the sequences of the three main lineage clusters to determine their lineage-defining mutations, if any. On average, sequences in the C.1 cluster accumulated roughly 16 mutations, whereas B.1.1.56 and B.1.1.54 have approximately 13–14 mutations relative to the Wuhan reference (MN908947.3) (Fig. 3a). This is relatively higher than the number of acquired mutations in other sequences as of 26 August 2020, which is consistent with these three lineages having emerged more recently than others in the study, hence accumulating more genomic changes. Sequences are assigned lineages based on the presence of certain lineage-defining mutations (Extended Data Fig. 7). The sequences belonging to B.1.1.54, B.1.1.56 and C.1 all have the mutations that define their B.1.1 parental lineage (C.1 was previously known as B.1.1.1.1) (Fig. 3b), including the 23403A>G (spike D614G) mutation, with additional mutations that differentiate them (Fig. 3b). Sequences in B.1.1.54 have the 12503T>C (NSP8: Y138H) and 29721C>T mutations in >90% frequency, similar to 22675C>T for B.1.1.56 and 4002C>T (NSP3: T428I), 10097G>A (3C-like proteinase: G15S), 13536C>T, 18747C>T and 23731C>T for C.1 (Fig. 3b). The early hospital-linked lineage B.1.106 was defined by the 16376C>T (helicase: P47L) mutation. Five of these mutations—12503T>C, 16376C>T, 18747C>T 29721C>T and 22675C>T—are predominantly present in South African SARS-CoV-2 genomes, with just a few occurrences found elsewhere globally (Fig. 3c and Extended Data Fig. 8), whereas other lineage-defining mutations are also common in other countries (Extended Data Fig. 8). There are two other highly prevalent nucleotide mutations on the spike protein in the B.1.56 and C.1. lineages: 22675C>T and 23731C>T; however, these are synonymous mutations and distinct from those identified in 501Y.V2 (ref. 5).
a, Violin plot showing the number of mutations in each cluster. b, Variant maps of the most common mutations in each cluster mapped against the SARS-CoV-2 genome structure. Most common mutations defined as mutations present in more than 90% of the genomes in that group. c, Change in frequency of some unique South African mutations over time in South Africa versus the rest of the world.
Major contributors to lineage amplifications in South Africa were hospital outbreaks. For example, lineage C.1 was amplified in a nosocomial outbreak in the North West Province in April 2020 (ref. 24) before spreading to KZN and other provinces. Another South African lineage, B.1.106, also emerged in a nosocomial outbreak in KZN Province in April 2020. This was a large outbreak that infected 88 healthcare staff and 47 patients and dominated most of the early infections in Durban, South Africa (Fig. 4b). This nosocomial outbreak attracted national attention as it was responsible for 14% of the infections in KZN and over 45% of the national deaths by early April 2020. We used genetic sequencing, together with active outbreak investigation, to understand how the virus entered and spread in this hospital4. This lineage also spread to the population and was associated with a second nosocomial outbreak in a nearby hospital that involved 26 healthcare workers (Fig. 4a). These two nosocomial outbreaks were identified within days of the first infection and were followed by active infection and prevention control measures4,18. The B.1.106 lineage largely subsided after the outbreak investigations and isolation of all infected individuals. The B.1.106 lineage’s prevalence at the population level decreased quickly after June 2020 (Fig. 4b).
a, Phylogenetic tree of B.1.1.106 sequences by nosocomial outbreak (designated CH1 and CH3) and district location. b, Proportion of sequences classified as B.1.106 over time in KZN. c, Mapping the spread of the B.1.106 lineage from phylogeographic reconstructions. Time scale is specified in decimal dates from 2019.8 (October 2019) to 2020.5 (June 2020). Shade circular patterns represent confidence intervals of location estimation.
We report an in-depth analysis of the spread of SARS-CoV-2 in South Africa from 6 March 2020 to 26 August 2020, showing that the bulk of introductions happened before lockdown and travel restrictions were implemented on 26 March 2020. However, despite drastic lockdown measures, the pandemic spread quickly, causing over 785,000 laboratory-confirmed infections by November 2020. To track the evolution of the virus in real time, we formed the NGS-SA25, a consortium of genomics and bioinformatics scientists who worked with national government laboratories to quickly generate and analyze data in the country. We produced 1,365 SARS-CoV-2 genomes and mapped the emergence of 16 novel lineages in South Africa. We found that three main lineages were responsible for almost half of all the infections in South Africa as of 15 September 2020. Despite a relative sequencing bias in KZN Province, we were able to detect these major lineages across multiple provinces. It is, therefore, likely that more extensive sampling throughout the country could detect the spread of these lineages nationally, especially during the period when lockdown levels were eased and mobility increased. Indeed, data from Cape Town also later identified 27 sequences available in GISAID of the C.1 lineage19 (EPI_ISL_660121-EPI_ISL_660150, EPI_ISL_660158). B.1.1.54, B.1.1.56 and C.1 were the most geographically widespread lineages in South Africa during the time of this study.
Genomic data were also used in real time to identify and control nosocomial outbreaks. The B.1.106 lineage, which was the first South African lineage to be identified, was leveraged to document how the virus spread inside a large hospital in Durban, KZN. The lessons learned in this outbreak were used to quickly control a second nosocomial outbreak in a nearby hospital. The active outbreak response, investigation and isolation of positive cases might have limited the spread of this lineage. Our analysis, therefore, shows that several SARS-CoV-2 lineages, each with unique mutations, emerged within localized epidemics during lockdown even as the introduction of new lineages from outside South Africa was being curbed.
That many of the mutations in our analysis are synonymous and that differences in Ct values do not seem to be affected by the infecting viral strain argue against selection for fitter variants, which contrast with reported characteristics of variant 501Y.V2 (ref. 5). All four of the main lineages reported in the current study contain the D614G mutation in the spike gene. Furthermore, the D614G mutation is found in 1,350 (99%) of the South African sequences. Although we are currently investigating any fitness cost associated with the different lineages, we found only three other non-synonymous mutations in spike (A688V, G769V and A1078S) with frequency ranging from 1.2% to 3.6% in this dataset, which suggests that the evolutionary stability of SARS-CoV-2 and, in particular, the spike protein was maintained in South Africa during the first wave of the pandemic. However, during this study, whether any of the low-prevalence spike mutations reported could have a fitness advantage in terms of transmission, viral replication or reduced antigenicity was unknown. That said, we remain vigilant that there remain small recurrent gaps in our genome sequences in potentially important regions, especially in some of our lower-quality sequences. We think that these gaps might have been introduced due to potential primer mismatch in small parts of the ORF1b, S and ORF3a genes (Extended Data Fig. 9). However, as we enter the period where re-infections and re-introduction of the viruses from international travelers is becoming more frequent, pre-existing immune responses could exert enough pressure on SARS-CoV-2 to select for resistance mutations. The dynamic nature of the COVID-19 epidemic in South Africa, and globally, supports the case for continued genomic surveillance of SARS-CoV-2. We are currently investigating limits to cross-reactivity among strains. Limited cross-reactivity could lead to effects such as antibody-dependent enhancement (ADE) in response to a vaccine with a non-native strain. ADE occurs in infections such as dengue when a previously infected individual is infected with a second strain of virus, which antibodies from the first infection can bind to, but not neutralize, viral proteins20. There is a chance that this could also happen to SARS-CoV-2 if the pandemic is not controlled over a long time, providing a greater opportunity for viral evolution, which could potentially affect the efficacy of current vaccines.
This study emphasizes the usefulness of integrating genomic surveillance methods to document and help control SARS-CoV-2 spread in local and national settings. Genomics data can also be used in real time to inform and consolidate national outbreak investigation and response strategies widely throughout Africa.
Methods
Ethical statement
We obtained de-identified remnant nasopharyngeal and oropharyngeal swab samples from patients testing positive for SARS-CoV-2 by reverse transcription quantitative polymerase chain reaction (RT–qPCR) from public health and private medical diagnostics laboratories (Supplementary Table 3). The project was approved by University of KwaZulu-Natal Biomedical Research Ethics Committee (protocol reference no. BREC/00001195/2020; project title: COVID-19 transmission and natural history in KwaZulu-Natal, South Africa: epidemiological investigation to guide prevention and clinical care). This project was also approved by the University of the Witwatersrand Human Research Ethics Committee (clearance certificate no. M180832; project title: Surveillance for outpatient influenza-like illness and asymptomatic virus colonization in South Africa). Sequence data from the Western Cape were approved by the Stellenbosch University Health Research Ethics Committee (reference no. N20/04/008_COVID-19; project title: COVID-19: sequencing the virus from South African patients). Patient consent was not required for the genomic surveillance; this requirement was waived by the research ethics committees.
Epidemiological data
We analyzed COVID-19 case counts in South Africa from publicly released data up to 15 September 2020, from the National Department of Health and the National Institute for Communicable Diseases in South Africa. This was accessible through the repository of the Data Science for Social Impact Research Group at the University of Pretoria (https://github.com/dsfsi/covid19za)26. The National Department of Health releases daily updates on the number of new confirmed cases, deaths and recoveries, with a breakdown by province. For correlation with government epidemic control measures, information from government press releases and speech transcripts was extracted. To illustrate the epidemic progression, the daily number of confirmed cases for South Africa was plotted alongside a timeline of lockdown levels and variation in estimated virus reproduction number until 15 September 2020.
Estimation of Re
The estimations for effective daily Re of SARS-CoV-2 in South Africa were obtained from the COVID-19-Re data repository (https://github.com/covid-19-Re/dailyRe-Data)17 as of 15 September 2020. The effective Re describes the average number of secondary infections caused by an infected individual. As described previously17, the relevant method of calculation of Re builds upon another method developed by Cori et al.27, accessible through the EpiEstim R package. Instead of using a time series of infection incidence, which cannot be observed directly, the relevant method infers the infection incidence time series based on secondary sources of information, such as COVID-19 confirmed case data, hospital admissions and deaths. This was considered in combination with two other sets of time variables: 1) the duration of SARS-CoV-2 incubation period and 2) the time delays between onset of symptoms and a positive test, a hospital admission or the death of a patient. The relevant method infers infection time series from the stated observed incidence data by deconvolution28,29.
SARS-CoV-2 samples and metadata
Residual samples from nasopharyngeal and oropharyngeal swabs collected from patients positive for COVID-19 that were obtained from all 11 districts of KZN Province were used for SARS-CoV-2 whole-genome sequencing. We obtained samples in the form either of primary swabs or extracted RNA. The swab samples were heat inactivated in a water bath at 60 °C for 30 min, in a biosafety level 3 laboratory, before RNA extraction. RNA was extracted using the Viral NA/gDNA Kit on the chemagic 360 system (PerkinElmer) using the automated chemagic 360 insturment (PerkinElmer) or manually using the Qiagen Viral RNA Mini Kit (Qiagen). Associated metadata for the samples included date and location (district) of sampling and sex and age of the patients.
Real Time RT–PCR
To detect the SARS-CoV-2 virus by PCR, the TaqPath COVID-19 CE-IVD RT–PCR Kit (Life Technologies) was used according to the manufacturer’s instructions. The assays target genomic regions (ORF1ab, S protein and N protein) of the SARS-CoV-2 genome. RT–PCR was performed on a QuantStudio 7 Flex Real-Time PCR instrument (Life Technologies). Ct values were analyzed using auto-analysis settings with the threshold lines falling within the exponential phase of the fluorescence curves and above any background signal.
Whole-genome sequencing and genome assembly
Complementary DNA (cDNA) synthesis was performed on the RNA using random primers followed by gene-specific multiplex PCR using the ARTIC protocol30. Briefly, extracted RNA was converted to cDNA using the Superscript IV First Strand synthesis system (Life Technologies) and random hexamer primers. SARS-CoV-2 whole-genome amplification by multiplex PCR was carried out using primers designed on Primal Scheme (http://primal.zibraproject.org/) to generate 400-bp amplicons with an overlap of 70 bp that covers the 30-kb SARS-CoV-2 genome. PCR products were cleaned up using AMPure XP purification beads (Beckman Coulter) and quantified using the Qubit dsDNA High Sensitivity assay on the Qubit 4.0 instrument (Life Technologies).
The Illumina Nextera Flex DNA Library Prep Kit was used according to the manufacturer’s protocol to prepare uniquely indexed paired-end libraries of genomic DNA. Sequencing libraries were normalized to 4 nM, pooled and denatured with 0.2 N sodium acetate. A 12 pM sample library was spiked with 1% PhiX (PhiX Control v3 adapter-ligated library used as a control). Libraries were loaded onto a 500-cycle v2 MiSeq Reagent Kit and run on the Illumina MiSeq instrument (Illumina).
Raw reads coming from Illumina sequencing were assembled using Genome Detective 1.126 (https://www.genomedetective.com/) and the Coronavirus Typing Tool31,32. The initial assembly obtained from Genome Detective was polished by aligning mapped reads to the references and filtering out low-quality mutations using the bcftools 1.7-2 mpileup method. All mutations were confirmed visually with BAM files using Geneious software (Biomatters). All of the sequences were deposited in GISAID (https://www.gisaid.org/)19, and the GISAID accession was included as part of Supplementary Table 1.
Compilation of the SARS-CoV-2 South Africa dataset
To present a comprehensive analysis of the genomic epidemiology of SARS-CoV-2 in South Africa, the genomes generated as of 15 September 2020 (n = 1,111) were combined with all other South African genomes available in GISAID on the same date (n = 298). Appropriate acknowledgement was given to the sequencing laboratories (Supplementary Table 2), and this resulted in a dataset of 1,409 genomes. Sampling locations of genomes in this dataset included all provinces in South Africa and all districts in KZN Province, which was the most sampled province (Extended Data Fig. 1). Collection dates spanned from 6 March 2020 (the first cases in South Africa) to 26 August 2020.
Quality control of genome sequences
Before phylogenetic reconstruction, we filtered out low-quality sequences from the dataset. We retrieved all South African SARS-CoV-2 genotypes from the GISAID database as of 26 August 2020 (n = 1,409). We filtered out all genotypes that met any of the following criteria: 1) sequences with less than 90% genotype coverage; 2) genotypes with too many mutations (defined as having >20 nucleotide mutations relative to the Wuhan reference), which would violate the SARS-CoV-2 molecular clock at the time of study; 3) genotypes with more than ten ambiguous bases; and 4) genotypes with clustered mutations, defined as mutations in close proximity to one another. These are the standard quality assessment parameters employed in NextClade (https://clades.nextstrain.org). To this end, we analyzed all 1,409 South African genotypes. A total of 16 South African genotypes were filtered out owing to low coverage, whereas an additional 28 were removed owing to poor sequence quality. All the genomes in this dataset had a total coverage of more than 90%, with 70.4% of them (n = 959) having a coverage of more than 99%, and 94.1% (n = 1,283) of them having a coverage of more than 95% relative to the reference, whereas 53.1% (n = 726) of genomes had no missing nucleotides, giving a coverage of 100%. The final dataset of South African sequences (n = 1,365) were further annotated with additional metadata information (sampling locations, unique lab IDs and outbreak numbers) (Extended Data Fig. 10). The bulk of the South African sequences (~81%) were sampled within the province of KZN, with sampling from all of the 11 districts within the province.
Global reference dataset
South African sequences were analyzed against a backdrop of globally representative SARS-CoV-2 genotypes. At the time of sequence analysis, more than 90,000 SARS-CoV-2 genotypes have been publicly shared. Owing to the sheer size of this dataset and oversampling in specific countries (for example, England), we had to downsample this dataset to a manageable size. Important lineage-defining genotypes, along with ten randomly sampled genotypes per location, were included in the phylogenetic reconstruction. The final 5,848 references contained 889 other African genotypes, 1,209 genotypes from Asia, 2,775 genotypes from Europe, 434 and 367 genotypes from North and South America, respectively, and 174 genotypes from Oceania.
Phylogenetic analysis of SARS-CoV-2 in South Africa
South African genotypes were analyzed against the global reference dataset using a custom build of the SARS-CoV-2 NextStrain build (https://github.com/nextstrain/ncov). The pipeline contains several Python scripts that manage the analysis workflow. In short, it allows for the filtering of genotypes, the alignment of genotypes in MAFFT33, phylogenetic tree inference in IQ-Tree34, tree dating and ancestral state construction and annotation. The resulting time-scaled phylogeny can be viewed interactively and has been shared publicly on the NGS-SA NextStrain page (https://nextstrain.org/groups/ngs-sa/COVID19-Africa-2020.09.16).
The raw ML tree topology that was produced by the NextStrain build was used to estimate the number of viral introductions through time into South Africa. TreeTime35 was used to transform this ML tree topology into a dated tree topology using a constant rate of 8.0 × 10−4 nucleotide substitutions per site per year, after the exclusion of outlier sequences. A migration model was fitted on the resulting time-scaled tree topology in TreeTime, mapping country locations to tips and internal nodes. The resulting annotated tree topology was used to infer the number of viral introductions into South Africa through time.
Lineage and clade classification
We used the dynamic lineage classification method proposed by Rambaut et al.14 in this study via the Phylogenetic Assignment of Named Global Outbreak Lineages (PANGOLIN) software suite (https://github.com/hCoV-2019/pangolin). This is aimed at identifying the most epidemiologically important lineages of SARS-CoV-2 at the time of analysis, allowing researchers to monitor the epidemic in a particular geographical region. Accordingly, with this recently proposed dynamic lineage classification, many factors might suggest a new lineage, including: 1) monophyletic clusters on a global tree; 2) the presence of a statistically significant support (bootstrap/ultrafast bootstrap) on the node of the new lineages; 3) introduction into a novel geographic region; 4) epidemiological support (location and travel history); and 5) characteristic single-nucleotide polymorphisms. Accordingly, with those characteristics, three main SARS-CoV-2 lineages are currently recognized: lineage A, defined by Wuhan/WH04/2020; lineage B, defined by Wuhan-Hu-1 strain; and lineage C, a sub-classification from B lineage. We also classified the SARS-CoV-2 genomes in our dataset using the clade classification proposed by NextStrain, divided into 19A, 19B, 20A, 20B and 20C clades36,37.
Dated phylogenetics
To estimate time-calibrated phylogenies dated from time-stamped genome data, we conducted phylogenetic analysis, using the Bayesian software package BEAST v.1.10.4 (ref. 38), on four smaller subsets of data for each of the four lineages identified in the ML phylogeny and containing isolates from South Africa (Cluster B.1.1.54, n = 320; Cluster B.1.1.56, n = 104; Cluster C.1, n = 151; Cluster B.1.106, n = 68).
ML trees from these four data subsets were inspected in TempEst v1.5.3 for the presence of a temporal (that is, molecular clock) signal39. Linear regression of root-to-tip genetic distances against sampling dates indicated that the SARS-CoV-2 sequences evolve in a relatively strong clock-like manner (r = 9.45 × 10−2; r = 0.34; and r = 0.74 and r = 0.50 from subsets B.1.1.54; B.1.1.56; B.1.106; and C.1, respectively) (Extended Data Fig. 5).
For this analysis, we employed the strict molecular clock model, the HKY + I, the nucleotide substitution model and the exponential growth coalescent model40. We computed MCMC triplicate runs of 100 million states each, sampling every 10,000 steps for each dataset. Convergence of MCMC chains was checked using Tracer v.1.7.1 (ref. 41). Maximum clade credibility (MCC) trees were summarized from the MCMC samples using TreeAnnotator after discarding 10% as burn-in.
Phylogeographic analysis
To model phylogenetic diffusion of South African lineages across the country, we used a flexible relaxed random walk diffusion model that accommodates branch-specific variation in rates of dispersal with a Cauchy distribution42. For each sequence, latitude and longitude were attributed to a point randomly sampled within the patient’s province or district of residence. We discretized sequence sampling locations by considering five of nine provinces in South Africa, and all 11 districts in KZN, the most sampled province, where sequences belonging to the three clusters were sampled (as shown in Extended Data Fig. 3).
MCMC chains were run for more than 100 million generations and sampled every 10,000th step, with convergence assessed using Tracer v1.7 (ref. 43). MCC trees were summarized using TreeAnnotator after discarding 10% as burn-in. We used the R package ‘seraphim’44,45 to extract and map spatiotemporal information embedded in posterior trees.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
All the SARS-CoV-2 genomes generated and presented in this study are publicly accessible through the GISAID platform (https://www.gisaid.org/). The GISAID accession IDs of the South Africa sequences and reference genomes analyzed in this study are provided as part of Supplementary Table 3, which also contains the metadata for the sequences. Other raw data for this study are provided as the supplementary dataset: https://github.com/krisp-kwazulu-natal/SARSCoV2_South_Africa_major_lineages.git. The reference SARS-CoV-2 genome (MN908947.3) was downloaded from the National Center of Biotechnology Information database (https://www.ncbi.nlm.nih.gov/).
Code availability
R code and bash scripts to reproduce the analyses and figures presented in this paper are available at https://github.com/krisp-kwazulu-natal/SARSCoV2_South_Africa_major_lineages.git.
References
Hale, T., Webster, S., Petherick, A., Phillips, T. & Kira, B. Oxford COVID-19 Government Response Tracker. Blavatnik School of Government. https://www.bsg.ox.ac.uk/research/research-projects/coronavirus-government-response-tracker (2020).
Marivate, V. et al. Coronavirus disease (COVID-19) case data—South Africa. https://doi.org/10.5281/ZENODO.3819126.(2020)
Korber, B. et al. Tracking changes in SARS-CoV-2 spike: evidence that D614G increases infectivity of the COVID-19 virus. Cell 182, 812–829 (2020).
Lessells, R., Moosa, Y. & De Oliveira, T. Report into a nosocomial outbreak of coronavirus disease 2019 (COVID‐19) at Netcare St. Augustine’s Hospital https://www.krisp.org.za/manuscripts/StAugustinesHospitalOutbreakInvestigation_FinalReport_15may2020_comp.pdf (2020).
Tegally, H. et al. Emergence and rapid spread of a new severe acute respiratory syndrome-related coronavirus 2 (SARS-CoV-2) lineage with multiple spike mutations in South Africa. Preprint at https://www.medrxiv.org/content/10.1101/2020.12.21.20248640v1 (2020).
Andersen, K. G., Rambaut, A., Lipkin, W. I., Holmes, E. C. & Garry, R. F. The proximal origin of SARS-CoV-2. Nat. Med. 26, 450–452 (2020).
Wu, F. et al. A new coronavirus associated with human respiratory disease in China. Nature 579, 265–269 (2020).
Deng, X. et al. Genomic surveillance reveals multiple introductions of SARS-CoV-2 into Northern California. Science 369, 582–587 (2020).
Gonzalez-Reiche, A. S. et al. Introductions and early spread of SARS-CoV-2 in the New York City area. Science 369, 297–301 (2020).
Oude Munnink, B. B. et al. Rapid SARS-CoV-2 whole-genome sequencing and analysis for informed public health decision-making in the Netherlands. Nat. Med. 26, 1405–1410 (2020).
Eden, J.-S. et al. An emergent clade of SARS-CoV-2 linked to returned travellers from Iran. Virus Evol. 6, veaa027 (2020).
Gudbjartsson, D. F. et al. Spread of SARS-CoV-2 in the Icelandic population. N. Engl. J. Med. 382, 2302–2315 (2020).
Leung, K. et al. Territorywide study of early coronavirus disease outbreak, Hong Kong, China. Emerg. Infect. Dis. 27, 196–204 (2021)
Rambaut, A. et al. A dynamic nomenclature proposal for SARS-CoV-2 lineages to assist genomic epidemiology. Nat. Microbiol. 5, 1403–1407 (2020).
Giandhari, J. et al. Early transmission of SARS-CoV-2 in South Africa: an epidemiological and phylogenetic report. Int. J. Infect. Dis. 103, 234–241 (2021).
Disaster Management Act: Regulations to address, prevent and combat the spread of Coronavirus COVID-19: Amendment. South African Government. https://www.gov.za/documents/disaster-management-act-regulations-address-prevent-and-combat-spread-coronavirus-covid-19 (2020).
Huisman, J. S. et al. Estimation and worldwide monitoring of the effective reproductive number of SARS-CoV-2. Preprint at medRxiv https://doi.org/10.1101/2020.11.26.20239368 (2020).
Nordling, L. Study tells ‘remarkable story’ about COVID-19’s deadly rampage through a South African hospital. Science https://www.sciencemag.org/news/2020/05/study-tells-remarkable-story-about-covid-19-s-deadly-rampage-through-south-african (2020).
Shu, Y. & McCauley, J. GISAID: global initiative on sharing all influenza data–from vision to reality. Eurosurveillance 22, 30494 (2017).
KRISP severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) virus sequencing in South-Africa. https://www.ncbi.nlm.nih.gov/sra/SRX8454220 (2020).
SARS-CoV-2 lineages. https://cov-lineages.org/descriptions.html (2020)
Meredith, L. W. et al. Rapid implementation of SARS-CoV-2 sequencing to investigate cases of health-care associated COVID-19: a prospective genomic surveillance study. Lancet Infect. Dis. 20, 1263–1272 (2020).
du Plessis, L. et al. Establishment and lineage dynamics of the SARS-CoV-2 epidemic in the UK. Science https://doi.org/10.1126/science.abf2946 (2021).
Cluster outbreak at North West hospital: 106 patients and nurses infected with Covid-19. https://www.iol.co.za/news/south-africa/north-west/cluster-outbreak-at-north-west-hospital-106-patients-and-nurses-infected-with-covid-19-49616607 (2020).
Msomi, N. et al. A genomics network established to respond rapidly to public health threats in South Africa. Lancet Microbe 1, e229–e230 (2020).
Marivate, V. & Combrink, H. M. Use of available data to inform the COVID-19 outbreak in South Africa: a case study. Data Sci. J. https://datascience.codata.org/articles/10.5334/dsj-2020-019/ (2020).
Cori, A., Ferguson, N. M., Fraser, C. & Cauchemez, S. A new framework and software to estimate time-varying reproduction numbers during epidemics. Am. J. Epidemiol. 178, 1505–1512 (2013).
Gostic, K. M. et al. Practical considerations for measuring the effective reproductive number, Rt. PLoS Comput. Biol. 16, e1008409 (2020).
Goldstein, E. et al. Reconstructing influenza incidence by deconvolution of daily mortality time series. Proc. Natl Acad. Sci. USA 106, 21825–21829 (2009).
Quick, J. nCoV-2019 sequencing protocol v3 (LoCost) V.3. https://protocols.io/view/ncov-2019-sequencing-protocol-v3-locost-bh42j8ye (2020).
Vilsker, M. et al. Genome Detective: an automated system for virus identification from high-throughput sequencing data. Bioinformatics 35, 871–873 (2019)
Cleemput, S. et al. Genome Detective Coronavirus Typing Tool for rapid identification and characterization of novel coronavirus genomes. Bioinformatics 36, 3552–3555 (2020).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Nguyen, L.-T., Schmidt, H. A., Von Haeseler, A. & Minh, B. Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2015).
Sagulenko, P., Puller, V. & Neher, R. A. TreeTime: maximum-likelihood phylodynamic analysis. Virus Evol. 4, vex042 (2018).
Hadfield, J. et al. NextStrain: real-time tracking of pathogen evolution. Bioinformatics 34, 4121–4123 (2018).
Hodcroft, E. B., Hadfield, Neher, R. A. & Bedford, T. Year-letter genetic clade naming for SARS-CoV-2 on NextStrain.org. https://nextstrain.org/blog/2020-06-02-SARSCoV2-clade-naming (2020).
Suchard, M. A. et al. Bayesian phylogenetic and phylodynamic data integration using BEAST 1.10. Virus Evol. 4, vey016 (2018).
Rambaut, A., Lam, T. T., Max Carvalho, L. & Pybus, O. G. Exploring the temporal structure of heterochronous sequences using TempEst (formerly Path-O-Gen). Virus Evol. 2, vew007 (2016).
Griffiths, R. C. & Tavaré, S. Sampling theory for neutral alleles in a varying environment. Philos. Trans. R. Soc. Lond. Ser. B Biol. Sci. 344, 403–410 (1994).
Rambaut, A., Drummond, A. J., Xie, D., Baele, G. & Suchard, M. A. Posterior summarization in Bayesian phylogenetics using Tracer 1.7. Syst. Biol. 67, 901–904 (2018).
Lemey, P., Rambaut, A., Welch, J. J. & Suchard, M. A. Phylogeography takes a relaxed random walk in continuous space and time. Mol. Biol. Evol. 27, 1877–1885 (2010).
Ferreira, M. A. R. & Suchard, M. A. Bayesian analysis of elapsed times in continuous-time Markov chains. Can. J. Stat. 36, 355–368 (2008).
Dellicour, S. et al. Phylodynamic assessment of intervention strategies for the West African Ebola virus outbreak. Nat. Commun. 9, 2222 (2018).
Dellicour, S., Rose, R., Faria, N. R., Lemey, P. & Pybus, O. G. SERAPHIM: studying environmental rasters and phylogenetically informed movements. Bioinformatics 32, 3204–3206 (2016).
Acknowledgements
This research was funded by the South African Medical Research Council, MRC SHIP and the Department of Science and Innovation of South Africa. The KwaZulu-Natal Research Innovation and Sequencing Platform is funded by a core award of the South African Technology Innovation Agency. We thank A. Rambaut and Á. O’Toole for scientific discussion on how to include the South African lineages on PANGOLIN dynamic classification. We also thank all NGS-SA laboratories in South Africa that were responsible for producing the SARS-CoV-2 genomes that were the focus of the analysis in this paper. A full list of originating laboratories and authors is provided in Supplementary Table 2. Finally, we thank all other global laboratories for generating and making public the SARS-CoV-2 sequences (through GISAID) that were used as the reference dataset in this study. A complete list of individual contributors of sequences is provided in the Supplementary Information.
Author information
Authors and Affiliations
Contributions
J.G., S.P., S.E. and A.I. produced SARS-CoV-2 genomic data. N.M., K.M., N.H., D.Y., D.G., A.v.G., S.W., A.J.G., I.G., A.S., G.V.Z., W.P. and D.H. collected samples and curated the metadata. R.J.L. and T.d.O. participated in outbreak response. H.T., E.W., R.J.L., J.G., S.P., E.J.S., S.E., F.P., A.I., J.N.B., V.F., M.G., J.L., L.C.J.A. and T.d.O. analyzed the data. D.H., N.H, D.M., D.G., E.J.S., M.G., J.L., L.C.J.A. and T.d.O. helped with data interpretation. H.T., E.W., M.G., J.L., L.C.J.A. and T.d.O. wrote the initial manuscript. All authors participated in review of the manuscript at all stages.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Jennifer Sargent was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Map density representation of where the genomes in this study were sampled.
a, The number of genomes sampled in each province in South Africa (no genomes from Northern Cape – grey), b, The number of genomes sampled in each district of KZN, the most sampled province.
Extended Data Fig. 2 Classification of viruses circulating in South Africa.
a, Classification of South Africa genomes (n = 1365) per date into Pangolin lineages (SA-specific ones specified by red boxes), and into Nextstrain clades. b, Detailed sampling information for the four lineages cluster identified to be almost unique to South Africa.
Extended Data Fig. 3 Prevalence of main lineage clusters in South Africa.
a, Distribution of genomes belonging to the lineage clusters by province. b, Distribution of genomes belonging to the lineage clusters by district of KZN.
Extended Data Fig. 4 Ct score investigation of samples in this study.
a, Showing the average Ct scores at three target genes for genomes generated at KRISP, and classified into their respective Pangolin lineages. Each box is delimited by two lines at the 25th percentile and 75th percentile, with the line inside the box represents the median, and whisker lines drawn from the box to the whisker boundaries. b, Showing the decreasing trend in genome coverage as Ct score increases, with the bulk of genomes > 90% (n = 476) falling in the Ct < 30 category.
Extended Data Fig. 5 Temporal signaling for each cluster (Tempest).
For SARS-CoV-2, we accept temporal signaling with correlation coefficient > 0.2. Cluster B.1.1.54 (A) had a low correlation coefficient and was therefore rejected from further Bayesian spatiotemporal analyses. Regression lines are shown with error buffers (shaded area) representing 90% confidence intervals.
Extended Data Fig. 6 Maximum likelihood tree of a global dataset showing genomes coloured by sampling location in South Africa.
For genomes sampled in KZN, they are further specified by which district they were sampled from. A closer look into cluster B.1.106, C.1 and B.1.1.56 illustrated as trees from BEAST temporal analyses, with a defined time-scale. The zoom-in tree for B.1.1.54 was extracted as a subset of the big ML tree.
Extended Data Fig. 7 Flowchart of Pangolin lineage A and B dividing into sub-lineages with their lineage-defining mutations specified.
The main lineage-defining mutations of the main lineage cluster investigated in this study are shown in relation to their parent lineage (B.1.1).
Extended Data Fig. 8 Mutation frequencies in SA vs rest of the world for lineage-defining mutations.
Mutation predominantly seen in South Africa are shown in red, whereas the others are shown in blue.
Extended Data Fig. 9 Mapping of SARS-CoV-2 genome alignment generated in this study onto the genome structure to determine the location of gaps in the sequences.
a, An alignment of 436 medium quality genomes (<1000 missing bases) showing small amounts of recurrent gaps (white spaces) in ORF1b, S, ORF3a. b, An alignment of 203 low quality genomes (<2900 missing bases) showing a more important amounts of recurrent gaps (white) in ORF1a, ORF1b, S, ORF3a, and ORF7a genes. The rest of our genomes (N = 726) had 100% coverage relative to the reference.
Extended Data Fig. 10 Curation of South Africa dataset from all available South African genomes available on GISAID as at 15th September 2020.
We show initial number of genomes (n = 1409), how many were excluded at each cleaning step and the final number of genomes (n = 1365) with subdivisions into their originating province.
Supplementary information
Supplementary Information
Supplementary Table 1.
Supplementary Table 2
Acknowledgement of all laboratories in South Africa and worldwide responsible for producing the SARS-CoV-2 genomes analyzed in this paper
Supplementary Table 3
Metadata
Rights and permissions
About this article
Cite this article
Tegally, H., Wilkinson, E., Lessells, R.J. et al. Sixteen novel lineages of SARS-CoV-2 in South Africa. Nat Med 27, 440–446 (2021). https://doi.org/10.1038/s41591-021-01255-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-021-01255-3
This article is cited by
-
Humoral responses to wild type and ancient BA.1 SARS-CoV-2 variant after heterologous priming vaccination with ChAdOx1 nCoV-19 and BNT162b2 booster dose
Clinical and Experimental Medicine (2024)
-
Community-based management of a five-arm randomised clinical trial in COVID-19 outpatients in South Africa: challenges and opportunities
Trials (2023)
-
Rural populations facilitated early SARS-CoV-2 evolution and transmission in Missouri, USA
npj Viruses (2023)
-
Pharmacoinformatic approach to identify potential phytochemicals against SARS-CoV-2 spike receptor-binding domain in native and variants of concern
Molecular Diversity (2023)
-
Subregional origins of emerging SARS-CoV-2 variants during the second pandemic wave in Côte d’Ivoire
Virus Genes (2023)