Abstract
Developmental processes underlying normal tissue regeneration have been implicated in cancer, but the degree of their enactment during tumor progression and under the selective pressures of immune surveillance, remain unknown. Here we show that human primary lung adenocarcinomas are characterized by the emergence of regenerative cell types, typically seen in response to lung injury, and by striking infidelity among transcription factors specifying most alveolar and bronchial epithelial lineages. In contrast, metastases are enriched for key endoderm and lung-specifying transcription factors, SOX2 and SOX9, and recapitulate more primitive transcriptional programs spanning stem-like to regenerative pulmonary epithelial progenitor states. This developmental continuum mirrors the progressive stages of spontaneous outbreak from metastatic dormancy in a mouse model and exhibits SOX9-dependent resistance to natural killer cells. Loss of developmental stage-specific constraint in macrometastases triggered by natural killer cell depletion suggests a dynamic interplay between developmental plasticity and immune-mediated pruning during metastasis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Raw data from western blots is included in Source Data. A ranked list of DEGs and complete GSEA results for all PhenoGraph clusters analyzed in this manuscript and provided in Supplementary Tables 4–15. A custom GSEA annotation file, assembled to query cell types and pathways related lung epithelial development and regeneration is provided in the Supplementary Dataset. All raw and processed scRNA-seq data with cell type annotations was deposited in NCBI’s Gene Expression Omnibus and are accessible through accession number GSE123904. Fully annotated count matrices are available for download at (https://s3.amazonaws.com/dp-lab-data-public/lung-development-cancer-progression/PATIENT_LUNG_ADENOCARCINOMA_ANNOTATED.h5 and https://s3.amazonaws.com/dp-lab-data-public/lung-development-cancer-progression/MOUSE_LUNG_ADENOCARCINOMA_METASTASIS_ANNOTATED.h5). All other datasets generated and analyzed in the current study are available from the corresponding authors upon request.
Code availability
All custom code, statistical analysis and visualizations were performed in Python and are demonstrated in a Jupyter notebook available for download at https://github.com/dpeerlab/lung-development-cancer-progression. The following open-source algorithms were additionally used as described in the methods: SEQC (https://github.com/ambrosejcarr/seqc), t-SNE (https://lvdmaaten.github.io/software/), MAGIC (https://github.com/dpeerlab/magic) and PhenoGraph (https://github.com/jacoblevine/PhenoGraph).
References
Beumer, J. & Clevers, H. Regulation and plasticity of intestinal stem cells during homeostasis and regeneration. Development 143, 3639–3649 (2016).
Kumar, P. A. et al. Distal airway stem cells yield alveoli in vitro and during lung regeneration following H1N1 influenza infection. Cell 147, 525–538 (2011).
Vaughan, A. E. et al. Lineage-negative progenitors mobilize to regenerate lung epithelium after major injury. Nature 517, 621–625 (2015).
Zuo, W. et al. p63(+)Krt5(+) distal airway stem cells are essential for lung regeneration. Nature 517, 616–620 (2015).
Zacharias, W. J. et al. Regeneration of the lung alveolus by an evolutionarily conserved epithelial progenitor. Nature 555, 251–255 (2018).
Zaret, K. S. & Grompe, M. Generation and regeneration of cells of the liver and pancreas. Science 322, 1490–1494 (2008).
Kotton, D. N. & Morrisey, E. E. Lung regeneration: mechanisms, applications and emerging stem cell populations. Nat. Med. 20, 822–832 (2014).
Murry, C. E. & Keller, G. Differentiation of embryonic stem cells to clinically relevant populations: lessons from embryonic development. Cell 132, 661–680 (2008).
Tata, P. R. et al. Developmental history provides a roadmap for the emergence of tumor plasticity. Dev. Cell. 44, 679–693 (2018).
Massagué, J. & Obenauf, A. C. Metastatic colonization by circulating tumour cells. Nature 529, 298–306 (2016).
Klein, A. M. et al. Droplet barcoding for single-cell transcriptomics applied to embryonic stem cells. Cell 161, 1187–1201 (2015).
Zilionis, R. et al. Single-cell barcoding and sequencing using droplet microfluidics. Nat. Protoc. 12, 44–73 (2017).
Azizi, E. et al. Single-cell map of diverse immune phenotypes in the breast tumor microenvironment. Cell 174, 1293–1308 (2018).
van Dijk, D. et al. Recovering gene interactions from single-cell data using data diffusion. Cell 174, 716–729 (2018).
Levine, J. H. et al. Data-driven phenotypic dissection of AML reveals progenitor-like cells that correlate with prognosis. Cell 162, 184–197 (2015).
Malladi, S. et al. Metastatic latency and immune evasion through autocrine inhibition of WNT. Cell 165, 45–60 (2016).
Tirosh, I. et al. Single-cell RNA-seq supports a developmental hierarchy in human oligodendroglioma. Nature 539, 309–313 (2016).
Lavin, Y. et al. Innate immune landscape in early lung adenocarcinoma by paired single-cell analyses. Cell 169, 750–765 (2017).
Shen, R. & Seshan, V. E. FACETS: allele-specific copy number and clonal heterogeneity analysis tool for high-throughput DNA sequencing. Nucleic Acids Res. 44, e131 (2016).
Cheng, D. T. et al. Memorial sloan kettering-integrated mutation profiling of actionable cancer targets (MSK-IMPACT): a hybridization capture-based next-generation sequencing clinical assay for solid tumor molecular oncology. J. Mol. Diagn. 17, 251–264 (2015).
Bremnes, R. M. et al. The role of tumor-infiltrating lymphocytes in development, progression, and prognosis of non-small cell lung cancer. J. Thorac. Oncol. 11, 789–800 (2016).
Morrisey, E. E. & Hogan, B. L. Preparing for the first breath: genetic and cellular mechanisms in lung development. Dev. Cell 18, 8–23 (2010).
Du, Y. N. et al. Lung gene expression analysis (LGEA): an integrative web portal for comprehensive gene expression data analysis in lung development. Thorax 72, 481–484 (2017).
Du, Y. N., Guo, M. Z., Whitsett, J. A. & Xu, Y. ‘LungGENS’: a web-based tool for mapping single-cell gene expression in the developing lung. Thorax 70, 1092–1094 (2015).
Treutlein, B. et al. Reconstructing lineage hierarchies of the distal lung epithelium using single-cell RNA-seq. Nature 509, 371–375 (2014).
Guha, A. et al. Neuroepithelial body microenvironment is a niche for a distinct subset of Clara-like precursors in the developing airways. Proc. Natl Acad. Sci. USA 109, 12592–12597 (2012).
Haghverdi, L., Buttner, M., Wolf, F. A., Buettner, F. & Theis, F. J. Diffusion pseudotime robustly reconstructs lineage branching. Nat. Methods 13, 845–848 (2016).
Setty, M. et al. Wishbone identifies bifurcating developmental trajectories from single-cell data. Nat. Biotechnol. 34, 637–645 (2016).
Jobe, A. H., Whitsett, J. & Abman, S. H. (eds) Fetal and Neonatal Lung Development: Clinical Correlates and Technologies for the Future (Cambridge University Press, 2016).
Nakamura, N. et al. Identification of tumor markers and differentiation markers for molecular diagnosis of lung adenocarcinoma. Oncogene 25, 4245–4255 (2006).
Smith, B. A. et al. A human adult stem cell signature marks aggressive variants across epithelial cancers. Cell. Rep. 24, 3353–3366 (2018).
Niakan, K. K. et al. Sox17 promotes differentiation in mouse embryonic stem cells by directly regulating extraembryonic gene expression and indirectly antagonizing self-renewal. Genes Dev. 24, 312–326 (2010).
Seguin, C. A., Draper, J. S., Nagy, A. & Rossant, J. Establishment of endoderm progenitors by SOX transcription factor expression in human embryonic stem cells. Cell Stem Cell 3, 182–195 (2008).
Okubo, T., Knoepfler, P. S., Eisenman, R. N. & Hogan, B. L. N-myc plays an essential role during lung development as a dosage-sensitive regulator of progenitor cell proliferation and differentiation. Development 132, 1363–1374 (2005).
Rawlins, E. L., Clark, C. P., Xue, Y. & Hogan, B. L. The Id2+ distal tip lung epithelium contains individual multipotent embryonic progenitor cells. Development 136, 3741–3745 (2009).
Gyorffy, B., Surowiak, P., Budczies, J. & Lanczky, A. Online survival analysis software to assess the prognostic value of biomarkers using transcriptomic data in non-small-cell lung cancer. PLoS ONE 8, e82241 (2013).
Cancer Genome Atlas Research Network. Comprehensive molecular profiling of lung adenocarcinoma. Nature 511, 543–550 (2014).
Winslow, M. M. et al. Suppression of lung adenocarcinoma progression by Nkx2-1. Nature 473, 101–104 (2011).
Jung, H., Hsiung, B., Pestal, K., Procyk, E. & Raulet, D. H. RAE-1 ligands for the NKG2D receptor are regulated by E2F transcription factors, which control cell cycle entry. J. Exp. Med. 209, 2409–2422 (2012).
Long, E. O. Negative signaling by inhibitory receptors: the NK cell paradigm. Immunol. Rev. 224, 70–84 (2008).
Er, E. E. et al. Pericyte-like spreading by disseminated cancer cells activates YAP and MRTF for metastatic colonization. Nat. Cell Biol. 20, 966–978 (2018).
Weber, K. et al. RGB marking facilitates multicolor clonal cell tracking. Nat. Med. 17, 504–509 (2011).
Finak, G. et al. MAST: a flexible statistical framework for assessing transcriptional changes and characterizing heterogeneity in single-cell RNA sequencing data. Genome Biol. 16, 278 (2015).
Nowotschin, S. et al. The emergent landscape of the mouse gut endoderm at single-cell resolution. Nature 569, 361 (2019).
Mathelier, A. et al. JASPAR 2014: an extensively expanded and updated open-access database of transcription factor binding profiles. Nucleic Acids Res. 42, D142–D147 (2014).
Mathelier, A. et al. JASPAR 2016: a major expansion and update of the open-access database of transcription factor binding profiles. Nucleic Acids Res. 44, D110–D115 (2016).
Novershtern, N. et al. Densely interconnected transcriptional circuits control cell states in human hematopoiesis. Cell 144, 296–309 (2011).
Jeffrey, K. L. et al. Positive regulation of immune cell function and inflammatory responses by phosphatase PAC-1. Nat. Immunol. 7, 274–283 (2006).
Halko, N. M. P. & Tropp, J. A. Finding structure with randomness: probabilistic algorithms for constructing apprxoimate matrix decompositions. SIAM Rev. 53, 217–288 (2011).
Valle, S., Li, W. H. & Qin, S. J. Selection of the number of principal components: the variance of the reconstruction error criterion with a comparison to other methods. Ind. Eng. Chem. Res. 38, 4389–4401 (1999).
van der Maaten, L. & Hinton, G. Visualizing data using t-SNE. J. Mach. Learn. Res. 9, 2579–2605 (2008).
Jacomy, M., Venturini, T., Heymann, S. & Bastian, M. ForceAtlas2, a continuous graph layout algorithm for handy network visualization designed for the Gephi software. PLoS ONE 9, e98679 (2014).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740 (2011).
Rock, J. R. et al. Basal cells as stem cells of the mouse trachea and human airway epithelium. Proc. Natl Acad. Sci. USA 106, 12771–12775 (2009).
Li, S. et al. Molecular signatures of antibody responses derived from a systems biology study of five human vaccines. Nat. Immunol. 15, 195–204 (2014).
Abbas, A. R. et al. Immune response in silico (IRIS): immune-specific genes identified from a compendium of microarray expression data. Genes Immun. 6, 319–331 (2005).
Mor-Vaknin, N., Punturieri, A., Sitwala, K. & Markovitz, D. M. Vimentin is secreted by activated macrophages. Nat. Cell Biol. 5, 59–63 (2003).
Zepp, J. A. et al. Distinct mesenchymal lineages and niches promote epithelial self-renewal and myofibrogenesis in the lung. Cell 170, 1134–1148 e1110 (2017).
Lee, J. H. et al. Anatomically and functionally distinct lung mesenchymal populations marked by Lgr5 and Lgr6. Cell 170, 1149–1163 (2017).
Xia, H. et al. Calcium-binding protein S100A4 confers mesenchymal progenitor cell fibrogenicity in idiopathic pulmonary fibrosis. J. Clin. Invest. 127, 2586–2597 (2017).
Degryse, A. L. et al. Repetitive intratracheal bleomycin models several features of idiopathic pulmonary fibrosis. Am. J. Physiol. Lung Cell Mol. Physiol. 299, L442–L452 (2010).
Tanjore, H. et al. Contribution of epithelial-derived fibroblasts to bleomycin-induced lung fibrosis. Am. J. Respir. Crit. Care Med. 180, 657–665 (2009).
Lawson, W. E. et al. Characterization of fibroblast-specific protein 1 in pulmonary fibrosis. Am. J. Respir. Crit. Care Med. 171, 899–907 (2005).
Li, Z. H., Dulyaninova, N. G., House, R. P., Almo, S. C. & Bresnick, A. R. S100A4 regulates macrophage chemotaxis. Mol. Biol. Cell 21, 2598–2610 (2010).
Moore, K. W., Malefyt, deWaal, Coffman, R. & O’Garra, R. L. A. Interleukin-10 and the interleukin-10 receptor. Annu. Rev. Immunol. 19, 683–765 (2001).
Priceman, S. J. et al. Targeting distinct tumor-infiltrating myeloid cells by inhibiting CSF-1 receptor: combating tumor evasion of antiangiogenic therapy. Blood 115, 1461–1471 (2010).
Lambrechts, D. et al. Phenotype molding of stromal cells in the lung tumor microenvironment. Nat. Med. 24, 1277–1289 (2018).
Zilionis, R. et al. Single-cell transcriptomics of human and mouse lung cancers reveals conserved myeloid populations across individuals and species. Immunity 50, 1317–1334 (2019).
Coifman, R. R. et al. Geometric diffusions as a tool for harmonic analysis and structure definition of data: diffusion maps. Proc. Natl Acad. Sci. USA 102, 7426–7431 (2005).
Setty, M. et al. Characterization of cell fate probabilities in single-cell data with Palantir. Nat. Biotechnol. 37, 451–460 (2019).
Davoli, T., Uno, H., Wooten, E. C. & Elledge, S. J. Tumor aneuploidy correlates with markers of immune evasion and with reduced response to immunotherapy. Science 355, eaaf8399 (2017).
Yarilin, D. et al. Machine-based method for multiplex in situ molecular characterization of tissues by immunofluorescence detection. Sci. Rep. 5, 9534 (2015).
Otsu, N. Threshold selection method from gray-level histograms. IEEE T. Syst. Man. Cyb. 9, 62–66 (1979).
He, K., Gkioxari, G., Dollar, P. & Girshick, R. Mask R-CNN. IEEE Trans. Pattern Analysis Machine Intelligence 42, 386–397 (2020).
Loken, M. R., Parks, D. R. & Herzenberg, L. A. Two-color immunofluorescence using a fluorescence-activated cell sorter. J. Histochem. Cytochem. 25, 899–907 (1977).
Acknowledgements
We thank the Thoracic Oncology Service, the Flow Cytometry Core, the Molecular Cytology Core, the Genomic Editing and Screening Core and the Integrated Genomics Operation at MSKCC for their assistance. We especially thank the tissue donors at MSKCC for participating in this study. We thank J. Nainys and V. Kiseliovas for providing hands-on training of single-cell library preparations. We thank E. Azizi and T. Nawy for their critical feedback on this manuscript and F. Bakhoum and V. Bakhoum for their support. This work was supported by National Institutes of Health grants U54-CA209975 (J.M. and D.P.), P01-CA129243 (J.M.), DP1- HD084071 (D.P.), R01CA164729 (D.P.), P30-CA008748 (J.M.); DoD Innovator Award W81XWH-12-0074 (J.M.); and awards from the Burroughs Wellcome Fund Career Award at the Scientific Interface (A.M.L.), the Lung Cancer Research Foundation (A.M.L.), the Alan and Sandra Gerry Metastasis and Tumor Ecosystems Center at MSKCC (A.M.L. and J.H.), the Kirschstein-NRSA F30-CA220954 Predoctoral Fellowship (N.R.C.) and by a T32-GM007729 Medical Scientist Training Program Grant (N.R.C.).
Author information
Authors and Affiliations
Contributions
A.M.L., J.M. and D.P. conceived and oversaw the project, performed data interpretation and wrote the manuscript. A.M.L. generated and analyzed all scRNA-seq data and performed mouse experiments for single cell analyses; J.H. generated additional NK depleted macrometastases for immunofluorescence, which were processed by A.M.L. and J.H. Matched primary-metastasis pairs were identified, analyzed and interpreted by J.H. and S.F.B. NK cell cytotoxicity assays in vitro were designed and performed by J.H. N.R.C. and J.B.X. developed and applied image cytometry methods for NK cell cytotoxicity assays in vitro. M.S., V.-P.L., A.J.C. and S.K. assisted with scRNA-seq analysis. Y.X. developed and applied algorithms to segment nuclei in fixed tissues. N.R., V.A., M.M., J.T.P. and C.M.R. oversaw procurement of tissue samples, clinical specimen processing and histopathologic data interpretation. I.M. and L.M. assisted with scRNA-seq library preparation.
Corresponding authors
Ethics declarations
Competing interests
J.M. is a scientific advisor and owns company stock in Scholar Rock. C.M.R. has consulted with AbbVie, Amgen, Ascentage, Astra Zeneca, BMS, Celgene, Daiichi Sankyo, Genentech/Roche, Ipsen, Loxo and Pharmar, and is on the scientific advisory boards of Elucida and Harpoon. S.F.B. owns equity in, receives compensation from, and serves as a consultant, board member and a scientific advisory board member for Volastra Therapeutics. He also has consulted for Sanofi. All other authors declare no competing conflicts.
Additional information
Peer review information Javier Carmona was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Patient attributes.
Patient resection site, smoking history, primary lesion size, disease stage, diagnostic pathology, oncogenic mutations and treatment history.
Extended Data Fig. 2 Single cell parameters and pre-processing.
Cells were filtered based on (a) cumulative number of transcript counts, (b) cell complexity and (c) fraction of mitochondrial mRNA content detected per cell as described in the Methods; shown here for one representative library. Excluded cells are labeled in red. Histograms showing the distribution of (d) total number of transcripts detected per cell and (e) number of unique genes detected per cell in retained cells colored by sample. f-h, t-SNE projection of the complete atlas of normal lung, primary tumor and metastatic LUAD (same projection as Fig. 1c, n = 40,505 cells), cells colored by (f) sample and (g) meta-cell class as determined by (h) unsupervised clustering of canonical gene signatures within each meta-cell class across all cells. Clustering of canonical cell type expression signatures (annotated in Supplementary Table 2), z-score normalized per gene across cells. Assignment to meta-cell classes (detailed in Methods) are colored on the dendrogram.
Extended Data Fig. 3 Phenotyping myeloid, epithelial and stromal cell types.
a, t-SNE projection of myeloid, epithelial and stromal cells (purple and tan populations from Extended Data Fig. 2g, n = 9,195 cells) colored and labeled by their PhenoGraph cluster assignment (Phenograph run on subset as detailed in Methods). b, Heatmap of gene signatures differentially expressed by PhenoGraph clusters with abs(NES) >2 and padj < 0.05, clustered (rows and columns) according to the cosine distance metric for visualization; hits not meeting these criteria are whited out. Numbered by Phenograph clusters (top) and colored by inferred cell type assignments (bottom, see Methods). NES, normalized enrichment score; padj, Bonferroni corrected, two-sided p-value. c, Pearson correlations between PhenoGraph cluster centroids and bulk mRNA profiles from purified immune subpopulations47,48 (n = 5,987 genes, Methods). Correlation coefficients are whited out if p > 0.01 for the Pearson test for non-correlation. d, t-SNE projection of all myeloid/epithelial/stromal cells (same as a) colored and labeled by inferred cell types. PhenoGraph clusters were mapped to cell types using (b-c) and are directly mapped back to the complete patient dataset in (Fig. 1c) using the same color scheme. e, Clustered heatmap of the average imputed expression per cell type of distinguishing markers, standardized by z-score. Rows are colored by annotated cell type.
Extended Data Fig. 4 Phenotyping NK, T and B cells in the lymphoid compartment.
a, t-SNE projection of all lymphoid cells (blue cells from Extended Data Fig. 2g, n = 25,726 cells) colored by PhenoGraph cluster. b, Pearson correlations between PhenoGraph cluster expression centroids and bulk mRNA data published from purified immune subpopulations47,48 computed based on intersecting, variably expressed genes (n = 5,613, Methods). Rows are colored and labeled by PhenoGraph clusters. Correlation coefficients are whited out if p > 0.01 for the Pearson test for non-correlation. c, Clustered heatmap of the average imputed expression per PhenoGraph cluster of canonical lymphoid markers, standardized by z-scores. Rows are colored by PhenoGraph clusters (left) and annotated cell types (right, see Methods). d, t-SNE projection of all lymphoid cells (same as a) colored and labeled by inferred cell type. PhenoGraph clusters were mapped to cell types using (b, c) and are directly mapped back to the complete patient dataset in (Fig. 1c) using the same color scheme. e, The cell distribution of NKG7 imputed expression, a canonical NK cell marker, across all annotated lymphoid cell types. f, Dot plots showing relative frequency of expressing cells and mean normalized expression (un-imputed data) of canonical markers per lymphoid cell type.
Extended Data Fig. 5 Reproducibility of cell types across patients.
a, Clustered heatmap of the imputed average expression of key cell type markers, for all cell types annotated in the complete patient dataset (Fig. 1c) and standardized by z-scores. Rows are colored by cell type annotation, number of patients in which the cell type was detected, and the total number of cells assigned to this cell type (left to right on left side of clustered heatmap). b, Kernel density plot depicting patient entropy as a measure of sample mixing across all patients within each cell type; computed with bootstrapping to correct for number of cells in each cluster (n = 100 random subsamples) as described in Methods. High entropy indicates most similar cells come from a well-mixed set of patient samples, whereas low entropy indicates most similar cells come from the same patient sample. Distributions are colored by annotated cell type. c, Estimates of tumor purity measured by scRNA-seq and targeted panel DNA sequencing of matched bulk tumor using FACETS, an allele-specific copy number analysis tool19. We test effectiveness of pairing between tumor purity estimates by reporting the Pearson correlation coefficient for n = 7 samples for which matched scRNA-seq and bulk, targeted panel DNA sequencing was available with one-sided P value testing for non-correlation. d, t-SNE projection of individual patient tumors from n = 5 representative patient samples; annotated number of cells per patient. scRNA-seq data for each tumor was processed independently as described in the Methods; each dot represents a cell colored by PhenoGraph cluster, labeled by inferred cell type. e, Number of patients in which each cell type was detected during individual patient analyses. This is concordant with patient frequencies observed in the pooled analysis, summarized in a. Power to detect minority cell types like neutrophils and plasma cells is reduced when analyzing patient samples one-by-one.
Extended Data Fig. 6 Epithelial cell types detected in the normal, adult human lung.
a, Force-directed layout of epithelial cells detected in the normal lung colored by PhenoGraph cluster and labeled with annotated cell type (n = 658 cells; Methods and below). b, Bar graphs representing the fraction of each cell type detected per patient. c, Clustered heatmap of the top 60 DEGs per cell type; imputed values standardized by z-scores are shown for visualization. DEGs were identified in un-imputed data using MAST43 as described in the Methods. Rows are colored by PhenoGraph cluster and labeled by annotated cell type. d, A clustered heatmap of differentially expressed gene signatures within each cell type. NES is shown for pathways in which abs(NES) > 2.5 and padj < 0.05; signatures not meeting this criterium are whited out. Columns are colored by PhenoGraph cluster. NES, normalized enrichment score; padj, Bonferroni corrected, two-sided p-value. Complete GSEA results per PhenoGraph Cluster, including nominal p-values, are provided in Supplementary Table 7. e, Histograms showing the fraction of cells per PhenoGraph cluster (that is cell loadings) expressing at or above the 75th percentile a fraction of each cell type-specific gene signature (AEC1, AEC2, Club and Ciliated) computed on imputed data; each distribution represents cells from one PhenoGraph cluster. Colors associated with each annotated epithelial cell lineage are maintained as in Fig. 2.
Extended Data Fig. 7 The relationship between primary tumor and normal lung epithelial cell types.
a, Top DEGs per PhenoGraph Clusters annotated in Fig. 2b, compared to all other cells, computed using MAST43. Each lineage-specific gene is colored by associated cell type, with diameter proportional to −log10(padj). See Supplementary Table 1 for lineage-specific genes. b, Clustered heatmap of gene signatures differentially enriched (abs(NES) >2 and padj < 0.05) in one or more PhenoGraph Clusters. NES, Normalized Enrichment Score. Column colors (top) correspond to annotated epithelial lineages. Signatures not meeting these criteria are whited out. See Supplementary Table 9 for complete GSEA results per PhenoGraph Cluster. c, Violin plots showing imputed expression of canonical lineage-specific transcription factors (columns) for each annotated epithelial cell lineage (color), scaled such that each plot has the same width; lines distinguish data quartiles. d, Force-directed layout of all epithelia (n = 2,140 cells) colored by extrema of the three most informative diffusion components (DCs, above) and by DC2 (below); GSEA of cells ranked along DC2 are positively enriched for embryonic stem cell gene signatures and pathways associated with proximal cell types, and negatively associated distal cell types. Complete GSEA results for DCs are provided in Supplementary Table 9. e, Fraction of normal- and primary tumor-derived cells comprising the union of all three DC extrema (center values, mean; error bars, 95% confidence interval; points, fraction of cells measured at n = 3 diffusion extrema). f, Fraction of each annotated cell type detected per diffusion component extrema and non-extrema. g, Cumulative imputed expression of a bulk-derived gene signature up-regulated in LUAD and not expressed in non-cancerous epithelium30 evaluated per cell in normal- vs. tumor-derived epithelium (center line, median; box limits, upper and lower quartiles; whiskers, 1.5X interquartile range; points, n = 2,140 individual cells); two-sided p < 0.001, Mann Whitney U test.
Extended Data Fig. 8 Identification of transcriptionally distinct metastatic subpopulations.
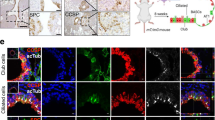
All data in this figure relates to the combined normal, primary tumor and metastatic epithelium. a, Clustered heatmap showing gene signatures differentially expressed across PhenoGraph clusters. Normalized enrichment score (NES) is colored for gene signatures in which abs(NES) > 1.5 and two-sided padj < 0.05. padj, Bonferroni corrected, two-sided p-value. Rows correspond to gene signatures, column corresponds to PhenoGraph clusters. See Supplementary Table 11 for complete GSEA results per cluster. Fraction of each PhenoGraph cluster derived per tissue source is visualized above each column. White stars (bottom) denote patient metastatic clusters based on fraction of metastatic cells (>10%). (b) Tissue source, PhenoGraph clusters (matching those characterized in a), and cell types (annotated as in Fig. 2), are visualized on a force directed layout (n = 3,786 cells). c, SOX2, SOX9 and DAPI immunofluorescence in three additional matched primary tumor-metastasis pairs. Scale bars, 100 µm. d, Fraction of each PhenoGraph cluster detected per metastasis sample (center line, median; box limits, upper and lower quartiles; whiskers, 1.5X interquartile range; points, fraction detected per n = 5 metastatic samples). e, Entropy of patient distribution in each PhenoGraph cluster, computed with bootstrapping to correct for number of cells per cluster (center line, median; box limits, upper and lower quartiles; whiskers, 1.5X interquartile range, n = 100 random subsamples of data). Metastatic clusters are shaded by PhenoGraph cluster ID and clusters are ordered by average entropy. Metastatic clusters associated with alveolar epithelial progenitor signature (type III), and two clusters comprised predominantly of primary tumor cells treated with neoadjuvant therapy are patient-specific. f, Left, Patient metastatic clusters ranked according to average expression of the lung epithelial development signature (n = 34 genes), iteratively leaving out one gene. Each row shows metastatic cluster ranking for each left-out gene. Right, kernel density plot of imputed and normalized expression of each left-out gene.
Extended Data Fig. 9 Developmental stage-specific differential immune sensitivity extended.
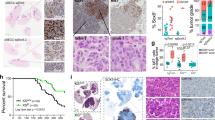
a, Boxplots showing the average expression of NK cell-specific genes in TCGA lung adenocarcinoma patients stratified by SOX2 or SOX9 expression. NK cells are more abundant in SOX9high tumors and conversely, less abundant in SOX2high tumors (two-sided p < 0.001 based on Mann Whitney U test). Center line represents median; box limits, upper and lower quartiles; whiskers, 1.5X interquartile range; points, outliers. b, Cropped Western blots showing SOX2 and SOX9 protein levels upon DOX induction in the H2087-LCC model; no independent repeats were performed. Un-cropped Western blots are provided in Source Data. c, mRNA expression of Sox2 and Sox9 in bulk KP mouse LUAD derivatives38, from primary tumors (circles) or metastases (x); red indicates the derivative is metatstatic. d, Endogenous expression of nuclear SOX9 enumerated by quantitative immunofluorescence in KP482T1 metastatic cells before and after co-culture with IL2-activated mouse NK cells. Fraction of SOX9 positive cells before and after NK cell co-culture are reported. Average of n = 3 technical replicates for each of 3 biological replicates. Between 5 and 9 locations were imaged and quantified per biological and technical replicate; resulting in quantitation of 4,534 cells before NK cell co-culture and 2,556 individual cells after NK cell co-culture. P-values from paired two-tailed t-tests (n = 3 biological replicates, degrees of freedom = 2, t = 8.33, p = 0.01 for SOX9 single positive comparison). e, Relative mRNA expression of NK activating ligands in H2087-LCC cells with and without induction of SOX2 or SOX9 (n = 3 technical replicates; center values, mean; error bars, 95% confidence interval).
Extended Data Fig. 10 Clonality and phenotypic landscape of NK cell-depleted macrometastases.
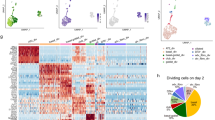
a, Integrated radiance of H2087-LCC cells intracardially injected in mice,+/− anti-GM1 antibody treatment to deplete NK cells measured over time. b, Schematic illustrating trichromatic marking system implemented to assay the clonality of NK cell-depleted metastases. Fluorescence of trichromic reporter visualized in metastatic outbreaks generated in NSG mice lacking NK cells. c, FACS plots shows distribution of cells expressing Cerulean, Venus and mCherry per NK cell-depleted metastasis (lower) as compared to single and multi-color controls (upper); repeated independently for n = 6 NK cell-depleted macrometastases. d, Force-directed layout of all single cells (n = 6,073 cells) transcriptionally profiled from 8 NK cell-depleted macrometastases colored by PhenoGraph cluster. e, Kernel density plot of the imputed expression of SOX2 and SOX9 in each NK cell-depleted PhenoGraph cluster; clusters are ranked by median SOX2 expression. f, A bipartite graph representing genome-wide correlations across all common, variably expressed genes (n = 2,895; Methods) between each NK cell-depleted PhenoGraph cluster (n = 18, circular nodes) and each developmental state observed in human tumors (n = 4, square nodes, annotated in Fig. 3c). The Pearson correlation is computed across all categorical assignments between the two independent sets and edges link NK cell-depleted PhenoGraph clusters to human developmental for Pearson R > 0.20 and two-sided p < 0.05; edge width is scaled by the magnitude of the correlation (observed range: 0.20–0.62). Pearson correlation coefficients are also reported in Supplementary Table 2. Shading is used to highlight nodes assigned the three metastatic states detailed in Fig. 4.
Supplementary information
Supplementary Information
Supplementary Fig. 1.
Supplementary Table 1
Supplementary Table 1
Supplementary Table 2
Supplementary Table 2
Supplementary Table 3
Supplementary Table 3
Supplementary Table 4
Supplementary Table 4
Supplementary Table 5
Supplementary Table 5
Supplementary Table 6
Supplementary Table 6
Supplementary Table 7
Supplementary Table 7
Supplementary Table 8
Supplementary Table 8
Supplementary Table 9
Supplementary Table 9
Supplementary Table 10
Supplementary Table 10
Supplementary Table 11
Supplementary Table 11
Supplementary Table 12
Supplementary Table 12
Supplementary Table 13
Supplementary Table 13
Supplementary Table 14
Supplementary Table 14
Supplementary Table 15
Supplementary Table 15
Supplementary Dataset
Supplementary Dataset
Source data
Source Data Fig. 3
Statistical source data
Source Data Fig. 4
Statistical source data
Source Data Fig. 5
Statistical source data
Source Data Fig. 6
Statistical source data
Source Data Extended Data Fig. 5
Statistical source data
Source Data Extended Data Fig. 9
Statistical source data
Source Data Extended Data Fig. 9
Unprocessed western blots
Source Data Extended Data Fig. 10
Statistical source data
Rights and permissions
About this article
Cite this article
Laughney, A.M., Hu, J., Campbell, N.R. et al. Regenerative lineages and immune-mediated pruning in lung cancer metastasis. Nat Med 26, 259–269 (2020). https://doi.org/10.1038/s41591-019-0750-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-019-0750-6
This article is cited by
-
Single-cell division tracing and transcriptomics reveal cell types and differentiation paths in the regenerating lung
Nature Communications (2024)
-
SOX on tumors, a comfort or a constraint?
Cell Death Discovery (2024)
-
ScRNA-seq of gastric cancer tissues reveals differences in the immune microenvironment of primary tumors and metastases
Oncogene (2024)
-
UPP1 promotes lung adenocarcinoma progression through the induction of an immunosuppressive microenvironment
Nature Communications (2024)
-
scRNA-seq revealed high stemness epithelial malignant cell clusters and prognostic models of lung adenocarcinoma
Scientific Reports (2024)