Abstract
The involvement of host immunity in the gut microbiota-mediated colonization resistance to Clostridioides difficile infection (CDI) is incompletely understood. Here, we show that interleukin (IL)-22, induced by colonization of the gut microbiota, is crucial for the prevention of CDI in human microbiota-associated (HMA) mice. IL-22 signaling in HMA mice regulated host glycosylation, which enabled the growth of succinate-consuming bacteria Phascolarctobacterium spp. within the gut microbiome. Phascolarctobacterium reduced the availability of luminal succinate, a crucial metabolite for the growth of C. difficile, and therefore prevented the growth of C. difficile. IL-22-mediated host N-glycosylation is likely impaired in patients with ulcerative colitis (UC) and renders UC-HMA mice more susceptible to CDI. Transplantation of healthy human-derived microbiota or Phascolarctobacterium reduced luminal succinate levels and restored colonization resistance in UC-HMA mice. IL-22-mediated host glycosylation thus fosters the growth of commensal bacteria that compete with C. difficile for the nutritional niche.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Source data for all Figures and Extended Data Figures are provided with the paper. The microbiome data in this study are available at the NCBI Sequence Read Archive under BioProject PRJNA594915. The metabolome data are available at the NIH Common Fund’s Data Repository and Coordinating Center website (supported by NIH grant, U01-DK097430), the Metabolomics Workbench (http://www.metabolomicsworkbench.org) where it has been assigned project ID PR000882 (anti-IL-22 antibody experiment in Fig. 2b and Extended Data Fig. 4), PR000881 (FMT experiment in Fig. 6f and Extended Data Fig. 9) and PR000869 (Phascolarctobacterium administration in Fig. 6i).
References
Britton, R. A. & Young, V. B. Role of the intestinal microbiota in resistance to colonization by Clostridium difficile. Gastroenterology 146, 1547–1553 (2014).
Buffie, C. G. et al. Precision microbiome reconstitution restores bile acid mediated resistance to Clostridium difficile. Nature 517, 205–208 (2015).
Theriot, C. M. et al. Antibiotic-induced shifts in the mouse gut microbiome and metabolome increase susceptibility to Clostridium difficile infection. Nat. Commun. 5, 3114 (2014).
Kamada, N. & Nunez, G. Role of the gut microbiota in the development and function of lymphoid cells. J. Immunol. 190, 1389–1395 (2013).
Sonnenberg, G. F. et al. Innate lymphoid cells promote anatomical containment of lymphoid-resident commensal bacteria. Science 336, 1321–1325 (2012).
Zheng, Y. et al. Interleukin-22 mediates early host defense against attaching and effacing bacterial pathogens. Nat. Med. 14, 282–289 (2008).
Sakamoto, K. et al. IL-22 controls iron-dependent nutritional immunity against systemic bacterial infections. Sci. Immunol. 2, eaai8371 (2017).
Pham, T. A. et al. Epithelial IL-22RA1-mediated fucosylation promotes intestinal colonization resistance to an opportunistic pathogen. Cell Host Microbe 16, 504–516 (2014).
Pickard, J. M. et al. Rapid fucosylation of intestinal epithelium sustains host-commensal symbiosis in sickness. Nature 514, 638–641 (2014).
Goto, Y. et al. Innate lymphoid cells regulate intestinal epithelial cell glycosylation. Science 345, 1254009 (2014).
Satoh-Takayama, N. et al. Microbial flora drives interleukin 22 production in intestinal NKp46+ cells that provide innate mucosal immune defense. Immunity 29, 958–970 (2008).
Sanos, S. L. et al. RORγT and commensal microflora are required for the differentiation of mucosal interleukin 22-producing NKp46+ cells. Nat. Immunol. 10, 83–91 (2009).
Sadighi Akha, A. A. et al. Interleukin-22 and CD160 play additive roles in the host mucosal response to Clostridium difficile infection in mice. Immunology 144, 587–597 (2015).
Hasegawa, M. et al. Interleukin-22 regulates the complement system to promote resistance against pathobionts after pathogen-induced intestinal damage. Immunity 41, 620–632 (2014).
Abt, M. C. et al. Innate immune defenses mediated by two ILC subsets are critical for protection against acute Clostridium difficile infection. Cell Host Microbe 18, 27–37 (2015).
Nagao-Kitamoto, H. et al. Functional characterization of inflammatory bowel disease-associated gut dysbiosis in gnotobiotic mice. Cell Mol. Gastroenterol. Hepatol. 2, 468–481 (2016).
Collins, J., Auchtung, J. M., Schaefer, L., Eaton, K. A. & Britton, R. A. Humanized microbiota mice as a model of recurrent Clostridium difficile disease. Microbiome 3, 35 (2015).
Sonnenberg, G. F. & Artis, D. Innate lymphoid cell interactions with microbiota: implications for intestinal health and disease. Immunity 37, 601–610 (2012).
Watanabe, Y., Nagai, F. & Morotomi, M. Characterization of Phascolarctobacterium succinatutens sp. nov., an asaccharolytic, succinate-utilizing bacterium isolated from human feces. Appl. Environ. Microbiol. 78, 511–518 (2012).
Topping, D. L. & Clifton, P. M. Short-chain fatty acids and human colonic function: roles of resistant starch and nonstarch polysaccharides. Physiol. Rev. 81, 1031–1064 (2001).
Ferreyra, J. A. et al. Gut microbiota-produced succinate promotes C. difficile infection after antibiotic treatment or motility disturbance. Cell Host Microbe 16, 770–777 (2014).
Rodemann, J. F., Dubberke, E. R., Reske, K. A., Seo da, H. & Stone, C. D. Incidence of Clostridium difficile infection in inflammatory bowel disease. Clin. Gastroenterol. Hepatol. 5, 339–344 (2007).
Reddy, S. S. & Brandt, L. J. Clostridium difficile infection and inflammatory bowel disease. J. Clin. Gastroenterol. 47, 666–671 (2013).
Berg, A. M., Kelly, C. P. & Farraye, F. A. Clostridium difficile infection in the inflammatory bowel disease patient. Inflamm. Bowel Dis. 19, 194–204 (2013).
Vancamelbeke, M. et al. Genetic and transcriptomic bases of intestinal epithelial barrier dysfunction in inflammatory bowel disease. Inflamm. Bowel Dis. 23, 1718–1729 (2017).
Arijs, I. et al. Mucosal gene expression of antimicrobial peptides in inflammatory bowel disease before and after first infliximab treatment. PLoS One 4, e7984 (2009).
Arijs, I. et al. Effect of vedolizumab (anti-α4β7-integrin) therapy on histological healing and mucosal gene expression in patients with UC. Gut 67, 43–52 (2018).
Johnson, J. L., Jones, M. B., Ryan, S. O. & Cobb, B. A. The regulatory power of glycans and their binding partners in immunity. Trends Immunol. 34, 290–298 (2013).
Stanley, P. What have we learned from glycosyltransferase knockouts in mice? J. Mol. Biol. 428, 3166–3182 (2016).
Wu, F. et al. Phascolarctobacterium faecium abundant colonization in human gastrointestinal tract. Exp. Ther. Med. 14, 3122–3126 (2017).
Zhang, L. et al. Insight into alteration of gut microbiota in Clostridium difficile infection and asymptomatic C. difficile colonization. Anaerobe 34, 1–7 (2015).
Hourigan, S. K. et al. Microbiome changes associated with sustained eradication of Clostridium difficile after single faecal microbiota transplantation in children with and without inflammatory bowel disease. Aliment. Pharmacol. Ther. 42, 741–752 (2015).
Levy, M. et al. Microbiota-modulated metabolites shape the intestinal microenvironment by regulating NLRP6 inflammasome signaling. Cell 163, 1428–1443 (2015).
Battaglioli, E. J. et al. Clostridioides difficile uses amino acids associated with gut microbial dysbiosis in a subset of patients with diarrhea. Sci. Transl. Med. 10, eaam7019 (2018).
Morgan, X. C. et al. Dysfunction of the intestinal microbiome in inflammatory bowel disease and treatment. Genome Biol. 13, R79 (2012).
Wolk, K. et al. IL-22 induces lipopolysaccharide-binding protein in hepatocytes: a potential systemic role of IL-22 in Crohn’s disease. J. Immunol. 178, 5973–5981 (2007).
Schmechel, S. et al. Linking genetic susceptibility to Crohn’s disease with Th17 cell function: IL-22 serum levels are increased in Crohn’s disease and correlate with disease activity and IL23R genotype status. Inflamm. Bowel Dis. 14, 204–212 (2008).
Mann, E. R. et al. Human gut dendritic cells drive aberrant gut-specific T cell responses in ulcerative colitis, characterized by increased IL-4 production and loss of IL-22 and IFN-γ. Inflamm. Bowel Dis. 20, 2299–2307 (2014).
Leung, J. M. et al. IL-22-producing CD4+ cells are depleted in actively inflamed colitis tissue. Mucosal Immunol. 7, 124–133 (2014).
Lamas, B. et al. CARD9 impacts colitis by altering gut microbiota metabolism of tryptophan into aryl hydrocarbon receptor ligands. Nat. Med. 22, 598–605 (2016).
Martin, J. C. et al. IL-22BP is produced by eosinophils in human gut and blocks IL-22 protective actions during colitis. Mucosal Immunol. 9, 539–549 (2016).
Pelczar, P. et al. A pathogenic role for T cell-derived IL-22BP in inflammatory bowel disease. Science 354, 358–362 (2016).
Xu, A. T. et al. High suppressor of cytokine signaling-3 expression impairs STAT3-dependent protective effects of interleukin-22 in ulcerative colitis in remission. Inflamm. Bowel Dis. 21, 241–250 (2015).
Li, Y. et al. Increased suppressor of cytokine signaling-3 expression predicts mucosal relapse in ulcerative colitis. Inflamm. Bowel Dis. 19, 132–140 (2013).
Dias, A. M. et al. Dysregulation of T cell receptor N-glycosylation: a molecular mechanism involved in ulcerative colitis. Hum. Mol. Genet. 23, 2416–2427 (2014).
Lopez, J. & Grinspan, A. Fecal microbiota transplantation for inflammatory bowel disease. Gastroenterol. Hepatol. 12, 374–379 (2016).
Paramsothy, S. et al. Multidonor intensive faecal microbiota transplantation for active ulcerative colitis: a randomised placebo-controlled trial. Lancet 389, 1218–1228 (2017).
Atarashi, K. et al. Th17 cell induction by adhesion of microbes to intestinal epithelial cells. Cell 163, 367–380 (2015).
Paik, J. et al. Potential for using a hermetically-sealed, positive-pressured isocage system for studies involving germ-free mice outside a flexible-film isolator. Gut Microbes 6, 255–265 (2015).
Hecht, G. et al. A simple cage-autonomous method for the maintenance of the barrier status of germ-free mice during experimentation. Lab. Anim. 48, 292–297 (2014).
Schloss, P. D. et al. Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 75, 7537–7541 (2009).
Kozich, J. J., Westcott, S. L., Baxter, N. T., Highlander, S. K. & Schloss, P. D. Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl. Environ. Microbiol. 79, 5112–5120 (2013).
Segata, N. et al. Metagenomic biomarker discovery and explanation. Genome Biol. 12, R60 (2011).
Hirayama, A. et al. Metabolic profiling reveals new serum biomarkers for differentiating diabetic nephropathy. Anal. Bioanal. Chem. 404, 3101–3109 (2012).
Lorenz, M. A., Burant, C. F. & Kennedy, R. T. Reducing time and increasing sensitivity in sample preparation for adherent mammalian cell metabolomics. Anal. Chem. 83, 3406–3414 (2011).
Cohen, P. S. & Laux, D. C. Bacterial adhesion to and penetration of intestinal mucus in vitro. Methods Enzymol. 253, 309–314 (1995).
Schulz, B. L., Packer, N. H. & Karlsson, N. G. Small-scale analysis of O-linked oligosaccharides from glycoproteins and mucins separated by gel electrophoresis. Anal. Chem. 74, 6088–6097 (2002).
Acknowledgements
The authors thank A. B. Shreiner for technical assistance, the University of Michigan Center for Gastrointestinal Research (DK034933), Host Microbiome Initiative, the GF Animal Facility and the Michigan Regional Comprehensive Metabolomics Resource Core (MRC2) (DK097153) for core services and I. L. Bergin from the In Vivo Animal Core at the University of Michigan Unit for Laboratory Animal Medicine for histological assessment. This work was supported by the National Institutes of Health (NIH) DK110146 and DK108901 (N.K.), Crohn’s and Colitis Foundation of America (N.K and H.N.-K.), Young Investigator Grant from the Global Probiotics Council (N.K.), University of Michigan Center for Gastrointestinal Research (DK034933) (N.K.), Joint Usage/Research Program of Medical Mycology Research Center Chiba University 18-1 (N.K. and Y.G.), Japan Society for the Promotion of Science Postdoctoral Fellowship for Research Abroad (H.N.-K. and S.K.), the Uehara Memorial Foundation Postdoctoral Fellowship Award (S.K.), Clinical and Translational Science Awards Program (S.K.), Prevent Cancer Foundation (S.K.), Japan Society for the Promotion of Science KAKENHI grants 16H04901, 17H05654 and 18H04805 (S.F.), Japan Science and Technology Agency PRESTO grant JPMJPR1537 (S.F.), Japan Science and Technology Agency ERATO grant JPMJER1902 (S.F.), Advanced Research and Development Programs for Medical Innovation CREST program grant JP19gm1010009 (S.F.), the Takeda Science Foundation (S.F.) and the Food Science Institute Foundation (S.F.). MS analysis of glycans was performed by the Swedish Infrastructure for Biological Mass Spectrometry, supported by the Swedish Research Council.
Author information
Authors and Affiliations
Contributions
H.N.-K. and N.K. conceived and designed experiments. H.N.-K. conducted most of the experiments with help from J.L.L., S.K., P.K. and A.M.S. M.G.G. performed microbiome analysis. C.I., A.H. and S.F. performed metabolome analysis. K.A.E. helped with GF animal experiments. P.D.R.H. provided human stool samples. C.J., K.A.T and N.G.K. conducted glycan analysis. Y.G., R.R.J., V.B.Y., E.C.M. and J.Y.K. helped with critical advice and discussion. H.N.-K. and N.K. analyzed the data. H.N.-K. and N.K. wrote the manuscript with contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Saheli Sadanand was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Healthy human microbiotas prevent C. difficile infection.
a, (Left) GF B6 mice were colonized with healthy control (HC) microbiotas for 2 weeks (human microbiota-associated (HMA) mice). GF or HC-HMA mice (GF: n = 9, HC#1:n = 8 and HC#2:n = 3, biologically independent animals) were then infected with C. difficile VPI 10463 spores (103 spores/mouse). C. difficile load in feces was determined on indicated days post-infection. Dots represent individual mice. Bars indicate median. Data were pooled from 2 independent experiments. (Right) The mortality of C. difficile infected GF or HC-HMA mice. Statistical significance was assessed by Log-rank test (two-sided). (b and c) (b) HC-HMA mice were treated with a cocktail of antibiotics or regular water and then infected with C. difficile VPI 10463 spores (n = 5, biologically independent animals). (Left) CFU in feces. Dots represent individual mice. Bars indicate median. Data were pooled from 2 independent experiments. Statistical significance was assessed by 2-way ANOVA with Bonferroni post-hoc test (two-sided). (Right) The mortality of mice. Statistical significance was assessed by Log-rank test (two-sided). c, Representative histological images and associated histological scores. Scale bar is 100 µm. Data are presented as mean ± s.e.m. from pooled two independent experiments. Statistical significance was assessed by Mann-Whitney U test (two-sided).
Extended Data Fig. 2 IL-22 neutralization inhibits the expression of antimicrobial proteins.
Reg3b and Reg3g mRNA levels in control or αIL-22 antibody-treated HC-HMA Rag1−/− mice were determined by qPCR. Expression was normalized to that of the murine Actb gene. Dots represent individual mice. Data are presented as mean ± s.e.m. (n = 8, biologically independent animals) from pooled 2 independent experiments. Statistical significance was assessed by Mann-Whitney U test (two-sided).
Extended Data Fig. 3 IL-22 shapes the gut microbial community.
HC-HMA-Rag1−/− mice were treated with control or αIL-22 antibody twice, on day -5 and day -3, before fecal samples were collected. Bacterial 16 S rRNA sequences were analyzed. a, Shannon index (α-diversity) and number of OTUs (richness) of control and αIL-22 antibody-treated HMA-Rag1−/− mice. Dots represent individual mice. Data are presented as mean ± s.e.m. (n = 3, biologically independent animals). Data are representative of 2 independent experiments. Statistical significance was assessed by Mann-Whitney U test (two-sided). b, Microbial community structures were analyzed using the Yue and Clayton dissimilarity distance metric (θYC) and are shown in a nonmetric, multidimensional scaling plot. c, Bacterial taxonomy at the family level in the feces.
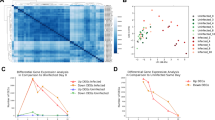
Extended Data Fig. 4 Luminal metabolomic analysis in IL-22–neutralized HMA mice.
a, Fecal samples were collected after treatment with αIL-22 antibody, twice and before CDI (day 0). A heat map showing the quantified metabolic profiles of control or αIL-22 antibody-treated HC-HMA-Rag1−/− mice. All concentrations of quantified metabolites were transformed into Z-scores and clustered according to their Euclidean distance. Gray areas in the heat map indicate that respective metabolites were not detected. b, Principal component analysis of the metabolome data. c, A loading scatter plot of the principal component analysis. d, Luminal metabolites were analyzed by CE-TOF/MS. Dots represent individual mice. Data are presented as mean ± s.e.m. (n = 13, biologically independent animals) from pooled 3 independent experiments. Statistical significance was assessed by Mann-Whitney U test (two-sided).
Extended Data Fig. 5 C. difficile growth on succinate.
In vitro growth of WT JIR8094 or Cd-CD2344- mutant C. difficile in a minimal medium supplemented with glucose or succinate. Data are presented as mean ± s.d. (n = 3 technical replicates). Data are representative of 3 independent experiments. Statistical significance was assessed by 2-way ANOVA with Bonferroni post-hoc test (two-sided).
Extended Data Fig. 6 Succinate is not required for the growth of C. difficile in germ-free mice.
GF C57BL/6 mice were infected with WT JIR8094 or Cd-CD2344− mutant C. difficile strains. C. difficile load in feces was determined on indicated days post-infection (n = 5, biologically independent animals). Dots represent individual mice. Bars indicate median. Data were from pooled 2 independent experiments. Statistical significance was assessed by 2-way ANOVA with Bonferroni post-hoc test (two-sided).
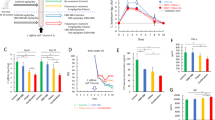
Extended Data Fig. 7 Gene expression profiles in UC patient cohorts.
a, The mRNA expression of IL22 mRNA in the colonic tissue from control subjects (n = 11), patients with inactive UC (n = 23) and patients with active UC (n = 74). Data were derived from GEO data set GSE75214. Dots represent biologically independent subjects. Data are presented as mean ± s.e.m.. Statistical significance was assessed by Kruskal-Wallis test with Dunn posttest (two-sided). (b and d) The mRNA expression of IL22, IL22RA, MGAT4A and MGAT4B mRNA in the colonic tissue from control subjects and patients with active UC. (c and e) Correlation between MGAT4A/MGAT4B and IL22RA mRNA expression in 3 groups. Statistical significance was measured by Pearson correlation test (two-sided). Dots represent biologically independent subjects. Data were derived from GEO data sets GSE16879 (control: n = 8 and active UC: n = 24) and GSE73661 (control: n = 12 and active UC: n = 67). Data are presented as mean ± s.e.m.. Statistical significance was assessed by Mann-Whitney U test (two-sided). (b and d). Statistical significance was measured by Pearson correlation test (two-sided) (c and e).
Extended Data Fig. 8 FMT restores the microbial composition in UC-HMA mice.
Significantly altered bacteria in pre–C. difficile infected UC-HMA mice with or without FMT were identified by LEfSe analysis. UC-HMA mice–enriched taxa have a positive LDA score (red, n = 6, biologically independent animals), and FMT-treated UC-HMA mice–enriched taxa have a negative LDA score (green, n = 6, biologically independent animals). A P value of < 0.05 and a score ≥ 2.0 were considered significant in Kruskal–Wallis and pairwise Wilcoxon tests (two-sided), respectively.
Extended Data Fig. 9 Luminal metabolomic analysis in FMT-treated UC-HMA mice.
a, Fecal samples were collected after treatment with FMT (day -3) and before CDI (day 0). A heat map showing the quantified metabolic profiles of UC-HMA or FMT-treated UC-HMA mice. All concentrations of quantified metabolites were transformed into Z-scores and clustered according to their Euclidean distance. Gray areas in the heat map indicate that respective metabolites were not detected. b, The principal component analysis of the metabolome data (n = 9, biologically independent animals). c, A loading scatter plot of the principal component analysis. d, Luminal metabolites were analyzed by CE-TOF/MS. Dots represent individual mice. Data are presented as mean ± s.e.m. (n = 9, biologically independent animals) from pooled 2 independent experiments. Statistical significance was assessed by Mann-Whitney U test (two-sided).
Extended Data Fig. 10 Phascolarctobacterium inoculation protect mice from CDI.
SPF C57BL/6 mice were treated with cefoperazone (0.5 mg/mL in drinking water). After 8 days, the mice were switched to regular water and allowed to recover for 2 days before being infected with C. difficile VPI spores. Mice were treated with P. faecium JCM 30894 and P. succinatutens JCM 16074 (106 CFU each strain) or with its culture medium by oral gavage, once, before C. difficile inoculation (1 day prior to CDI, pre-treatment) or 4 times post-inoculation (1, 3, 7, 10 days post-infection, post-treatment). Dots represent individual mice (n = 5, biologically independent animals). The mortality of C. difficile–infected mice was assessed. Statistical significance was assessed by Log-rank test (two-sided) from pooled 2 independent experiments.
Supplementary information
Source data
Source Data Fig. 1
Statistical Source Data to Fig. 1
Source Data Fig. 2
Statistical Source Data to Fig. 2
Source Data Fig. 3
Statistical Source Data to Fig. 3
Source Data Fig. 4
Statistical Source Data to Fig. 4
Source Data Fig. 5
Statistical Source Data to Fig. 5
Source Data Fig. 6
Statistical Source Data to Fig. 6
Source Data Extended Data Fig. 1
Statistical Source Data to Extended Data Fig. 1
Source Data Extended Data Fig. 2
Statistical Source Data to Extended Data Fig. 2
Source Data Extended Data Fig. 3
Statistical Source Data to Extended Data Fig. 3
Source Data Extended Data Fig. 4
Statistical Source Data to Extended Data Fig. 4
Source Data Extended Data Fig. 5
Statistical Source Data to Extended Data Fig. 5
Source Data Extended Data Fig. 6
Statistical Source Data to Extended Data Fig. 6
Source Data Extended Data Fig. 7
Statistical Source Data to Extended Data Fig. 7
Source Data Extended Data Fig. 9
Statistical Source Data to Extended Data Fig. 9
Source Data Extended Data Fig. 10
Statistical Source Data to Extended Data Fig. 10
Rights and permissions
About this article
Cite this article
Nagao-Kitamoto, H., Leslie, J.L., Kitamoto, S. et al. Interleukin-22-mediated host glycosylation prevents Clostridioides difficile infection by modulating the metabolic activity of the gut microbiota. Nat Med 26, 608–617 (2020). https://doi.org/10.1038/s41591-020-0764-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-020-0764-0
This article is cited by
-
Metabolic network of the gut microbiota in inflammatory bowel disease
Inflammation and Regeneration (2024)
-
A commensal protozoan attenuates Clostridioides difficile pathogenesis in mice via arginine-ornithine metabolism and host intestinal immune response
Nature Communications (2024)
-
A non-human primate model for human norovirus infection
Nature Microbiology (2024)
-
IL-22 alters gut microbiota composition and function to increase aryl hydrocarbon receptor activity in mice and humans
Microbiome (2023)
-
Microbiota-mediated colonization resistance: mechanisms and regulation
Nature Reviews Microbiology (2023)