Abstract
The MORDOR I trial1, conducted in Niger, Malawi and Tanzania, demonstrated that mass azithromycin distribution to preschool children reduced childhood mortality1. However, the large but simple trial design precluded determination of the mechanisms involved. Here we examined the gut microbiome of preschool children from 30 Nigerien communities randomized to either biannual azithromycin or placebo. Gut microbiome γ-diversity was not significantly altered (P = 0.08), but the relative abundances of two Campylobacter species, along with another 33 gut bacteria, were significantly reduced in children treated with azithromycin at the 24-month follow-up. Metagenomic analysis revealed functional differences in gut bacteria between treatment groups. Resistome analysis showed an increase in macrolide resistance gene expression in gut microbiota in communities treated with azithromycin (P = 0.004). These results suggest that prolonged mass azithromycin distribution to reduce childhood mortality reduces certain gut bacteria, including known pathogens, while selecting for antibiotic resistance.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author upon request. The microbial sequencing reads have been deposited with the NCBI Sequence Read Archive under BioProject no. PRJNA549968. Additional processed data reported in this study are available upon request. Limitations apply to variables that may compromise participant privacy or consent.
References
Keenan, J. D. et al. Azithromycin to reduce childhood mortality in sub-Saharan Africa. N. Engl. J. Med. 378, 1583–1592 (2018).
Jimenez, S. G., Heine, R. G., Ward, P. B. & Robins-Browne, R. M. Campylobacter upsaliensis gastroenteritis in childhood. Pediatr. Infect. Dis. J. 18, 988–992 (1999).
Bourke, B., Chan, V. L. & Sherman, P. Campylobacter upsaliensis: waiting in the wings. Clin. Microbiol. Rev. 11, 440–449 (1998).
Sire, J. M. et al. Community-acquired infectious diarrhoea in children under 5 years of age in Dakar, Senegal. Paediatr. Int. Child Health 33, 139–144 (2013).
Truong, D. T. et al. MetaPhlAn2 for enhanced metagenomic taxonomic profiling. Nat. Methods 12, 902–903 (2015).
Segata, N. et al. Metagenomic biomarker discovery and explanation. Genome Biol. 12, R60 (2011).
Franzosa, E. A. et al. Species-level functional profiling of metagenomes and metatranscriptomes. Nat. Methods 15, 962–968 (2018).
Yoshioka, S. & Newell, P. D. Disruption of de novo purine biosynthesis in Pseudomonas fluorescens Pf0-1 leads to reduced biofilm formation and a reduction in cell size of surface-attached but not planktonic cells. PeerJ. 4, e1543 (2016).
Zhao, H., Patel, V., Helmann, J. D. & Dörr, T. Don’t let sleeping dogmas lie: new views of peptidoglycan synthesis and its regulation. Mol. Microbiol. 106, 847–860 (2017).
Typas, A., Banzhaf, M., Gross, C. A. & Vollmer, W. From the regulation of peptidoglycan synthesis to bacterial growth and morphology. Nat. Rev. Microbiol. 10, 123–136 (2011).
O’Neill, L. A., Kishton, R. J. & Rathmell, J. A guide to immunometabolism for immunologists. Nat. Rev. Immunol. 16, 553–565 (2016).
Doan, T. et al. Macrolide resistance in MORDOR I: a cluster-randomized trial in Niger. N. Engl. J. Med. 380, 2271–2273 (2019).
Oldenburg, C. E. et al. Gut resistome after oral antibiotics in preschool children in BurkinaFaso: a randomized controlled trial. Clin. Infect. Dis. https://doi.org/10.1093/cid/ciz455 (2019).
O’Brien, K. S. et al. Antimicrobial resistance following mass azithromycin distribution for trachoma: a systematic review. Lancet Infect. Dis. 19, e14–e25 (2019).
Doan, T. et al. Macrolide resistance in MORDOR I - A cluster-randomized trial in Niger. N. Engl. J. Med. 380, 2271–2273 (2019).
McMullan, B. J. & Mostaghim, M. Prescribing azithromycin. Aust. Prescr. 38, 87–89 (2015).
Engberg, J., Aarestrup, F. M., Taylor, D. E., Gerner-Smidt, P. & Nachamkin, I. Quinolone and macrolide resistance in Campylobacter jejuni and C. coli: resistance mechanisms and trends in human isolates. Emerg. Infect. Dis. 7, 24–34 (2001).
Nachamkin, I., Ung, H. & Li, M. Increasing fluoroquinolone resistance in Campylobacter jejuni, Pennsylvania, USA, 1982–2001. Emerg. Infect. Dis. 8, 1501–1503 (2002).
Same, R. G. & Tamma, P. D. Campylobacter infections in children. Pediatr. Rev. 39, 533–541 (2018).
Kotloff, K. L. et al. The Global Enteric Multicenter Study (GEMS) of diarrheal disease in infants and young children in developing countries: epidemiologic and clinical methods of the case/control study. Clin. Infect. Dis. 55(Suppl 4), S232–S245 (2012).
Amour, C. et al. Epidemiology and impact of Campylobacter infection in children in 8 low-resource settings: results from the MAL-ED study. Clin. Infect. Dis. 63, 1171–1179 (2016).
Nachamkin, I. Chronic effects of Campylobacter infection. Microbes Infect. 4, 399–403 (2002).
Kotloff, K. L. et al. The incidence, aetiology, and adverse clinical consequences of less severe diarrhoeal episodes among infants and children residing in low-income and middle-income countries: a 12-month case-control study as a follow-on to the Global Enteric Multicenter Study (GEMS). Lancet Glob. Health 7, e568–e584 (2019).
Buss, S. N. et al. Multicenter evaluation of the BioFire FilmArray gastrointestinal panel for etiologic diagnosis of infectious gastroenteritis. J. Clin. Microbiol. 53, 915–925 (2015).
Patrick, M. E. et al. Features of illnesses caused by five species of Campylobacter, Foodborne Diseases Active Surveillance Network (FoodNet)—2010–2015. Epidemiol. Infect. 146, 1–10 (2018).
Bolinger, H. & Kathariou, S. The current state of macrolide resistance in Campylobacter spp.: trends and impacts of resistance mechanisms. Appl. Environ. Microbiol. 83, e00416-17 (2017).
Founou, L. L., Amoako, D. G., Founou, R. C. & Essack, S. Y. Antibiotic resistance in food animals in Africa: a systematic review and meta-analysis. Microb. Drug Resist. 24, 648–665 (2018).
Schiaffino, F. et al. Antibiotic resistance of Campylobacter species in a pediatric cohort study. Antimicrob. Agents Chemother. 63, e01911-18 (2019).
Keenan, J. D. et al. Longer-term assessment of azithromycin for reducing childhood mortality in Africa. N. Engl. J. Med. 380, 2207–2214 (2019).
Doan, T. et al. Gut microbial diversity in antibiotic-naive children after systemic antibiotic exposure: a randomized controlled trial. Clin. Infect. Dis. 64, 1147–1153 (2017).
Chandramohan, D. et al. Effect of adding azithromycin to seasonal malaria chemoprevention. N. Engl. J. Med. 380, 2197–2206 (2019).
Amza, A. et al. Does mass azithromycin distribution impact child growth and nutrition in Niger? A cluster-randomized trial. PLoS Negl. Trop. Dis. 8, e3128 (2014).
Sié, A. et al. Effect of antibiotics on short-term growth among children in Burkina Faso: a randomized trial. Am. J. Trop. Med. Hyg. 99, 789–796 (2018).
Amza, A. et al. A cluster-randomized controlled trial evaluating the effects of mass azithromycin treatment on growth and nutrition in Niger. Am. J. Trop. Med. Hyg. 88, 138–143 (2013).
Doan, T. et al. Mass azithromycin distribution and community microbiome: a cluster-randomized trial. Open Forum Infect. Dis. 5, ofy182 (2018).
Doan, T. et al. Metagenomic DNA sequencing for the diagnosis of intraocular infections. Ophthalmology 124, 1247–1248 (2017).
Fu, L., Niu, B., Zhu, Z., Wu, S. & Li, W. CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics 28, 3150–3152 (2012).
Ziv, J. & Lempel, A. A universal algorithm for sequential data compression. IEEE Trans. Inf. Theory 23, 337–343 (1977).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Kim, D., Song, L., Breitwieser, F. P. & Salzberg, S. L. Centrifuge: rapid and sensitive classification of metagenomic sequences. Genome Res. 26, 1721–1729 (2016).
Lakin, S. M. et al. MEGARes: an antimicrobial resistance database for high throughput sequencing. Nucleic Acids Res. 45, D574–D580 (2017).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Hunt, M. et al. ARIBA: rapid antimicrobial resistance genotyping directly from sequencing reads. Microb. Genom. 3, e000131 (2017).
Gibreel, A. et al. Macrolide resistance in Campylobacter jejuni and Campylobacter coli: molecular mechanism and stability of the resistance phenotype. Antimicrob. Agents Chemother. 49, 2753–2759 (2005).
Luangtongkum, T. et al. Antibiotic resistance in Campylobacter: emergence, transmission and persistence. Future Microbiol. 4, 189–200 (2009).
Jost, L. Entropy and diversity. Oikos 113, 363–375 (2006).
Jost, L. Partitioning diversity into independent alpha and beta components. Ecology 88, 2427–2439 (2007).
Acknowledgements
We thank S. Kumar for the helpful discussions regarding the gut pathogens identified in this study. We thank the program officers from the trial’s sponsor: Bill and Melinda Gates Foundation, R. Izadnegahdar, J. Jacobson, T. Kanyok, E. Shutes, C. Damman, A. Sarangi and L. Lamberti. We also thank the members of the Data and Safety Monitoring Committee: University of Washington, J. Walson; Liverpool School of Tropical Medicine, A. Hightower; Loyola University, E. Anderson; Berhan Public Health & Eye Care Consultancy, W. Alemayehu; Tulane University, L. Rajan. This work was supported by the Bill and Melinda Gates Foundation (award no. OPP1032340 to T.M.L.), the Peierls Foundation (T.M.L.), a Research to Prevent Blindness Career Development Award (award no. 127613A to T.D.) and an unrestricted grant from Research to Prevent Blindness. The funding organizations had no role in the analysis or interpretation of the data or the preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
T.D. and T.M.L designed the experiments and supervised the project. T.D., L.Z., M.E.C., S.S. and C. Chen performed the experiments. T.C.P. performed the randomization. A.M.A., R.M., S.K., A.A., C. Cook, E.L., J.D.K. and T.M.L. oversaw the field work and sample collection. E.D.C. assisted with sample sequencing. T.D., A.H., L.W. and T.C.P. wrote the scripts and performed the bioinformatics analyses with contributions from T.M.L. I.N. assisted with data interpretation. T.D. and T.M.L. wrote the initial draft; all coauthors reviewed the manuscript and agreed to publication.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information: Alison Farrell is the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
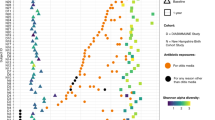
Extended Data Fig. 1 Relative abundances of species significantly different between treatment groups.
Normalized abundance, averaged at the village level, arranged by supervised hierarchical clustering. Thirty-five differential species identified with a false discovery rate < 0.01 and log2(fold change) > 2 (two-sided negative binomial Wald test with Benjamini–Hochberg correction for multiple comparisons).
Extended Data Fig. 2 Resistome of the gut microbiome between treatment groups.
Box plots, with individual values shown as dots, for all classes of resistance determinants, except macrolides, detected in the rectal samples of preschool children at baseline and at 24 months for 30 villages. Each data point is the prevalence of resistance determinants per village. The boxes indicate the median and quartiles of prevalence; the whiskers indicate the estimated 95% CI. Az, azithromycin; Pl, placebo.
Extended Data Fig. 3 Campylobacter macrolide resistance between treatment groups.
Box plots, with individual values shown as dots, of the prevalence of any nucleotide change from A at position 2,075 of the 23S rRNA transcript detected in the rectal samples of preschool children at baseline and at 24 months (n = 30 villages; P = 0.66, ANCOVA). Each data point is the prevalence of resistance transcripts per village. The boxes indicate the median and quartiles of prevalence; the whiskers indicate the estimated 95% CI.
Supplementary information
Supplementary Information
Supplementary Tables 1–6 and Statistical Analysis Plan
Source data
Source Data Fig. 2
Statistical Data Analysis
Source Data Fig. 3
Statistical Data Analysis
Source Data Fig. 4
Statistical Data Analysis
Extended Data Fig. 1
Statistical Data Analysis
Extended Data Fig. 2
Statistical Data Analysis
Extended Data Fig. 3
Statistical Data Analysis
Rights and permissions
About this article
Cite this article
Doan, T., Hinterwirth, A., Worden, L. et al. Gut microbiome alteration in MORDOR I: a community-randomized trial of mass azithromycin distribution. Nat Med 25, 1370–1376 (2019). https://doi.org/10.1038/s41591-019-0533-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-019-0533-0
This article is cited by
-
Infant diarrheal disease in rhesus macaques impedes microbiome maturation and is linked to uncultured Campylobacter species
Communications Biology (2024)
-
Gut microbial CAZymes markers for depression
Translational Psychiatry (2024)
-
Improved eukaryotic detection compatible with large-scale automated analysis of metagenomes
Microbiome (2023)
-
Antibiotic perturbations to the gut microbiome
Nature Reviews Microbiology (2023)
-
Changes of gut microbiota reflect the severity of major depressive disorder: a cross sectional study
Translational Psychiatry (2023)