Abstract
Proinflammatory cytokines in the tumor microenvironment can promote tumor growth, yet their value as therapeutic targets remains underexploited. We validated the functional significance of the cardiotrophin-like cytokine factor 1 (CLCF1)–ciliary neurotrophic factor receptor (CNTFR) signaling axis in lung adenocarcinoma (LUAD) and generated a high-affinity soluble receptor (eCNTFR–Fc) that sequesters CLCF1, thereby inhibiting its oncogenic effects. eCNTFR–Fc inhibits tumor growth in multiple xenograft models and in an autochthonous, highly aggressive genetically engineered mouse model of LUAD, driven by activation of oncogenic Kras and loss of Trp53. Abrogation of CLCF1 through eCNTFR–Fc appears most effective in tumors driven by oncogenic KRAS. We observed a correlation between the effectiveness of eCNTFR–Fc and the presence of KRAS mutations that retain the intrinsic capacity to hydrolyze guanosine triphosphate, suggesting that the mechanism of action may be related to altered guanosine triphosphate loading. Overall, we nominate blockade of CLCF1–CNTFR signaling as a novel therapeutic opportunity for LUAD and potentially for other tumor types in which CLCF1 is present in the tumor microenvironment.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information files or are available from the corresponding author upon reasonable request. Statistical Source Data underlying all figures are provided as a separate file with a tab for each panel and the unprocessed western blots as Source Data. A Nature Research Reporting Summary for this article is available as a Supplementary Information file.
References
Ostrem, J. M. & Shokat, K. M. Direct small-molecule inhibitors of KRAS: from structural insights to mechanism-based design. Nat. Rev. Drug Disco. 15, 771–785 (2016).
Cox, A. D., Fesik, S. W., Kimmelman, A. C., Luo, J. & Der, C. J. Drugging the undruggable RAS: mission possible? Nat. Rev. Drug Disco. 13, 828–851 (2014).
Herbst, R. S., Morgensztern, D. & Boshoff, C. The biology and management of non-small cell lung cancer. Nature 553, 446–454 (2018).
Lin, J. J., Riely, G. J. & Shaw, A. T. Targeting ALK: precision medicine takes on drug resistance. Cancer Disco. 7, 137–155 (2017).
Chong, C. R. & Janne, P. A. The quest to overcome resistance to EGFR-targeted therapies in cancer. Nat. Med. 19, 1389–1400 (2013).
Camidge, D. R., Pao, W. & Sequist, L. V. Acquired resistance to TKIs in solid tumours: learning from lung cancer. Nat. Rev. Clin. Oncol. 11, 473–481 (2014).
Forde, P. M. et al. Neoadjuvant PD-1 blockade in resectable lung cancer. N. Engl. J. Med. 379, e14 (2018).
Gandhi, L. et al. Pembrolizumab plus chemotherapy in metastatic non-small-cell lung cancer. N. Engl. J. Med. 378, 2078–2092 (2018).
Hellman, M. D. et al. Nivolumab plus ipilimumab in lung cancer with a high tumor mutational burden. N. Engl. J. Med. 378, 2093–2104 (2018).
Camidge, D. R., Doebele, R. C. & Kerr, K. M. Comparing and contrasting predictive biomarkers for immunotherapy and targeted therapy of NSCLC. Nat. Rev. Clin. Oncol. 16, 341–355 (2019).
Kalluri, R. The biology and function of fibroblasts in cancer. Nat. Rev. Cancer 16, 582–598 (2016).
De Palma, M., Biziato, D. & Petrova, T. V. Microenvironmental regulation of tumour angiogenesis. Nat. Rev. Cancer 17, 457–474 (2017).
Su, S. et al. CD10(+)GPR77(+) cancer-associated fibroblasts promote cancer formation and chemoresistance by sustaining cancer stemness. Cell 172, 841–856, e816 (2018).
Kunita, A. et al. MicroRNA-21 in cancer-associated fibroblasts supports lung adenocarcinoma progression. Sci. Rep. 8, 8838 (2018).
Labernadie, A. et al. A mechanically active heterotypic E-cadherin/N-cadherin adhesion enables fibroblasts to drive cancer cell invasion. Nat. Cell Biol. 19, 224–237 (2017).
Albrengues, J. et al. Epigenetic switch drives the conversion of fibroblasts into proinvasive cancer-associated fibroblasts. Nat. Commun. 6, 10204 (2015).
Chen, W. J. et al. Cancer-associated fibroblasts regulate the plasticity of lung cancer stemness via paracrine signalling. Nat. Commun. 5, 3472 (2014).
Vicent, S. et al. Cross-species functional analysis of cancer-associated fibroblasts identifies a critical role for CLCF1 and IL-6 in non-small cell lung cancer in vivo. Cancer Res. 72, 5744–5756 (2012).
Senaldi, G. et al. Novel neurotrophin-1/B cell-stimulating factor-3: a cytokine of the IL-6 family. Proc. Natl Acad. Sci. USA 96, 11458–11463 (1999).
Lelievre, E. et al. Signaling pathways recruited by the cardiotrophin-like cytokine/cytokine-like factor-1 composite cytokine: specific requirement of the membrane-bound form of ciliary neurotrophic factor receptor alpha component. J. Biol. Chem. 276, 22476–22484 (2001).
Hu, X. et al. Ciliary neurotrophic factor receptor alpha subunit-modulated multiple downstream signaling pathways in hepatic cancer cell lines and their biological implications. Hepatology 47, 1298–1308 (2008).
Kober, P., Bujko, M., Oledzki, J., Tysarowski, A. & Siedlecki, J. A. Methyl-CpG binding column-based identification of nine genes hypermethylated in colorectal cancer. Mol. Carcinog. 50, 846–856 (2011).
Fan, K. et al. Hypomethylation of CNTFR alpha is associated with proliferation and poor prognosis in lower grade gliomas. Sci. Rep. 7, 7079 (2017).
Lu, J. et al. CNTF receptor subunit alpha as a marker for glioma tumor-initiating cells and tumor grade: laboratory investigation. J. Neurosurg. 117, 1022–1031 (2012).
Pento, J. T. Monoclonal antibodies for the treatment of cancer. Anticancer Res. 37, 5935–5939 (2017).
Chiu, M. L. & Gilliland, G. L. Engineering antibody therapeutics. Curr. Opin. Struct. Biol. 38, 163–173 (2016).
Ciombor, K. K. & Berlin, J. Aflibercept–a decoy VEGF receptor. Curr. Oncol. Rep. 16, 368 (2014).
Weinblatt, M. E. et al. A trial of etanercept, a recombinant tumor necrosis factor receptor:Fc fusion protein, in patients with rheumatoid arthritis receiving methotrexate. N. Engl. J. Med. 340, 253–259 (1999).
Economides, A. N. et al. Cytokine traps: multi-component, high-affinity blockers of cytokine action. Nat. Med. 9, 47–52 (2003).
Hunter, S. A. & Cochran, J. R. Cell-binding assays for determining the affinity of protein–protein interactions: technologies and considerations. Methods Enzymol. 580, 21–44 (2016).
Kim, J. W. & Cochran, J. R. Targeting ligand–receptor interactions for development of cancer therapeutics. Curr. Opin. Chem. Biol. 38, 62–69 (2017).
Silver, J. S. & Hunter, C. A. gp130 at the nexus of inflammation, autoimmunity, and cancer. J. Leukoc. Biol. 88, 1145–1156 (2010).
Davis, S. et al. Released form of CNTF receptor-alpha component as a soluble mediator of CNTF responses. Science 259, 1736–1739 (1993).
Auguste, P. et al. Alanine substitution for Thr268 and Asp269 of soluble ciliary neurotrophic factor (CNTF) receptor alpha component defines a specific antagonist for the CNTF response. J. Biol. Chem. 271, 26049–26056 (1996).
Van Deventer, J. A. & Wittrup, K. D. Yeast surface display for antibody isolation: library construction, library screening, and affinity maturation. Methods Mol. Biol. 1131, 151–181 (2014).
Aguinaldo, A. M. & Arnold, F. Staggered extension process (StEP) in vitro recombination. Methods Mol. Biol. 192, 235–239 (2002).
Boder, E. T., Midelfort, K. S. & Wittrup, K. D. Directed evolution of antibody fragments with monovalent femtomolar antigen-binding affinity. Proc. Natl Acad. Sci. USA 97, 10701–10705 (2000).
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. E. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protoc. 10, 845–858 (2015).
Perret, D. et al. Two different contact sites are recruited by cardiotrophin-like cytokine (CLC) to generate the CLC/CLF and CLC/sCNTFR alpha composite cytokines. J. Biol. Chem. 279, 43961–43970 (2004).
Rousseau, F. et al. Ciliary neurotrophic factor, cardiotrophin-like cytokine, and neuropoietin share a conserved binding site on the ciliary neurotrophic factor receptor alpha chain. J. Biol. Chem. 283, 30341–30350 (2008).
Kintzing, J. R., Filsinger Interrante, M. V. & Cochran, J. R. Emerging strategies for developing next-generation protein therapeutics for cancer treatment. Trends Pharm. Sci. 37, 993–1008 (2016).
Ip, N. Y. et al. CNTF and LIF act on neuronal cells via shared signaling pathways that involve the IL-6 signal transducing receptor component gp130. Cell 69, 1121–1132 (1992).
Leibinger, M., Diekmann, H., Fischer, D. & Andreadaki, A. Neuronal STAT3 activation is essential for CNTF- and inflammatory stimulation-induced CNS axon regeneration. Cell Death Dis. 4, e805.
Yao, Z. et al. BRAF mutants evade ERK-dependent feedback by different mechanisms that determine their sensitivity to pharmacologic inhibition. Cancer Cell 28, 370–383 (2015).
Yao, Z. et al. Tumours with class 3 BRAF mutants are sensitive to the inhibition of activated RAS. Nature 548, 234–238 (2017).
Hunter, J. C. et al. Biochemical and structural analysis of common cancer-associated KRAS mutations. Mol. Cancer Res. 13, 1325–1335 (2015).
Kempf, E., Rousseau, B., Besse, B. & Paz-Ares, L. KRAS oncogene in lung cancer: focus on molecularly driven clinical trials. Eur. Respir. Rev. 25, 71–76 (2016).
Tebbutt, N. C. et al. Reciprocal regulation of gastrointestinal homeostasis by SHP2 and STAT-mediated trefoil gene activation in gp130 mutant mice. Nat. Med. 8, 1089–1097 (2002).
Mainardi, S. et al. SHP2 is required for growth of KRAS-mutant non-small cell lung cancer in vivo. Nat. Med. 24, 961–967 (2018).
Ruess, D. A. et al. Mutant KRAS-driven cancers depend on PTPN11/SHP2 phosphatase. Nat. Med. 24, 954–960 (2018).
Nichols, R. J. et al. RAS nucleotide cycling underlies the SHP2 phosphatase dependence of mutant BRAF-, NF1- and RAS-driven cancers. Nat. Cell Biol. 20, 1064–1073 (2018).
Wong, G. S. et al. Targeting wild-type KRAS-amplified gastroesophageal cancer through combined MEK and SHP2 inhibition. Nat. Med. 24, 968–977 (2018).
Jackson, E. L. et al. The differential effects of mutant p53 alleles on advanced murine lung cancer. Cancer Res. 65, 10280–10288 (2005).
Marini, K. D. et al. Inhibition of activin signaling in lung adenocarcinoma increases the therapeutic index of platinum chemotherapy. Sci. Transl. Med. 10, eaat350 (2018).
Janes, M. R. et al. Targeting KRAS mutant cancers with a covalent G12C-specific inhibitor. Cell 172, 578–589 e517 (2018).
Haigis, K. M. KRAS alleles: the devil is in the detail. Trends Cancer 3, 686–697 (2017).
Lito, P., Solomon, M., Li, L. S., Hansen, R. & Rosen, N. Allele-specific inhibitors inactivate mutant KRAS G12C by a trapping mechanism. Science 351, 604–608 (2016).
Saggio, I. et al. Nonradioactive receptor binding assay for ciliary neurotrophic factor. Anal. Biochem. 221, 387–391 (1994).
Sansone, P. et al. IL-6 triggers malignant features in mammospheres from human ductal breast carcinoma and normal mammary gland. J. Clin. Invest. 117, 3988–4002 (2007).
Brooks, G. D. et al. IL6 trans-signaling promotes KRAS-driven lung carcinogenesis. Cancer Res. 76, 866–876 (2016).
Wu, X., Cao, Y., Xiao, H., Li, C. & Lin, J. Bazedoxifene as a novel GP130 inhibitor for pancreatic cancer therapy. Mol. Cancer Ther. 15, 2609–2619 (2016).
Shi, Y. et al. Targeting LIF-mediated paracrine interaction for pancreatic cancer therapy and monitoring. Nature 569, 131–135 (2019).
Liu, J. et al. An integrated TCGA pan-cancer clinical data resource to drive high-quality survival outcome analytics. Cell 173, 400–416 (2018).
Jonkers, J. et al. Synergistic tumor suppressor activity of BRCA2 and p53 in a conditional mouse model for breast cancer. Nat. Genet. 29, 418–425 (2001).
Jackson, E. L. et al. Analysis of lung tumor initiation and progression using conditional expression of oncogenic K-ras. Genes Dev. 15, 3243–3248 (2001).
Wong, J. et al. High-resolution, small animal radiation research platform with X-ray tomographic guidance capabilities. Int. J. Radiat. Oncol. Biol. Phys. 71, 1591–1599 (2008).
Johnstone, C. D. et al. Multi-institutional MicroCT image comparison of image-guided small animal irradiators. Phys. Med. Biol. 62, 5760–5776 (2017).
Johnstone, C. D. & Bazalova-Carter, M. MicroCT imaging dose to mouse organs using a validated monte carlo model of the small animal radiation research platform (SARRP). Phys. Med. Biol. 63, 115012 (2018).
Lee, S. J. et al. Regulation of hypoxia-inducible factor 1alpha (HIF-1alpha) by lysophosphatidic acid is dependent on interplay between p53 and Kruppel-like factor 5. J. Biol. Chem. 288, 25244–25253 (2013).
Acknowledgements
We thank D. Gwinn especially and all of the other members of the Sweet-Cordero and Cochran labs for helpful suggestions as well as S. Hunter, T. Hunter and Y. Shi for assistance with the STAT3 immunohistochemical assay. C.P.M. was supported by a fellowship from the Howard Hughes Medical Institute, by the National Cancer Institute of the National Institutes of Health under the Ruth L. Kirschstein National Service Research Award (NRSA) F31 (F31CA236324) and by the Stanford Medical Scientist Training Program T32 Grant. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. J.W.K. was funded by a graduate fellowship from the Stanford Bio-X Program. K.K. was supported by a Postdoc Mobility grant (P300PB-174377) from the Swiss National Science Foundation. E.A.S.C. and J.R.C. were funded by a grant from the Lungevity Foundation and Upstage Lung Cancer. J.R.C. was also funded by a Stanford Coulter Foundation Translational Partnership Award. E.A.S.C. also received funding from the American Lung Association. This work was also funded by a multi-investigator grant from the National Cancer Institute (R01CA225103) to E.A.S.C. and J.R.C.
Author information
Authors and Affiliations
Contributions
J.W.K., C.P.M., J.R.C. and E.A.S.C. conceived and designed the study. J.W.K. and C.P.M. performed most of the experiments. K.K., K.M. and D.R.S. assisted with the experimental performance. A.L.K. assisted with pathology evaluation and toxicity studies. A.G.L. assisted with human survival data analysis. J.S., I.F., L.P.A. and M.H.G. procured and provided human samples. S.G.L. and L.C.S. processed human samples. S.V. provided initial data on CLCF1 in mouse CAFs. J.W.K., C.P.M., J.R.C. and E.A.S.C. wrote and modified the manuscript. All authors gave intellectual input to the study and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
J.W.K., C.P.M., J.R.C. and E.A.S.C. are included as inventors on intellectual property related to the work described in this manuscript. J.R.C. is a co-founder and Director of xCella Biosciences, which is developing protein therapeutics for oncology.
Additional information
Peer review information Javier Carmona was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 High levels of CLCF1 in KRAS mutant lung adenocarcinoma (LUAD) are associated with decreased patient survival.
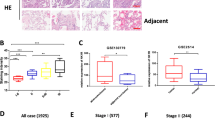
(a) CLCF1 expression is plotted as log2 transcripts per million (TPM) on the y-axis. Each cohort and group are plotted on the x-axis as LUAD (normal; tumor) and lung squamous cell carcinoma (LUSC) (normal; tumor), respectively. Each violin plot is overlaid with a boxplot depicting the maxima, upper quartile, median, lower quartile, and minima for each group. The sample sizes are shown below each plot as n = 59, 515, 511, and 501, respectively. P values were calculated with two-tailed unpaired Student’s t-test and corrected for multiple comparisons with Bonferroni adjustment. (b-e) Kaplan–Meier survival curves for LUAD samples. Samples sizes are listed on each plot. Hazard ratio and its respective P values were calculated with a multivariate Cox hazard regression correcting for age of diagnosis, gender, and cancer stage. Blue solid line represents the normal group (gene expression < 75th percentile) and red solid line represents the high group (gene expression > 75th percentile). (b) CLCF1 expression from samples with KRAS mutation. (c) CLCF1 expression from samples with no KRAS mutations. (d) CNTFR expression from samples with KRAS mutation. (e) CNTFR expression from samples with no KRAS mutations. (f) IL6ST (gp130) expression from samples with KRAS mutation. (g) IL6ST (gp130) expression from samples with no KRAS mutations. (h) LIFR expression from samples with KRAS mutation. (i) LIFR expression from samples with no KRAS mutations. HR = adjusted hazard ratio; CI = confidence interval for the adjusted hazard ratio.
Extended Data Fig. 2 CLCF1 secreted by cancer-associated fibroblasts and LUAD cell lines.
(a) qRT–PCR of CLCF1 mRNA expression in human paired cancer-associated fibroblasts (CAFs) relative to normal lung fibroblasts (NLFs) from the same patient. qRT–PCR data are representative of n = 3 independent experiments. GAPDH and HPRT expression were used for normalization. *** P < 0.001 using two-tailed unpaired Student’s t-test. Data represented as mean ± S.D. (b) Representative cropped western blot analysis of conditioned media CLCF1 levels of indicated LUAD cell lines (representative of n = 2 independent experiments). (c) ELISA results showing CLCF1 levels (pg/mL) in conditioned media from indicated cell lines (n = 3 independent experiments). Data represented as mean ± S.D.
Extended Data Fig. 3 Recombinant production of CLCF1 and soluble β-receptors.
(a) Representative analytical RP-HPLC chromatogram of purified CLCF1. A gradient of 10–60% solvent B (90% acetonitrile/10% water/0.1% trifluoroacetic acid) over 38 min was used. Asterisk (*) indicates peak corresponding to CLCF1. (b) SDS–PAGE of purified CLCF1 under non-reducing and reducing conditions. (c) SDS–PAGE of purified soluble β-receptor constructs under reducing and non-reducing conditions. Asterisk (*) indicates the corresponding band for each protein. All experiments were repeated at least three times with similar results.
Extended Data Fig. 4 CNTFR knockdown decreases proliferation in vitro and tumor growth in vivo.
(a) Proliferation curves for H23, H358, H1975, and H2009, respectively, after CNTFR knockdown by the AlamarBlue proliferation assay (n = 3 independent experiments with three technical replicates per group). *** P < 0.001 using two-way analysis of variance (ANOVA) and Dunnett’s multiple comparison test (DMCT). Data represented as mean ± S.D. (b) Representative images of colony-formation assay in indicated LUAD cell lines (n = 3 independent experiments with similar results). (c) Representative images of spheres from cells grown in anchorage-independent conditions in indicated LUAD cell lines (n = 3 independent experiments with similar results). (d) Tumor volume quantification of H23 xenografts with indicated shRNAs [shGFP (n = 8), shCNTFR 1 (n = 6), and shCNTFR 2 (n = 4) biologically independent samples]. *** P < 0.001 using two-way ANOVA and DMCT. Data represented as mean ± S.E.M. (e) Tumor volume quantification of H2009 xenografts with indicated shRNAs (n = 8 biologically independent samples). *** P < 0.001 using two-way ANOVA and DMCT. Data represented as mean ± S.E.M. (f-g) Representative H&E staining and IHC for phospho-histone H3 (PH3) and cleaved caspase-3 (CC3) from (f) H23 and (g) H2009 xenografts (n = 3 independent experiments with similar results). The quantification of PH3- and CC3-positive foci is presented in Fig. 1o. Scale bars, 50 µm.
Extended Data Fig. 5 CLCF1 knockdown decreases tumor growth and CNTFR knockdown decreases ERK, S6, and STAT3 signaling in vivo.
(a) Tumor volume quantification of final time point of H2009 xenografts with indicated shRNAs (n = 8 biologically independent samples). *** P < 0.001 using one-way ANOVA and Dunnett’s multiple comparison test (DMCT). Whiskers identify the maximum and minimum values; boxes indicate the 75th and 25th percentile and the line the median. (b-d) Representative IHC for phospho-ERK (P-ERK), Phospho-S6RP (P-S6), and phospho-STAT3 (P-STAT3) from A549, H23, and H2009 xenografts. Note: P-STAT3 baseline levels were below the level of detection for H23 (n = 3 independent experiments with similar results). Scale bars, 50 µm.
Extended Data Fig. 6 Schematic representation of the overall engineering strategy.
1) Parental CNTFR was randomly mutagenized via error-prone PCR and the resulting yeast-displayed library was screened to isolate high affinity CLCF1 binders using equilibrium sorting conditions. 2) Twenty clones were randomly selected from enriched yeast pools following sort 3 and sort 4, and the mutant CNTFR DNA was extracted and shuffled together using staggered extension process (StEP). The resulting library was screened against CLCF1 using equilibrium and kinetic off-rate sorting conditions to isolate yeast displaying high affinity binders. After 3 rounds of sorting, a combination of four consensus mutations was determined to comprise the highest affinity CLCF1 binder. 3) A further round of random mutagenesis was performed on this variant (variant 4) and the resulting yeast displayed library was screened to isolate a population that showed reduced binding to LIFR while retaining binding for CLCF1, as shown in Fig. 4e. From this screening approach, two mutations, Y177H and K178N, were identified (Fig. 4f) that conferred reduced LIFR binding. 4) The addition of point mutations T268A and D269A conferred reduced gp130 binding. 5) A combination of these 8 mutations comprised the final CNTFR variant (eCNTFR), an engineered protein that possessed high affinity binding to CLCF1 (R110Q, T174P, S237F, and I287F), and lack of binding to LIFR (Y177H and K178N) and gp130 (T268A and D269A).
Extended Data Fig. 7 Tripartite receptor complex formation on yeast surface.
Two soluble constructs, one containing a hexahistidine tag and another fused to a mouse Fc domain, were prepared for each of the β subunits (LIFR-Fc, LIFR-His, gp130–Fc, and gp130–His). (a) As shown by the flow cytometry dot plots, i) when yeast-displayed CNTFR was incubated with LIFR-Fc (10 nM), no binding signal was detected. ii) Upon addition of CLCF1 (10 nM), LIFR-Fc binding signal increases indicating that in the presence of CLCF1, CNTFR interacts with LIFR. iii) When gp130–His was included in addition to CLCF1 and LIFR-Fc, LIFR-Fc binding signal to yeast-displayed CNTFR increased further. (b) i) Similarly, without CLCF1, gp130–Fc showed no detectable binding to yeast-displayed CNTFR. ii) Upon addition of CLCF1, gp130–Fc signal increased. iii) When LIFR-His was added gp130–Fc binding increased further, suggesting coordinated binding between LIFR and gp130 to the CLCF1–CNTFR complex. All experiments were repeated at least three times with similar results.
Extended Data Fig. 8 Library sorting process for non-LIFR binders.
(a) Representative flow cytometry dot plot showing yeast library sorting process: isolation of non-LIFR binders (Sort 2 with 1 nM LIFR-Fc), isolation of CLCF1 binders (Sort 3 with 0.5 nM CLCF1–His), and isolation of non-LIFR binders (Sort 4 with 10 nM LIFR-Fc). To retain the binding affinity for CLCF1, strategy was alternated between positive screening for 0.5 nM CLCF1 and negative screening for increasing concentrations of LIFR-Fc. (b) Flow cytometry histogram representing yeast-displayed CNTFR variant containing T268A and D269A (CNTFR_AA) binding to CLCF1 (10 nM), gp130–Fc (10 nM), and LIFR–fc (10 nM) relative to untreated negative control (red). CNTFR_AA binds to CLCF1 and LIFR but not to gp130. For gp130–Fc and LIFR-Fc binding studies 10 nM of CLCF1 was added to induce receptor complex formation. All experiments were repeated at least three times with similar results.
Extended Data Fig. 9 Recombinant production of soluble CNTFR proteins.
(a) Overlaid analytical size exclusion chromatography chromatograms of purified wtCNTFR–HIS, wtCNTFR–Fc, eCNTFR–HIS, and eCNTFR–Fc. (b) SDS–PAGE of purified CNTFR constructs under reducing and non-reducing conditions. All experiments were repeated at least three times with similar results.
Extended Data Fig. 10 Yeast-displayed affinity matured eCNTFR does not bind to CNTF but retains high affinity binding to mCLCF1.
(a) Binding curves representing yeast-displayed wild-type CNTFR (wtCNTFR) and engineered CNTFR (eCNTFR) binding to CNTF measured by flow cytometry (n = 3 independent experiments). Data represented as mean ± S.D. (b) Binding curves representing yeast-displayed wtCNTFR and eCNTFR binding to mCLCF1 measured by flow cytometry. While eCNTFR still binds to mCLCF1 with comparable affinity to hCLCF1, there was no detectable binding to CNTF (n = 3 independent experiments). Data represented as mean ± S.D. (c) Representative cropped western blot of two independent experiments showing A549 cells treated with mCLCF1 and phosphorylation of STAT3 (Y705) in a time-dependent manner.
Supplementary information
Supplementary Information
Supplementary Figs. 1–5 and Tables 1–8.
Rights and permissions
About this article
Cite this article
Kim, J.W., Marquez, C.P., Kostyrko, K. et al. Antitumor activity of an engineered decoy receptor targeting CLCF1–CNTFR signaling in lung adenocarcinoma. Nat Med 25, 1783–1795 (2019). https://doi.org/10.1038/s41591-019-0612-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-019-0612-2
This article is cited by
-
FFAR4 activation inhibits lung adenocarcinoma via blocking respiratory chain complex assembly associated mitochondrial metabolism
Cellular & Molecular Biology Letters (2024)
-
Systematic Analysis of the Prognostic Significance and Roles of the Integrin Alpha Family in Non-Small Cell Lung Cancers
Advances in Therapy (2023)
-
Identification of the effects of COVID-19 on patients with pulmonary fibrosis and lung cancer: a bioinformatics analysis and literature review
Scientific Reports (2022)
-
RNA-based therapies: A cog in the wheel of lung cancer defense
Molecular Cancer (2021)
-
An engineered ligand trap inhibits leukemia inhibitory factor as pancreatic cancer treatment strategy
Communications Biology (2021)