Abstract
Genetically engineered T cell therapy can induce remarkable tumor responses in hematologic malignancies. However, it is not known if this type of therapy can be applied effectively to epithelial cancers, which account for 80–90% of human malignancies. We have conducted a first-in-human, phase 1 clinical trial of T cells engineered with a T cell receptor targeting HPV-16 E7 for the treatment of metastatic human papilloma virus-associated epithelial cancers (NCT02858310). The primary endpoint was maximum tolerated dose. Cell dose was not limited by toxicity with a maximum dose of 1 × 1011 engineered T cells administered. Tumor responses following treatment were evaluated using RECIST (Response Evaluation Criteria in Solid Tumors) guidelines. Robust tumor regression was observed with objective clinical responses in 6 of 12 patients, including 4 of 8 patients with anti-PD-1 refractory disease. Responses included extensive regression of bulky tumors and complete regression of most tumors in some patients. Genomic studies, which included intra-patient tumors with dichotomous treatment responses, revealed resistance mechanisms from defects in critical components of the antigen presentation and interferon response pathways. These findings demonstrate that engineered T cells can mediate regression of common carcinomas, and they reveal immune editing as a constraint on the curative potential of cellular therapy and possibly other immunotherapies in advanced epithelial cancer.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
Sequencing data files from WES and RNA have been deposited in the Database of Genotypes and Phenotypes (dbGaP) (accession no. phs002286.v1.p1). Gene sets for pathways analysis were from the Ingenuity Pathway Analysis database (version Summer Release 2018, https://digitalinsights.qiagen.com). WES mutation analysis and copy number variant tables are provided in Supplementary Datasets 1 and 2. Source data for Figs. 1 and 2, along with Extended Data Figs. 1–9, are provided in Supplementary Dataset 3. External requests for data will be evaluated by the corresponding author and requests may be subject to NIH policy. Source data are provided with this paper.
Code availability
RNA-seq expression data and WES data were processed with the CCR Collaborative Bioinformatics Resource (CCBR) in-house pipeline (https://github.com/CCBR/Pipeliner).
References
Robbins, P. F. et al. A pilot trial using lymphocytes genetically engineered with an NY-ESO-1-reactive T cell receptor: long-term follow-up and correlates with response. Clin. Cancer Res. 21, 1019–1027 (2015).
June, C. H., O’Connor, R. S., Kawalekar, O. U., Ghassemi, S. & Milone, M. C. CAR T cell immunotherapy for human cancer. Science 359, 1361–1365 (2018).
Guedan, S., Ruella, M. & June, C. H. Emerging cellular therapies for cancer. Annu. Rev. Immunol. 37, 145–171 (2019).
Hinrichs, C. S. Molecular pathways: breaking the epithelial cancer barrier for chimeric antigen receptor and T cell receptor gene therapy. Clin. Cancer Res. 22, 1559–1564 (2016).
Hinrichs, C. S. & Restifo, N. P. Reassessing target antigens for adoptive T cell therapy. Nat. Biotechnol. 31, 999–1008 (2013).
Stevanović, S. et al. Landscape of immunogenic tumor antigens in successful immunotherapy of virally induced epithelial cancer. Science 356, 200–205 (2017).
Doran, S. L. et al. T cell receptor gene therapy for human papillomavirus-associated epithelial cancers: a first-in-human, phase I/II study. J. Clin. Oncol. 37, 2759–2768 (2019).
Jin, B. Y. et al. Engineered T cells targeting E7 mediate regression of human papillomavirus cancers in a murine model. JCI Insight 3, e99488 (2018).
Schwartzentruber, D. J. Guidelines for the safe administration of high-dose interleukin-2. J. Immunother. 24, 287–293 (2001).
Center for Biologics Evaluation and Research. YESCARTA (axicabtagene ciloleucel) (FDA, 2019).
Center for Biologics Evaluation and Research. KYMRIAH (tisagenlecleucel) (FDA, 2019).
Spies, T. et al. A gene in the human major histocompatibility complex class II region controlling the class I antigen presentation pathway. Nature 348, 744–747 (1990).
Cerundolo, V. et al. Presentation of viral antigen controlled by a gene in the major histocompatibility complex. Nature 345, 449–452 (1990).
Farrar, M. A. & Schreiber, R. D. The molecular cell biology of interferon-gamma and its receptor. Annu. Rev. Immunol. 11, 571–611 (1993).
Schneider, W. M., Chevillotte, M. D. & Rice, C. M. Interferon-stimulated genes: a complex web of host defenses. Annu Rev. Immunol. 32, 513–545 (2014).
Tran, E., Longo, D. L. & Urba, W. J. A milestone for CAR T cells. New Engl. J. Med. 377, 2593–2596 (2017).
Ribas, A. & Wolchok, J. D. Cancer immunotherapy using checkpoint blockade. Science 359, 1350–1355 (2018).
Gettinger, S. et al. Impaired HLA class I antigen processing and presentation as a mechanism of acquired resistance to immune checkpoint inhibitors in lung cancer. Cancer Discov. 7, 1420–1435 (2017).
Zaretsky, J. M. et al. Mutations associated with acquired resistance to PD-1 blockade in melanoma. New Engl. J. Med. 375, 819–829 (2016).
Shin, D. S. et al. Primary resistance to PD-1 blockade mediated by JAK1/2 mutations. Cancer Discov. 7, 188–201 (2017).
Schreiber, R. D., Old, L. J. & Smyth, M. J. Cancer immunoediting: integrating immunity’s roles in cancer suppression and promotion. Science 331, 1565–1570 (2011).
Dunn, G. P., Koebel, C. M. & Schreiber, R. D. Interferons, immunity and cancer immunoediting. Nat. Rev. Immunol. 6, 836–848 (2006).
Tran, E., Robbins, P. F. & Rosenberg, S. A. ‘Final common pathway’ of human cancer immunotherapy: targeting random somatic mutations. Nat. Immunol. 18, 255–262 (2017).
Scheper, W. et al. Low and variable tumor reactivity of the intratumoral TCR repertoire in human cancers. Nat. Med. 25, 89–94 (2019).
Schumacher, T. N., Scheper, W. & Kvistborg, P. Cancer neoantigens. Annu. Rev. Immunol. 37, 173–200 (2019).
Huber, M. H. et al. A phase II study of ifosfamide in recurrent squamous cell carcinoma of the head and neck. Am. J. Clin. Oncol. 19, 379–382 (1996).
Sandler, A. et al. Ifosfamide in the treatment of advanced or recurrent squamous cell carcinoma of the head and neck: a phase II Hoosier Oncology Group trial. Am. J. Clin. Oncol. 21, 195–197 (1998).
Sutton, G. P., Blessing, J. A., Photopulos, G., Berman, M. L. & Homesley, H. D. Gynecologic Oncology Group experience with ifosfamide. Semin. Oncol. 17, 6–10 (1990).
Riemer, A. B. et al. A conserved E7-derived cytotoxic T lymphocyte epitope expressed on human papillomavirus 16-transformed HLA-A2+ epithelial cancers. J. Biol. Chem. 285, 29608–29622 (2010).
Zhang, L. et al. Enhanced efficacy and limited systemic cytokine exposure with membrane-anchored interleukin-12 T cell therapy in murine tumor models. J. Immunother. Cancer 8, e000210 (2020).
Helman, S. R. et al. Human papillomavirus T cell cross-reactivity in cervical cancer: implications for immunotherapy clinical trial design. JAMA Netw. Open 1, e180706 (2018).
Jin, J. et al. Enhanced clinical-scale manufacturing of TCR transduced T cells using closed culture system modules. J. Transl. Med. 16, 13 (2018).
Khong, H. T. & Rosenberg, S. A. Pre-existing immunity to tyrosinase-related protein (TRP)-2, a new TRP-2 isoform, and the NY-ESO-1 melanoma antigen in a patient with a dramatic response to immunotherapy. J. Immunol. 168, 951–956 (2002).
Cormier, J. N. et al. Natural variation of the expression of HLA and endogenous antigen modulates CTL recognition in an in vitro melanoma model. Int. J. Cancer 80, 781–790 (1999).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at https://arxiv.org/abs/1303.3997v2 (2013).
Acknowledgements
This research was funded by the NIH Intramural Research Program and through a NIH Cooperative Research and Development Agreement with Kite, a Gilead Company (C.S.H.). Support was also provided by the NHLBI-funded National Gene Vector Biorepository at Indiana University under contract no. 75N92019D00018 and by federal funding through the NCI, NIH, under contract no. 75N91019D00024 (C.S.H.). The content of this publication does not necessarily reflect the views or policies of the Department of Health and Human Services, nor does mention of trade names, commercial products or organizations imply endorsement by the US Government. The clinical-grade E7 TCR retroviral vector was manufactured by S. Feldman, NCI Surgery Branch.
Author information
Authors and Affiliations
Contributions
N.B.N., S.M.N. and C.S.H. conceived the research, interpreted the data and co-wrote the manuscript. N.B.N., A.L.S., S.S., T.J.M., J.B.L., A.C.W., C.S., S.L.D., S.K., S.S., S.H.A. and A.B. performed experiments and/or analyzed the data. C.L.T. and C.S.H. were involved in preclinical studies. S.M.N., J.A.K., M.H.B., E.F., W.C.F., D.S., N.G., S.H. and C.S.H. were involved in the clinical, nursing, referral and/or clinical manufacturing team. All authors proofread the final manuscript.
Corresponding author
Ethics declarations
Competing interests
C.S.H. is an inventor on the NIH patent for the E7 TCR and other NIH patents in the field of immunotherapy. C.S.H. receives research funding through an NCI Collaborative Research and Development Agreement with Kite Pharma. The other authors declare no competing interests.
Additional information
Peer review information Saheli Sadanand was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1
CONSORT flow diagram.
Extended Data Fig. 2 Durable, complete regression of multiple index and non-index tumors.
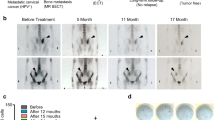
Radiographic imaging studies from Patient 5 and Patient 12. a, For Patient 5, computed tomography (CT) scans are shown. b, For Patient 12, the top row are CT scans and the middle and bottom rows are positron emission tomography-CT scans. The timepoint for the scans is indicated above each column. Yellow arrows indicate tumors. c, The number of discrete tumors present at baseline and serial timepoints after treatment are shown.
Extended Data Fig. 3 Peripheral blood engraftment of E7 TCR-T cells in Patient 12 at late time points.
The frequency of E7 TCR-T cells in the peripheral blood of Patient 12 at late time points after treatment is graphed.
Extended Data Fig. 4 Peripheral blood engraftment of E7 TCR-T cells.
Flow cytometry was used to determine engraftment of E7 TCR-T cells in the peripheral blood of patients at the first response assessment timepoint (6 weeks). a, The frequency of E7 TCR-T cells in peripheral blood correlated with dose (P = 0.0006, Kruskal-Wallis one-way ANOVA) but not with response (P = 0.347, two-sided unpaired t-test). b, The concentration of E7 TCR-T cells in peripheral blood correlated with dose (P = 0.0226, one-way ANOVA) but not with response (P = 0.4796, two-sided unpaired t-test).
Extended Data Fig. 5 Serum antibodies against E7 TCR-T cells.
The dotted line indicates the background detection level. Hamster anti-mouse TCR antibody was used as a positive control (Pos). Healthy donor serum was used as the negative control (Neg).
Extended Data Fig. 6 Infusion product phenotypic characterization.
The expression of cell surface antigens by E7 TCR-T cells was determined by flow cytometry. a, Stacked bar graph of the frequency of single-positive CD4 and CD8 T cells. b, Pie chart that depict memory T cell subset composition. Tn (naïve, CD45RA+CCR7+), Tcm (central memory, CD45RA−CCR7+), Tem (effector memory, CD45RA−CCR7−), Temra (effector memory RA+, CD45RA+CCR7−). c, Bar graph of the frequency of expression of the chemokine receptors in the symbol legend. d, The frequency of each marker or subset in the E7 TCR-T cells administered to patients who responded (R) or did not respond (NR) to treatment. P-values are two-sided unpaired t-tests.
Extended Data Fig. 7 Phenotype of engrafted peripheral blood E7 TCR-T cells.
a, Flow cytometric analysis of memory T cell subsets, chemokine receptors, CD4/CD8 T cell subsets, and Ki-67 expression by engrafted E7 TCR-T cells is shown. The T cell subset or phenotypic marker is indicated above each graph. Gating is on CD3+, mTCRB+, live, lymphocytes. CD4 and CD8 frequencies are for single positive cells. Tn (naïve, CD45RA+CCR7+), Tcm (central memory, CD45RA−CCR7+), Tem (effector memory, CD45RA−CCR7−), Temra (effector memory RA+, CD45RA+CCR7−). b, Phenotype at first response assessment (6 weeks) of engrafted E7 TCR-T cells in patients who responded (R) or did not respond (NR) to treatment. The T cell subset or phenotypic marker is indicated above each graph. Red symbols represent patients with tumor responses. P-values are from two-sided unpaired t-tests.
Extended Data Fig. 8 Infusion product functional characterization.
a, Percent cell killing at 8 hours as measured by impedance-based cytolysis assay. The effector to target ratio (E:T) is indicated on the x-axis. The target cell is indicated in the symbol legend. Error bars represent the standard deviation of 2–5 technical replicates. CaSki is HLA-A*02:01+ HPV-16+. 624 is HLA-A*02:01+ HPV-16-. b, The frequency of transduced cells that expressed Ki-67 as determined by flow cytometry. c, Cytokine production following coculture of the infusion product from the patient indicated in the graph title with 293-A2 cells pulsed with the target peptide indicated in the symbol legend. Error bars represent the standard deviation of 2-3 independent experiments (with 2 technical replicates in each experiment); all replicates are shown. d, ELISPOT assay measurement of cytokine-secreting cells in the infusion product. The cytokine measured is indicated by the graph title. The target cells and symbol legend are the same as in panel C. 2 independent experiments are shown. e-h, Functional characteristics of the infusion products administered to patients who responded (R) or did not respond (NR) to treatment. P-values are two-sided unpaired t-tests.
Extended Data Fig. 9 Serum cytokines and chemokines following E7 TCR-T cell infusion.
Serum concentrations of IL-1α, IL-1β, IL-4, IL-5, IL-6, IL-7, IL-8, IL-10, IL-12/23 p40, IL-12 p70, IL-13, IL-15, IL-16, IL-17A, IL-21, IL-22, IL-23, IL-27, IL-31, IFNγ, TNFα, TNFβ, GMCSF, VEGF-A, Eotaxin, Eotaxin-3, CXCL10, MCP-1, MCP-4, MIP-1α, MIP-1β, TARC, and MIP-3α were determined. Chemokines and cytokines for which the peak values were significantly different between responding and non-responding patients are shown. a, Graph of serum IL-17 and VEGF-A concentrations in patients indicated in symbol legend at the time points indicated on the x-axis. Red color indicates patients with tumor responses. b, Peak IL-17 and VEGF-A serum concentrations in patients who responded (R) or who did not respond (NR) to treatment. The symbol legend is shown in panel A. Statistical significance was determined by a two-sided unpaired t-test.
Extended Data Fig. 10 Infiltration of tumor biopsy specimens with E7 TCR-T cells.
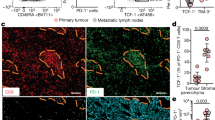
Serial samples from endoscopic biopsy of Patient 12’s tumor were examined. Immunohistochemistry was performed to detect CD3 protein, and RNAscope was performed to detect the E7 TCR α-chain (mTRAC) and HPV-16 E7 transcripts. A sequential labeling technique was employed to simultaneously examine protein and RNA expression on a single slide (each sample was stained once and is shown). The time point for each biopsy is indicated at the top of each row. The protein or transcript labeled is indicated to the left of each column. The color legend indicates labeling for the overlays on the bottom two rows. The yellow arrows point to E7 TCR-T cells as detected by CD3 protein membrane labeling and cytoplasmic mTRAC transcript labeling.
Supplementary information
Supplementary Information
Supplementary Fig. 1, Tables 1–3, and clinical trial protocol.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Rights and permissions
About this article
Cite this article
Nagarsheth, N.B., Norberg, S.M., Sinkoe, A.L. et al. TCR-engineered T cells targeting E7 for patients with metastatic HPV-associated epithelial cancers. Nat Med 27, 419–425 (2021). https://doi.org/10.1038/s41591-020-01225-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-020-01225-1
This article is cited by
-
Novel insights into TCR-T cell therapy in solid neoplasms: optimizing adoptive immunotherapy
Experimental Hematology & Oncology (2024)
-
HLA-class II restricted TCR targeting human papillomavirus type 18 E7 induces solid tumor remission in mice
Nature Communications (2024)
-
Digital twins elucidate critical role of Tscm in clinical persistence of TCR-engineered cell therapy
npj Systems Biology and Applications (2024)
-
Activating STING/TBK1 suppresses tumor growth via degrading HPV16/18 E7 oncoproteins in cervical cancer
Cell Death & Differentiation (2024)
-
CAR T cell therapy for patients with solid tumours: key lessons to learn and unlearn
Nature Reviews Clinical Oncology (2024)