Abstract
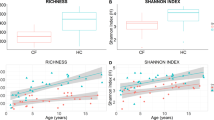
Most infants with cystic fibrosis (CF) have pancreatic exocrine insufficiency that results in nutrient malabsorption and requires oral pancreatic enzyme replacement. Newborn screening for CF has enabled earlier diagnosis, nutritional intervention and enzyme replacement for these infants, allowing most infants with CF to achieve their weight goals by 12 months of age1. Nevertheless, most infants with CF continue to have poor linear growth during their first year of life1. Although this early linear growth failure is associated with worse long-term respiratory function and survival2,3, the determinants of body length in infants with CF have not been defined. Several characteristics of the CF gastrointestinal (GI) tract, including inflammation, maldigestion and malabsorption, may promote intestinal dysbiosis4,5. As GI microbiome activities are known to affect endocrine functions6,7, the intestinal microbiome of infants with CF may also impact growth. We identified an early, progressive fecal dysbiosis that distinguished infants with CF and low length from infants with CF and normal length. This dysbiosis included altered abundances of taxa that perform functions that are important for GI health, nutrient harvest and growth hormone signaling, including decreased abundance of Bacteroidetes and increased abundance of Proteobacteria. Thus, the GI microbiota represent a potential therapeutic target for the correction of low linear growth in infants with CF.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
All metagenomic DNA sequence data required to assess the conclusions of this research are available without restriction from the Sequence Read Archive at the National Center for Biotechnology Information under BioProject accession PRJNA510445.
References
Leung, D. H. et al. Effects of diagnosis by newborn screening for cystic fibrosis on weight and length in the first year of life. JAMA Pediatr. 171, 546–554 (2017).
Konstan, M. W. et al. Growth and nutritional indexes in early life predict pulmonary function in cystic fibrosis. J. Pediatr. 142, 624–630 (2003).
Yen, E. H., Quinton, H. & Borowitz, D. Better nutritional status in early childhood is associated with improved clinical outcomes and survival in patients with cystic fibrosis. J. Pediatr. 162, 530–535 (2013).
Hoffman, L. R. et al. Escherichia coli dysbiosis correlates with gastrointestinal dysfunction in children with cystic fibrosis. Clin. Infect. Dis. 58, 396–399 (2014).
Manor, O. et al. Metagenomic evidence for taxonomic dysbiosis and functional imbalance in the gastrointestinal tracts of children with cystic fibrosis. Sci. Rep. 6, 22493 (2016).
Yan, J. & Charles, J. F. Gut microbiota and IGF-1. Calcif. Tissue Int. 102, 406–414 (2018).
Yan, J. et al. Gut microbiota induce IGF-1 and promote bone formation and growth. Proc. Natl Acad. Sci. USA 113, E7554–E7563 (2016).
Elborn, J. S. Cystic fibrosis. Lancet 388, 2519–2531 (2016).
Borowitz, D. & Gelfond, D. Intestinal complications of cystic fibrosis. Curr. Opin. Pulm. Med. 19, 676–680 (2013).
De Lisle, R. C. & Borowitz, D. The cystic fibrosis intestine. Cold Spring Harb. Perspect. Med. 3, a009753 (2013).
Gelfond, D. & Borowitz, D. Gastrointestinal complications of cystic fibrosis. Clin. Gastroenterol. Hepatol. 11, 333–342 (2013).
Stalvey, M. S. et al. Growth in prepubertal children with cystic fibrosis treated with ivacaftor. Pediatrics 139, e20162522 (2017).
Nielsen, S. et al. Disrupted progression of the intestinal microbiota with age in children with cystic fibrosis. Sci. Rep. 6, 24857 (2016).
Ooi, C. Y. et al. Impact of CFTR modulation with ivacaftor on gut microbiota and intestinal inflammation. Sci. Rep. 8, 17834 (2018).
Segata, N. et al. Metagenomic microbial community profiling using unique clade-specific marker genes. Nat. Methods 9, 811–814 (2012).
Vatanen, T. et al. The human gut microbiome in early-onset type 1 diabetes from the TEDDY study. Nature 562, 589–594 (2018).
Schirmer, M. et al. Compositional and temporal changes in the gut microbiome of pediatric ulcerative colitis patients are linked to disease course. Cell Host Microbe 24, 600–610 (2018).
Backhed, F. et al. Dynamics and stabilization of the human gut microbiome during the first year of life. Cell Host Microbe 17, 690–703 (2015).
Pannaraj, P. S. et al. Association between breast milk bacterial communities and establishment and development of the infant gut microbiome. JAMA Pediatr. 171, 647–654 (2017).
Gupta, R. W. et al. Histamine-2 receptor blockers alter the fecal microbiota in premature infants. J. Pediatr. Gastroenterol. Nutr. 56, 397–400 (2013).
Imhann, F. et al. Proton pump inhibitors affect the gut microbiome. Gut 65, 740–748 (2016).
Subramanian, S. et al. Persistent gut microbiota immaturity in malnourished Bangladeshi children. Nature 510, 417–421 (2014).
Wilson, D. C. & Pencharz, P. B. Nutrition and cystic fibrosis. Nutrition 14, 792–795 (1998).
Matel, J. L. & Milla, C. E. Nutrition in cystic fibrosis. Semin. Respir. Crit. Care Med. 30, 579–586 (2009).
Langdon, A., Crook, N. & Dantas, G. The effects of antibiotics on the microbiome throughout development and alternative approaches for therapeutic modulation. Genome Med. 8, 39 (2016).
Treem, W. R., Ahsan, N., Shoup, M. & Hyams, J. S. Fecal short-chain fatty acids in children with inflammatory bowel disease. J. Pediatr. Gastroenterol. Nutr. 18, 159–164 (1994).
Owira, P. M. & Winter, T. A. Colonic energy salvage in chronic pancreatic exocrine insufficiency. JPEN J. Parenter. Enteral. Nutr. 32, 63–71 (2008).
Vogt, J. A. & Wolever, T. M. Fecal acetate is inversely related to acetate absorption from the human rectum and distal colon. J. Nutr. 133, 3145–3148 (2003).
Boets, E. et al. Quantification of in vivo colonic short chain fatty acid production from inulin. Nutrients 7, 8916–8929 (2015).
Vernocchi, P. et al. Gut microbiota signatures in cystic fibrosis: loss of host CFTR function drives the microbiota enterophenotype. PLoS ONE 13, e0208171 (2018).
Debyser, G. et al. Faecal proteomics: a tool to investigate dysbiosis and inflammation in patients with cystic fibrosis. J. Cyst. Fibros. 15, 242–250 (2016).
Hildebrandt, M. A. et al. High-fat diet determines the composition of the murine gut microbiome independently of obesity. Gastroenterology 137, 1716–1724 (2009).
Huang, Y., Guo, F., Li, Y., Wang, J. & Li, J. Fecal microbiota signatures of adult patients with different types of short bowel syndrome. J. Gastroenterol. Hepatol. 32, 1949–1957 (2017).
Matamouros, S. et al. Adaptation of commensal proliferating Escherichia coli to the intestinal tract of young children with cystic fibrosis. Proc. Natl Acad. Sci. USA 115, 1605–1610 (2018).
Wang, Y. et al. Opportunistic bacteria confer the ability to ferment prebiotic starch in the adult cystic fibrosis gut. Gut Microbes 10, 367–381 (2019).
Piper, H. G. Intestinal microbiota in short bowel syndrome. Semin. Pediatr. Surg. 27, 223–228 (2018).
Bassis, C. M. et al. Comparison of stool versus rectal swab samples and storage conditions on bacterial community profiles. BMC Microbiol. 17, 78 (2017).
Tedjo, D. I. et al. The effect of sampling and storage on the fecal microbiota composition in healthy and diseased subjects. PLoS ONE 10, e0126685 (2015).
Choo, J. M., Leong, L. E. & Rogers, G. B. Sample storage conditions significantly influence faecal microbiome profiles. Sci. Rep. 5, 16350 (2015).
Yu, Z. & Morrison, M. Improved extraction of PCR-quality community DNA from digesta and fecal samples. BioTechniques 36, 808–812 (2004).
Morgan, X. C. et al. Dysfunction of the intestinal microbiome in inflammatory bowel disease and treatment. Genome Biol. 13, R79 (2012).
Breiman, L. Random forests. Mach. Learn. 45, 5–32 (2001).
Deng, H. & Runger, G. Gene selection with guided regularized random forest. Pattern Recognit. 46, 3483–3489 (2013).
Tran, M. et al. The acid steatocrit: a much improved method. J. Pediatr. Gastroenterol. Nutr. 19, 299–303 (1994).
Van den Neucker, A. et al. Clinical use of acid steatocrit. Acta Paediatr. 86, 466–469 (1997).
Acknowledgements
This project was supported by grants from the National Institutes of Health (R01DK095869, K24HL141669, 5P30DK089507) and the Cystic Fibrosis Foundation (SINGH15R0). E.B. is a Faculty Fellow of the Edmond J. Safra Center for Bioinformatics at Tel Aviv University. We gratefully acknowledge the contributions of the principal investigators (including D. Borowitz, B. Ramsey and D. Gelfond), study site coordinators and participants of the BONUS and Healthy Infants Study.
Author information
Authors and Affiliations
Contributions
L.R.H., S.I.M. and E.B. designed the study. C.E.P., K.R.H., A.T.V. and H.S.H. isolated and sequenced metagenomes of stool samples. C.E.P. performed fecal fat assays. A.E., M.J.B. and E.J.W. performed bioinformatic analyses. A.E. and M.J.B. performed statistical analyses with assistance from B.D.M. and S.L.H. H.S.H., A.E., C.E.P., M.J.B., D.H.L., E.B., S.I.M. and L.R.H. analyzed data and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Alison Farrell was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Principal coordinates analysis of the fecal microbiota of infants with CF and controls at month 4.
The structure of the fecal microbiomes of infants in this study at month 4 presented in a multidimensional scaling plot is dominated by the large abundances of B. longum, B. breve and E. coli. One sample is represented for each of 109 infants with CF and 25 controls. Each colored dot represents the microbiota of a different study sample, as indicated.
Extended Data Fig. 2 Phylogenetic plot comparing average microbiota at multiple taxonomic levels at months 4 and 12 for infants with CF and controls.
As indicated in the legend at the lower right, greyscale bars indicate relative abundance, and red or black bars on the outside of the circular graph indicate whether taxa were enriched in infants with CF (black) or controls (red) at month 12.
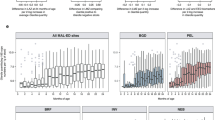
Extended Data Fig. 3 Mean percent relative abundance of genera in Phylum Proteobacteria in fecal samples from infants with CF and normal length, infants with CF and low length and controls.
Proteobacteria remained relatively high in infants with CF, especially those with low length, owing primarily to replacement with E. coli. In addition, abundances of Klebsiella and Enterobacter remained relatively high. Bar heights represents mean percent relative abundance of Proteobacteria at each timepoint, and the mean relative abundance contribution for selected genera are indicated within each bar.
Extended Data Fig. 4 Average percent relative abundance of different bacterial phyla in fecal samples from infants with CF with normal length, infants with CF with low length and controls.
Phylum-level average microbiota at each collection time point of the control infants (left) compared with normal-length infants with CF (middle) and low-length infants with CF (right).
Extended Data Fig. 5 Fecal microbiota development is significantly delayed in infants with CF relative to controls when omitting infants prescribed any acid suppressors and when models are trained on a sparse set of taxonomic features.
a, Infants prescribed any acid suppressors, including proton pump inhibitors and/or H2 blockers were omitted. b, Models trained on a sparse set of taxonomic features. The distributions of relative microbiota age are shown (x axis), following the approach in Subramanian et al.22 for each subject group (infants with CF or controls) using abundances at all taxonomic levels as the normalized error in a sample’s predicted microbiota age when using a computational model constructed for the other group. For example, negative relative microbiota age indicates delayed development compared with the group used to construct the model. The y axis shows density of samples that mapped to a given relative microbiota age at the indicated timepoints. Colored ratios summarize the fraction of replicate full-feature models that produced a distribution of relative microbiota ages that was significantly negatively (green for CF samples relative to control models) or positively (blue for control samples relative to CF models) different from zero. q < 0.01, one-sided Wilcoxon signed-rank test.
Extended Data Fig. 6 Levels of fecal fat percentage and fecal calprotectin during the first year of life.
Lines above the boxes represent significant differences between time points within a cohort (colored lines) or between cohorts at the same time point (black lines). q ≤ 0.01, two-sided Wilcoxon rank-sum test. The boxplot hinges indicate the first and third quartiles, and the whiskers indicate 1.5 times the IQR above and below.
Extended Data Fig. 7 Fecal microbiota development is significantly delayed in infants with CF with low length relative to infants with CF with normal length at month 12.
The distributions of relative microbiota age (x axis) are shown, following the approach in Subramanian et al.22 for each subject group (infants with CF and low length or infants with CF and normal length) as described in Fig. 2. Colored ratios summarize the fraction of replicate full-feature models that produced a distribution of relative microbiota ages that was significantly negatively (red for CF low-length samples relative to CF normal-length models) or positively (purple for CF normal-length samples relative to CF low-length models) different from zero. q < 0.01, one-sided Wilcoxon signed-rank test.
Extended Data Fig. 8 Prevalence of selected butyrate-producing species with significantly different prevalence between infants with CF compared with controls at month 12.
n = 23 for controls and 152 for infants with CF, q < 0.05, chi-squared test.
Supplementary information
Supplementary Information
Supplementary Table 1–7
Rights and permissions
About this article
Cite this article
Hayden, H.S., Eng, A., Pope, C.E. et al. Fecal dysbiosis in infants with cystic fibrosis is associated with early linear growth failure. Nat Med 26, 215–221 (2020). https://doi.org/10.1038/s41591-019-0714-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-019-0714-x
This article is cited by
-
Improved eukaryotic detection compatible with large-scale automated analysis of metagenomes
Microbiome (2023)
-
Cystic Fibrosis-Related Gut Dysbiosis: A Systematic Review
Digestive Diseases and Sciences (2023)
-
CF-Seq, an accessible web application for rapid re-analysis of cystic fibrosis pathogen RNA sequencing studies
Scientific Data (2022)
-
Identifying and preventing cardiovascular disease in patients with cystic fibrosis
Nature Cardiovascular Research (2022)
-
Infants with cystic fibrosis have altered fecal functional capacities with potential clinical and metabolic consequences
BMC Microbiology (2021)