Abstract
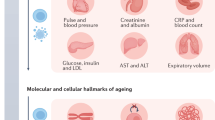
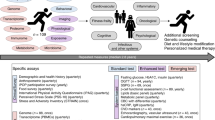
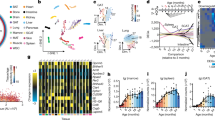
The molecular changes that occur with aging are not well understood1,2,3,4. Here, we performed longitudinal and deep multiomics profiling of 106 healthy individuals from 29 to 75 years of age and examined how different types of ‘omic’ measurements, including transcripts, proteins, metabolites, cytokines, microbes and clinical laboratory values, correlate with age. We identified both known and new markers that associated with age, as well as distinct molecular patterns of aging in insulin-resistant as compared to insulin-sensitive individuals. In a longitudinal setting, we identified personal aging markers whose levels changed over a short time frame of 2–3 years. Further, we defined different types of aging patterns in different individuals, termed ‘ageotypes’, on the basis of the types of molecular pathways that changed over time in a given individual. Ageotypes may provide a molecular assessment of personal aging, reflective of personal lifestyle and medical history, that may ultimately be useful in monitoring and intervening in the aging process.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Raw data included in this study are hosted on the NIH Human Microbiome 2 project site (https://portal.hmpdacc.org) with no restrictions on their use. Exome sequencing data are also available at dbGaP under study accession phs001719.v1.p1. Both raw and processed data are also hosted on the Stanford iPOP site (http://med.stanford.edu/ipop.html). For additional information regarding the study, please contact the corresponding author.
References
Lopez-Otin, C., Blasco, M. A., Partridge, L., Serrano, M. & Kroemer, G. The hallmarks of aging. Cell 153, 1194–1217 (2013).
Aunan, J. R., Cho, W. C. & Soreide, K. The biology of aging and cancer: a brief overview of shared and divergent molecular hallmarks. Aging Dis. 8, 628–642 (2017).
Hung, C. W., Chen, Y. C., Hsieh, W. L., Chiou, S. H. & Kao, C. L. Ageing and neurodegenerative diseases. Ageing Res. Rev. 9(Suppl. 1), S36–S46 (2010).
Kalyani, R. R. & Egan, J. M. Diabetes and altered glucose metabolism with aging. Endocrinol. Metab. Clin. North Am. 42, 333–347 (2013).
Steenman, M. & Lande, G. Cardiac aging and heart disease in humans. Biophys. Rev. 9, 131–137 (2017).
Field, A. E. et al. DNA methylation clocks in aging: categories, causes, and consequences. Mol. Cell 71, 882–895 (2018).
Horvath, S. DNA methylation age of human tissues and cell types. Genome Biol. 14, R115 (2013).
Lu, A. T. et al. DNA methylation GrimAge strongly predicts lifespan and healthspan. Aging 11, 303–327 (2019).
Pani, L. N. et al. Effect of aging on A1C levels in individuals without diabetes: evidence from the Framingham Offspring Study and the National Health and Nutrition Examination Survey 2001–2004. Diabetes Care 31, 1991–1996 (2008).
Bonomini, F., Rodella, L. F. & Rezzani, R. Metabolic syndrome, aging and involvement of oxidative stress. Aging Dis. 6, 109–120 (2015).
Gutch, M., Kumar, S., Razi, S. M., Gupta, K. K. & Gupta, A. Assessment of insulin sensitivity/resistance. Indian J. Endocrinol. Metab. 19, 160–164 (2015).
Knowles, J. W. et al. Measurement of insulin-mediated glucose uptake: direct comparison of the modified insulin suppression test and the euglycemic, hyperinsulinemic clamp. Metabolism 62, 548–553 (2013).
Zhou, W. et al. Longitudinal of host–microbe dynamics in prediabetes. Nature 569, 663–671 (2019).
Schüssler-Fiorenza Rose, S. M. et al. A longitudinal big data approach for precision health. Nat. Med. 25, 792–804 (2019).
Wetzels, J. F., Kiemeney, L. A., Swinkels, D. W., Willems, H. L. & den Heijer, M. Age- and gender-specific reference values of estimated GFR in Caucasians: the Nijmegen Biomedical Study. Kidney Int. 72, 632–637 (2007).
Edelberg, J. M., Cai, D. & Xaymardan, M. Translation of PDGF cardioprotective pathways. Cardiovasc. Toxicol. 3, 27–35 (2003).
Pola, R. et al. Age-dependent VEGF expression and intraneural neovascularization during regeneration of peripheral nerves. Neurobiol. Aging 25, 1361–1368 (2004).
Kolovou, G. et al. Ageing mechanisms and associated lipid changes. Curr. Vasc. Pharmacol. 12, 682–689 (2014).
Biagi, E. et al. Through ageing, and beyond: gut microbiota and inflammatory status in seniors and centenarians. PLoS ONE 5, e10667 (2010).
Galkin, F. et al. Human microbiome aging clocks based on deep learning and tandem of permutation feature importance and accumulated local effects. Preprint at bioRxiv https://doi.org/10.1101/507780 (2018).
Bonafe, M. et al. Polymorphic variants of insulin-like growth factor I (IGF-I) receptor and phosphoinositide 3-kinase genes affect IGF-I plasma levels and human longevity: cues for an evolutionarily conserved mechanism of life span control. J. Clin. Endocrinol. Metab. 88, 3299–3304 (2003).
Mori, K. et al. Fetuin-A is associated with calcified coronary artery disease. Coron. Artery Dis. 21, 281–285 (2010).
Reiner, A. P. et al. PROC, PROCR and PROS1 polymorphisms, plasma anticoagulant phenotypes, and risk of cardiovascular disease and mortality in older adults: the Cardiovascular Health Study. J. Thromb. Haemost. 6, 1625–1632 (2008).
Franceschi, C. & Campisi, J. Chronic inflammation (inflammaging) and its potential contribution to age-associated diseases. J. Gerontol. A Biol. Sci. Med. Sci. 69(Suppl. 1), S4–S9 (2014).
Peters, M. J. et al. The transcriptional landscape of age in human peripheral blood. Nat. Commun. 6, 8570 (2015).
Bae, E., Kim, H. E., Koh, E. & Kim, K. S. Phosphoglucomutase1 is necessary for sustained cell growth under repetitive glucose depletion. FEBS Lett. 588, 3074–3080 (2014).
Liu, Z. et al. A new aging measure captures morbidity and mortality risk across diverse subpopulations from NHANES IV: a cohort study. PLoS Med. 15, e1002718 (2018).
Liu, Z. et al. Correction: a new aging measure captures morbidity and mortality risk across diverse subpopulations from NHANES IV: a cohort study. PLoS Med. 16, e1002760 (2019).
Carter, T. A. et al. Mechanisms of aging in senescence-accelerated mice. Genome Biol. 6, R48 (2005).
Ross, G. R. et al. Enhanced store-operated Ca2+ influx and ORAI1 expression in ventricular fibroblasts from human failing heart. Biol. Open 6, 326–332 (2017).
Kreienkamp, R. et al. A cell-intrinsic interferon-like response links replication stress to cellular aging caused by progerin. Cell Rep. 22, 2006–2015 (2018).
Ma, X. et al. The nuclear receptor RXRA controls cellular senescence by regulating calcium signaling. Aging Cell 17, e12831 (2018).
Cevenini, E. et al. Human models of aging and longevity. Expert Opin. Biol. Ther. 8, 1393–1405 (2008).
Lean, M. E. J., Anderson, A. S., Morrison, C. & Currall, J. Evaluation of a dietary targets monitor. Eur. J. Clin. Nutr. 57, 667–673 (2003).
Hagströmer, M., Oja, P. & Sjöström, M. The International Physical Activity Questionnaire (IPAQ): a study of concurrent and construct validity. Public Health Nutr. 9, 755–762 (2006).
Acknowledgements
We thank numerous colleagues who contributed to this study, including all the authors of the Zhou et al. study13. We thank the Stanford Center for Genomics and Personalized Medicine (SCGPM) for providing the computing and data storage resources. We also thank the Jackson Laboratory for Genomic Medicine for contributing to this project. This work was supported by grants from the National Institutes of Health (NIH) Common Fund Human Microbiome Project (U54DK102556), by a grant award to the SCGPM Genome Sequencing Service Center (NIH, S10OD020141), by the Stanford Clinical and Translational Science Award to Spectrum (NIH, UL1TR001085) and by the Diabetes Genomics and Analysis Core of the Stanford Diabetes Research Center (P30DK116074). S.M.S.-F.R. was supported by NIH grant K08 ES028825.
Author information
Authors and Affiliations
Contributions
S.A., W.Z., M.R.S., K.C. and M. Avina performed experimental bench work and collected data. W.Z. and S.A. analyzed the data and generated results, with contributions from S.M.S.-F.R. M. Avina managed and coordinated the biobank sample inventory. M. Ashland and W.Z. managed the cohort and coordinated clinical visits. A.B. and M.S. obtained funding and provided additional study resources. S.A., W.Z., A.B., S.M.S.-F.R. and M.S. wrote and revised the manuscript. M.S. supervised the overall study.
Corresponding author
Ethics declarations
Competing interests
M.S. is a cofounder of Personalis, Qbio, Sensomics, Mirvie, Filtricine, Protos and January. He is on the scientific advisory board of Jungla, Jupiter and Genapsys.
Additional information
Peer review information Michael Basson was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Significant analytes associated with aging in the cross-section cohort (n = 106).
Left: Number of significant multi-omics molecules correlating with age based on p-value threshold (before multiple hypothesis correction). Right: The categories and their corresponding percentage (frequency) of metabolites significantly associated with the age. Significance is based on the Spearman rank tests.
Extended Data Fig. 2 Scatter plot of number of significant molecules with the age in longitudinal data of 43 individuals.
(Intentionally blank).
Extended Data Fig. 3 Scatter plot showing associations of the magnitude (cumulative trend values of contributing molecules) of each ageotpes with BMI based on 43 individuals.
Associations are not significant. Significance is calculated in the linear regression model.
Extended Data Fig. 4 Scatter plot showing associations of the magnitude (cumulative trend values of contributing molecules) of each ageotpes with Age based on 43 individuals.
Associations are not significant. Significance is calculated in the linear regression model.
Extended Data Fig. 5 Scatter plot showing associations of the magnitude (cumulative trend values of contributing molecules) of each ageotpes with insulin-resistant/sensitive status based on 43 individuals.
Associations are not significant. Significance is calculated in the linear regression model.
Supplementary information
Supplementary Tables
Supplementary Tables 1–18
Rights and permissions
About this article
Cite this article
Ahadi, S., Zhou, W., Schüssler-Fiorenza Rose, S.M. et al. Personal aging markers and ageotypes revealed by deep longitudinal profiling. Nat Med 26, 83–90 (2020). https://doi.org/10.1038/s41591-019-0719-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-019-0719-5
This article is cited by
-
Extracellular vesicle-derived TP53BP1, CD34, and PBX1 from human peripheral blood serve as potential biomarkers for the assessment and prediction of vascular aging
Hereditas (2024)
-
A double inducible cell ablation system for eliminating senescent astrocytes via apoptosis
Molecular Biology Reports (2024)
-
Neutrophil, lymphocyte count, and neutrophil to lymphocyte ratio predict multimorbidity and mortality—results from the Baltimore Longitudinal Study on Aging follow-up study
GeroScience (2024)
-
Validation of biomarkers of aging
Nature Medicine (2024)
-
Machine learning in precision diabetes care and cardiovascular risk prediction
Cardiovascular Diabetology (2023)