Abstract
Huntington’s disease (HD) is a dominantly inherited neurodegenerative disorder caused by a CAG trinucleotide expansion in the huntingtin gene (HTT), which codes for the pathologic mutant HTT (mHTT) protein. Since normal HTT is thought to be important for brain function, we engineered zinc finger protein transcription factors (ZFP-TFs) to target the pathogenic CAG repeat and selectively lower mHTT as a therapeutic strategy. Using patient-derived fibroblasts and neurons, we demonstrate that ZFP-TFs selectively repress >99% of HD-causing alleles over a wide dose range while preserving expression of >86% of normal alleles. Other CAG-containing genes are minimally affected, and virally delivered ZFP-TFs are active and well tolerated in HD neurons beyond 100 days in culture and for at least nine months in the mouse brain. Using three HD mouse models, we demonstrate improvements in a range of molecular, histopathological, electrophysiological and functional endpoints. Our findings support the continued development of an allele-selective ZFP-TF for the treatment of HD.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All requests for data will be reviewed by Sangamo Therapeutics, Inc. and the institutions involved to verify whether the request is subject to any intellectual property or confidentiality obligations. If deemed necessary, a material transfer agreement between requestor and Sangamo Therapeutics, Inc. and/or collaborating institutions may be required for sharing of some data. Any data that can be freely shared will be released. All microarray data that support the findings of this study have been deposited in the National Center for Biotechnology Information Gene Expression Omnibus (GEO) and are accessible through the GEO Series accession Nos. GSE127820 and GSE127821.
References
Ross, C. A. & Tabrizi, S. J. Huntington’s disease: from molecular pathogenesis to clinical treatment. Lancet Neurol. 10, 83–98 (2011).
Bates, G. P. et al. Huntington disease. Nat. Rev. Dis. Prim. 1, 15005 (2015).
The Huntington’s Disease Collaborative Research Group. A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington’s disease chromosomes. Cell 72, 971–983 (1993).
Zuccato, C., Valenza, M. & Cattaneo, E. Molecular mechanisms and potential therapeutical targets in Huntington’s disease. Physiol. Rev. 90, 905–981 (2010).
Yamamoto, A., Lucas, J. J. & Hen, R. Reversal of neuropathology and motor dysfunction in a conditional model of Huntington’s disease. Cell 101, 57–66 (2000).
Boudreau, R. L. et al. Nonallele-specific silencing of mutant and wild-type huntingtin demonstrates therapeutic efficacy in Huntington’s disease mice. Mol. Ther. 17, 1053–1063 (2009).
Harper, S. Q. et al. RNA interference improves motor and neuropathological abnormalities in a Huntington’s disease mouse model. Proc. Natl Acad. Sci. USA 102, 5820–5825 (2005).
Kordasiewicz, HollyB. et al. Sustained therapeutic reversal of Huntington’s disease by transient repression of huntingtin synthesis. Neuron 74, 1031–1044 (2012).
Stanek, L. M. et al. Silencing mutant huntingtin by AAV-mediated RNAi ameliorates disease manifestations in the YAC128 mouse model of Huntington’s disease. Hum. Gene Ther. 25, 461–74 (2014).
Grondin, R. et al. Six-month partial suppression of huntingtin is well tolerated in the adult rhesus striatum. Brain 135, 1197–1209 (2012).
McBride, J. L. et al. Preclinical safety of RNAi-mediated HTT suppression in the rhesus macaque as apotential therapy for Huntington’s disease. Mol. Ther. 19, 2152–2162 (2011).
Stiles, D. K. et al. Widespread suppression of huntingtin with convection-enhanced delivery of siRNA. Exp. Neurol. 233, 463–471 (2012).
Duyao, M. P. et al. Inactivation of the mouse Huntington’s disease gene homolog Hdh. Science 269, 407–410 (1995).
Nasir, J. et al. Targeted disruption of the Huntington’s disease gene results in embryonic lethality and behavioral and morphological changes in heterozygotes. Cell 81, 811–823 (1995).
Dragatsis, I., Levine, M. S. & Zeitlin, S. Inactivation of Hdh in the brain and testis results in progressive neurodegeneration and sterility in mice. Nat. Genet. 26, 300–306 (2000).
Arteaga-Bracho, E. E. et al. Postnatal and adult consequences of loss of huntingtin during development:implications for Huntington’s disease. Neurobiol. Dis. 96, 144–155 (2016).
Wang, G., Liu, X., Gaertig, M. A., Li, S. & Li, X. J. Ablation of huntingtin in adult neurons is nondeleterious but its depletion in young mice causes acute pancreatitis. Proc. Natl Acad. Sci. USA 113, 3359–3364 (2016).
Dietrich, P., Johnson, I. M., Alli, S. & Dragatsis, I. Elimination of huntingtin in the adult mouse leads to progressive behavioral deficits, bilateral thalamic calcification, and altered brain iron homeostasis. PLoS Genet. 13, e1006846 (2017).
Lopes, F. et al. Identification of novel genetic causes of Rett syndrome-like phenotypes. J. Med. Genet. 53, 190–199 (2016).
Monteys, A. M., Wilson, M. J., Boudreau, R. L., Spengler, R. M. & Davidson, B. L. Artificial miRNAs targeting mutant huntingtin show preferential silencing in vitro and in vivo. Mol. Ther. Nucleic Acids 4, e234 (2015).
Pfister, E. L. et al. Five siRNAs targeting three SNPs in huntingtin may provide therapy for three-quarters of Huntington’s disease patients. Curr. Biol. 19, 774–778 (2009).
Southwell, A. L. et al. In vivo evaluation of candidate allele-specific mutant huntingtin gene silencing antisense oligonucleotides. Mol. Ther. 22, 2093–2106 (2014).
Gagnon, K. T. et al. Allele-selective inhibition of mutant huntingtin expression with antisense oligonucleotides targeting the expanded CAG repeat. Biochemistry 49, 10166–10178 (2010).
Yu, D. et al. Single-stranded RNAs use RNAi to potently and allele-selectively inhibit mutant huntingtin expression. Cell 150, 895–908 (2012).
Reiter, F., Wienerroither, S. & Stark, A. Combinatorial function of transcription factors and cofactors. Curr. Opin. Genet. Dev. 43, 73–81 (2017).
Marti, E. RNA toxicity induced by expanded CAG repeats in Huntington’s disease. Brain Pathol. 26, 779–786 (2016).
Garriga-Canut, M. et al. Synthetic zinc finger repressors reduce mutant huntingtin expression in the brain of R6/2 mice. Proc. Natl Acad. Sci. USA 109, 3136–3145 (2012).
Agustin-Pavon, C., Mielcarek, M., Garriga-Canut, M. & Isalan, M. Deimmunization for gene therapy: host matching of synthetic zinc finger constructs enables long-term mutant Huntingtin repression in mice. Mol. Neurodegener. 11, 64 (2016).
Gersbach, C. A. & Perez-Pinera, P. Activating human genes with zinc finger proteins, transcription activator-like effectors and CRISPR/Cas9 for gene therapy and regenerative medicine. Expert Opin. Ther. Targets 18, 835–839 (2014).
Boch, J. et al. Breaking the code of DNA binding specificity of TAL-type III effectors. Science 326, 1509–1512 (2009).
Thakore, P. I., Black, J. B., Hilton, I. B. & Gersbach, C. A. Editing the epigenome: technologies for programmable transcription and epigenetic modulation. Nat. Methods 13, 127–137 (2016).
Margolin, J. F. et al. Kruppel-associated boxes are potent transcriptional repression domains. Proc. Natl Acad. Sci. USA 91, 4509–4513 (1994).
Landwehrmeyer, G. B. et al. Data analytics from Enroll-HD, a global clinical research platform for Huntington’s disease. Mov. Disord. Clin. Pract. 4, 212–224 (2017).
Consortium, H. Di Developmental alterations in Huntington’s disease neural cells and pharmacological rescue in cells and mice. Nat. Neurosci. 20, 648–660 (2017).
Mattis, V. B. et al. HD iPSC-derived neural progenitors accumulate in culture and are susceptible to BDNF withdrawal due to glutamate toxicity. Hum. Mol. Genet. 24, 3257–3271 (2015).
Menalled, L. et al. Systematic behavioral evaluation of Huntington’s disease transgenic and knock-in mouse models. Neurobiol. Dis. 35, 319–336 (2009).
Heikkinen, T. et al. Characterization of neurophysiological and behavioral changes, MRI brain volumetry and 1H MRS in zQ175 knock-in mouse model of Huntington’s disease. PLoS ONE 7, e50717 (2012).
Carty, N. et al. Characterization of HTT inclusion size, location, and timing in the zQ175 mouse model of Huntington’s disease: an in vivo high-content imaging study. PLoS ONE 10, e0123527 (2015).
Menalled, L. B. et al. Comprehensive behavioral and molecular characterization of a new knock-in mousemodel of Huntington’s Disease: zQ175. PLoS ONE 7, e49838 (2012).
Plotkin, J. L. & Surmeier, D. J. Corticostriatal synaptic adaptations in Huntington’s disease. Curr. Opin. Neurobiol. 33, 53–62 (2015).
Russell, D. S. et al. Change in PDE10 across early Huntington disease assessed by [18F]MNI-659 and PET imaging. Neurology 86, 748–754 (2016).
Niccolini, F. et al. Striatal molecular alterations in HD gene carriers: a systematic review and meta-analysis of PET studies. J. Neurol. Neurosurg. Psychiatry 89, 185–196 (2018).
Haggkvist, J. et al. Longitudinal small-animal PET imaging of the zQ175 mouse model of Huntington disease shows in vivo changes of molecular targets in the striatum and cerebral cortex. J. Nucl. Med. 58, 617–622 (2017).
Monod, J., Wyman, J. & Changeux, J. P. On the nature of allosteric transitions: a plausible model. J. Mol. Biol. 12, 88–118 (1965).
Pavletich, N. P. & Pabo, C. O. Zinc finger-DNA recognition: crystal structure of a Zif268-DNA complex at 2.1 A. Science 252, 809–817 (1991).
Nekludova, L. & Pabo, C. O. Distinctive DNA conformation with enlarged major groove is found in Zn-finger-DNA and other protein-DNA complexes. Proc. Natl Acad. Sci. USA 91, 6948–6952 (1994).
Mirny, L. A. Nucleosome-mediated cooperativity between transcription factors. Proc. Natl Acad. Sci. USA 107, 22534–22539 (2010).
Iyengar, S. & Farnham, P. J. KAP1 protein: an enigmatic master regulator of the genome. J. Biol. Chem. 286, 26267–26276 (2011).
Lupo, A. et al. KRAB-zinc finger proteins: a repressor family displaying multiple biological functions. Curr. Genom. 14, 268–278 (2013).
Hinde, E., Cardarelli, F. & Gratton, E. Spatiotemporal regulation of oligomerization and dynamics in live cells. Sci. Rep. 5, 12001 (2015).
Sathasivam, K. et al. Aberrant splicing of HTT generates the pathogenic exon 1 protein in Huntington disease. Proc. Natl Acad. Sci. USA 110, 2366–2370 (2013).
Banez-Coronel, M. et al. RAN translation in Huntington disease. Neuron 88, 667–677 (2015).
Capelli, P., Pivetta, C., Soledad Esposito, M. & Arber, S. Locomotor speed control circuits in the caudal brainstem. Nature 551, 373–377 (2017).
Wang, N. et al. Neuronal targets for reducing mutant huntingtin expression to ameliorate disease in a mouse model of Huntington’s disease. Nat. Med. 20, 536–541 (2014).
Indersmitten, T., Tran, C. H., Cepeda, C. & Levine, M. S. Altered excitatory and inhibitory inputs to striatal medium-sized spiny neurons and cortical pyramidal neurons in the Q175 mouse model of Huntington’s disease. J. Neurophysiol. 113, 2953–2966 (2015).
Datson, N. A. et al. The expanded CAG repeat in the huntingtin gene as target for therapeutic RNA modulation throughout the HD mouse brain. PLoS ONE 12, e0171127 (2017).
Russell, D. S. et al. The phosphodiesterase 10 positron emission tomography tracer, [18F]MNI-659, as a novel biomarker for early Huntington disease. JAMA Neurol. 71, 1520–1528 (2014).
Wilson, H. et al. Loss of extra-striatal phosphodiesterase 10A expression in early premanifest Huntington’s disease gene carriers. J. Neurol. Sci. 368, 243–248 (2016).
Bankiewicz, K. S. et al. AAV viral vector delivery to the brain by shape-conforming MR-guided infusions. J. Control Release 240, 434–442 (2016).
Waldvogel, H. J., Kim, E. H., Tippett, L. J., Vonsattel, J. P. & Faull, R. L. The neuropathology of Huntington’s disease. Curr. Top. Behav. Neurosci. 22, 33–80 (2015).
Urnov, F. D., Rebar, E. J., Holmes, M. C., Zhang, H. S. & Gregory, P. D. Genome editing with engineered zinc finger nucleases. Nat. Rev. Genet. 11, 636–646 (2010).
Dull, T. et al. A third-generation lentivirus vector with a conditional packaging system. J. Virol. 72, 8463–8471 (1998).
Trettel, F. et al. Dominant phenotypes produced by the HD mutation in STHdhQ111 striatal cells. Hum. Mol. Genet. 9, 2799–2809 (2000).
Bradley, C. K. et al. Derivation of Huntington’s disease-affected human embryonic stem cell lines. Stem Cells Dev. 20, 495–502 (2011).
Morlan, J., Baker, J. & Sinicropi, D. Mutation detection by real-time PCR: a simple, robust and highly selective method. PloS ONE 4, e4584 (2009).
Latorra, D., Campbell, K., Wolter, A. & Hurley, J. M. Enhanced allele-specific PCR discrimination in SNP genotyping using 3’ locked nucleic acid (LNA) primers. Hum. Mutat. 22, 79–85 (2003).
You, Y., Moreira, B. G., Behlke, M. A. & Owczarzy, R. Design of LNA probes that improve mismatch discrimination. Nucleic Acids Res. 34, e60 (2006).
Plautz, S. A., Boanca, G., Riethoven, J. J. & Pannier, A. K. Microarray analysis of gene expression profiles in cells transfected with nonviral vectors. Mol. Ther. 19, 2144–2151 (2011).
White, J. K. et al. Huntingtin is required for neurogenesis and is not impaired by the Huntington’s disease CAG expansion. Nat. Genet. 17, 404–410 (1997).
Mangiarini, L. et al. Exon 1 of the HD gene with an expanded CAG repeat is sufficient to cause a progressive neurological phenotype in transgenic mice. Cell 87, 493–506 (1996).
Minderer, M. et al. Chronic imaging of cortical sensory map dynamics using a genetically encoded calcium indicator. J. Physiol. 590, 99–107 (2012).
Zolotukhin, S. et al. Recombinant adeno-associated virus purification using novel methods improves infectious titer and yield. Gene Ther. 6, 973–985 (1999).
Langer, O. et al. Precursor synthesis and radiolabelling of the dopamine D2 receptor ligand [11C] raclopride from [11C] methyl triflate. Journal of Labelled Compounds and Radiopharmaceuticals: The Official Journal of the International Isotope Society 42, 1183–1193 (1999).
Nagy, K. et al. Performance evaluation of the small-animal nanoScan PET/MRI system. J. Nucl. Med. 54, 1825–1832 (2013).
Szanda, I. et al. National Electrical Manufacturers Association NU-4 performance evaluation of the PET component of the NanoPET/CT preclinical PET/CT scanner. J. Nucl. Med. 52, 1741–1747 (2011).
Acknowledgements
We thank A. Goodwin, C. Schmidt and I. Cardaun for AAV production. We thank J. Fitzpatrick, W. Arias and T. Schwagarus for stereotaxic injections. We thank J. Weidner for IHC of brain sections. We thank J. Babiarz and E. Leung for bioinformatics analysis. We thank I. Cardaun and S. Kanamkudam for zQ175 primary neuron culture and qPCR. We thank G. Nichol, D. Ando, K. Meyer, A. Manning-Bog and V. Beaumont for discussions. We thank M. Chiocco, D. Klatte, V. Choi, D. Howland, S. Noble, J. Zeitler, M. Stilwell, S. Abrahamson, B. Riley and M. Holmes for reviewing the manuscript. This work was funded by CHDI Foundation and Shire Human Genetic Therapies, Inc., a subsidiary of Shire PLC.
Author information
Authors and Affiliations
Contributions
B.Z., E.J.R., I.M.-S. and H.S.Z. developed the conceptual framework for these studies. J.C.M., L.Z., D.E.P. and E.J.R. contributed to ZFP design. S.J.H., S.L. and R.A. contributed to ZFP assembly. B.Z., K.M., Q.Y., J.R.P., J.M.C., G.Q., M.C.M., R.A., J.L., B.J.V. and A.N. contributed to ZFP testing in Q111/Q7 mouse cells and patient-derived fibroblasts. B.Z. and D.A.S. performed bioinformatics. B.Z., D.A.S., I.A. and D.G. performed microarray analysis. S.F., D.L., L.K., Y.A. and R.S. contributed to ZFP testing in HD NSCs and neurons. B.Z., H.-O.B.N., Y.S., K.T., C.T., A.G., S.K., J.B., L.M, L.P. T.H., K.K.L. and I.M.-S. contributed to zQ175 studies. M.D., J.K. and D.J.S. conducted the electrophysiology studies. J.B., L.M., M.M.S., J.H., L.T., M.T., A.V. and C.H. contributed to the autoradiography and imaging studies. B.Z., Q.Y., A.E.K. and S.R. contributed to the Q50 and R6/2 studies. B.Z., S.F., S.K., J.B., L.M., L.P., I.M.-S. and H.S.Z. designed the experiments. B.Z., F.D.U., P.D.G., E.J.R., I.M.-S. and H.S.Z. supervised the experiments. B.Z. prepared the Figures. B.Z., E.J.R., I.M.-S. and H.S.Z. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
This work was jointly supported by Sangamo Therapeutics, Inc. and CHDI Foundation, Inc. B.Z., K.M., D.A.S., Q.Y., L.Z., D.E.P., S.J.H., S.L., H.-O.B.N., G.Q., M.C.M., R.A., R.S. and E.J.R. are current employees and shareholders of Sangamo Therapeutics, Inc. S.F., D.L., J.R.P., J.C.M., I.A., D.G., L.K., J.M.C., Y.A., J.L., B.J.V., A.N., F.D.U., P.D.G. and H.S.Z. are former employees and shareholders of Sangamo Therapeutics, Inc. and were employed by Sangamo Therapeutics, Inc. when this work was conducted. S.K., J.B., L.M., L.P. and I.M.-S. are employed by CHDI Management, Inc. as advisors to CHDI Foundation, Inc., and declare no conflict of interest. CHDI Foundation, Inc. is a nonprofit biomedical research organization exclusively dedicated to developing therapeutics that substantially improve the lives of those affected by Huntington’s disease, and conducts research in a number of different ways. For the purposes of this manuscript, all research at Psychogenics, Evotec and Charles River Laboratories was conceptualized, planned and directed by the listed authors and conducted under a fee-for-service agreement.
Additional information
Peer review information: Joao Monteiro was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Validation of custom allele-specific RT–qPCR assays and dose-dependent ZFP protein expression.
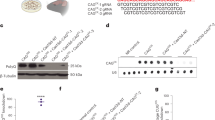
a, Validation of the rs363099 RT–qPCR assay. GM02151 (CAG18/45) and GM04723 (CAG15/67) fibroblasts are heterozygous for the SNP rs363099T/C. In these lines, the rs363099T variant is in phase with normal HTT and rs363099C is in phase with mHTT. Separate plasmid templates harboring the rs363099C or rs363099T amplicons were generated. The rs363099C template was serially diluted twofold, from 5 to 0.156 femtograms (fg), in the presence of a fixed amount (1.25 fg) of the rs363099T template (ratio of rs363099C/rs363099T, 0.125/1–4/1; left panel). Conversely, the rs363099T template was serially diluted twofold, from 5 to 0.156 fg, in the presence of a fixed amount (1.25 fg) of the rs363099C template (ratio of rs363099T/rs363099C, 0.125/1–4/1; right panel). Plasmids were diluted into total cDNA prepared from fibroblasts for which HTT and mHTT expression had been repressed using a strong biallelic HTT repressor. Plasmid inputs were chosen so that the RT–qPCR signal from the 1/1 plasmid ratio gave approximately the same signal as HTT and mHTT in untreated cells. rs363099C- (red) or rs363099T-specific (blue) RT–qPCR assays (see Methods) were performed independently for each of the dilution samples. Relative quantities were scaled to the values obtained for the 1/1 ratio of the rs363099C and rs363099T templates. b, Validation of the rs362331 RT–qPCR assay. ND30259 (CAG21/38) fibroblasts are heterozygous for the C/T SNP rs362331. The rs362331T variant is in phase with normal HTT and rs362331C is in phase with mHTT. Separate plasmid templates harboring rs362331C or rs362331T amplicons were generated and evaluated in the same manner as in a using a reciprocal serial dilution range of 2–0.0625 fg in the presence of 0.5 fg of fixed template. rs362331C- (red) or rs362331T-specific (blue) RT–qPCR assays were performed independently for each of the dilution samples. Relative quantities were scaled to the values obtained for the 1/1 ratio of the rs362331C and rs362331T templates. a,b, n = 3 technical replicates; mean ± s.d. Similar results were replicated in at least one independent experiment for each assay combination. c, Representative cropped image of ZFP-A, -B and -C protein levels assessed by immunoblotting in GM04723 fibroblasts transfected at doses of 1,000, 300, 100, 30 and 10 ng ZFP-TF mRNA. Similar results were replicated in one independent experiment. The uncropped scans are provided as source data. d, Relative quantitation of protein levels from c. ZFP/GAPDH ratios for each sample were scaled to the signal from the 1,000-ng ZFP-C transfected sample.
Extended Data Fig. 2 Evaluation of candidate ZFPs in HdhQ111/HdhQ7 striatal cells and HD fibroblasts.
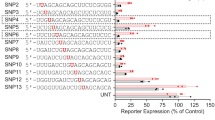
a,b, Comparison of allele selectivity exhibited by candidate ZFP-TF designs in (a) the mouse cells used in a previous study27 (striatal cells derived from the HdhQ111/HdhQ7 mouse model, CAG111/4) versus (b) the human HD fibroblast screening system used in the current study (GM02151, CAG18/45). Only three tested designs (ZFP-A, -B and -D) exhibit highly allele-selective repression in the human HD fibroblast system (>75% repression of the mutant allele with <10% repression of the WT allele, denoted by green circles). In contrast, 25 designs manifest such behavior in the less stringent mouse cells. n = 3 biologically independent samples; mean ± s.d. c, Evaluation of endogenous mHTT allele selectivity for the CAG-targeted ZF6xHunt–Kox-1 (ZF6) and ZF11xHunt–Kox-1 (ZF11) from ref. 27 in GM02151 fibroblasts. Normalized transcript levels for normal (CAG18, blue), mutant (CAG45, red) and total HTT (gray) in fibroblasts transfected with 1,000, 300 or 100 ng ZFP or GFP mRNA. M, mock transfection. n = 4 technical replicates; mean ± s.d.
Extended Data Fig. 3 Validation of NSC and neuronal marker expression in differentiated CAG17/48 hESCs.
a,d, Immunofluorescence staining of (a) PAX6 (red) and NESTIN (green) expression in NSCs differentiated from GENEA020 ESCs and (d) βIII-TUBULIN expression (red) in differentiated neurons (DAPI, blue). Similar results were replicated in three independent experiments. Scale bar, 20 μM. b,c, Normalized transcript levels for a panel of (b) pluripotency markers (OCT4, NANOG and REX1) and (c) NSC markers (PAX6, SOX1 and NES) in ESCs and differentiated NSCs. e,f, Normalized transcript levels for a panel of (e) NSC markers (PAX6, SOX1 and NES) and (f) neuronal markers (MAP2, GAD1 and FOXG1) in differentiated NSCs and neurons. Expression levels were normalized to GAPDH and scaled to the expression of (b) ESCs, (c,e) NSCs or (f) neurons. ESCs, n = 2 differentiated lots; NSCs, n = 2 differentiated lots; neurons, n = 4 differentiated lots; mean ± s.d.
Extended Data Fig. 4 ZFP specificity assessment in vitro by microarray and RT–qPCR.
a, Microarray assessment of GM02151 (CAG18/45) HD fibroblasts transfected with ZFP-A, -B or -C. Each dot represents the mean fold change compared to control-treated cells for a given gene (x value) and the associated P value (y value) determined by an eBayes ANOVA analysis. Genes with ≥ 2 FC change and P < 0.01 are colored red (downregulated) or green (upregulated). ZFP-A, -B and -C: n = 6 biologically independent samples; control: n = 5 biologically independent samples. b,c, RT–qPCR analysis of normal (blue), mutant (red) and total HTT for transfected (b) GM02151 fibroblast and (c) GENEA020 neuron replicates evaluted for microarray analysis. n = 6 biologically independent samples; mean ± s.d.; one-way ANOVA with Tukey’s multiple comparison test; **** P < 0.0001. Allele-specific RT–qPCR was performed as in Fig. 2b (GENEA020 NSCs) and Fig. 1i (GM02151 fibroblasts). d, Twelve genes were selected for RT–qPCR validation of the microarray results. Three genes were chosen for each ZFP based on two criteria: (1) at least one ZFP treatment resulted in ≥ 2 FC change with P < 0.01, and (2) the gene contained ≥6× CAG repeats (allowing up to three mismatches per 6× CAG) within 1 kb of a TSS (see Methods and Supplementary Table 7). An additional three genes were chosen that had large CAG arrays and which were not regulated by any ZFP in the microarray study (‘No ZFP’ group). The distances of the CAG repeat from the TSS, total CAG count and contiguous CAG count are indicated (see Supplementary Table 7). Normal and mutant HTT CAG alleles for GM02151 fibroblasts are shown for reference. e, Regulation of the CAG repeat genes in d was confirmed by RT–qPCR. Normalized expression levels were scaled to the control treatment. n = 6 biologically independent samples; mean ± s.d. f, Venn diagram of regulated genes in a. No genes were commonly regulated (≥ 2 FC change; P < 0.01) by ZFP-A (n = 10) and ZFP-B (n = 5); there was partial overlap with genes regulated by ZFP-C (n = 53).
Extended Data Fig. 5 Additional endpoints evaluated in the R6/2 mouse study.
a, Timeline overview of additional endpoints assessed in WT and AAV2/6-treated R6/2 mice. b–d, Body weight (b), accelerating rotarod (c) and forelimb grip strength (d) were assessed at the indicated timepoints for untreated non-transgenic littermates (WT, blue, n = 20), ZFP-B-treated R6/2 (red, n = 14) or GFP-treated R6/2 mice (gray, n = 14). Data are shown for female (left), male (middle) and gender combined (right) analyses. b, Two-way ANOVA with Tukey’s multiple comparison test; results are presented for treatment versus WT comparisons; * P < 0.05, ** P < 0.01, *** P < 0.001, **** P < 0.0001; mean ± s.d. c, Repeated measures with mixed-effects models; genotype main effect P < 0.0001, treatment effect P = 0.898. Center line, median; limits, 25–75th percentiles; whiskers, min–max; all points shown. d, Repeated measures with mixed-effects models; genotype main effect P = 0.0025; treatment effect P = 0.114; mean ± s.d. c,d, Unpaired two-tailed t-test results are shown for each timepoint: * P < 0.05, **** P < 0.0001.
Extended Data Fig. 6 zQ175 bridging data for ZFP-D and control treatments.
a,b, mRNA (a) or soluble protein (b) levels of WT and mHTT in zQ175 het neurons 10 d after infection with AAV2/1 + 2 ZFP-D as assessed by RT–qPCR (a), Singulex (mouse HTT) or MSD (mHTT) (b). LOD, limit of detection; Bkg, background limit. n = 3 biologically independent samples; mean ± s.d.; one-way ANOVA with Tukey’s multiple comparison test; mean ± s.d.; **** P < 0.0001. c,d, Quantitation of perinuclear (c) or nuclear inclusions (d) in ZFP+ NEUN+ cells for 2–6- (left) or 6–12- (right) month-old cohorts of zQ175 het mice treated with AAV2/1 + 2 ΔDBD or ZFP-D. BLOQ, below limit of quantitation. n = 4 mice; mean ± s.d.; two-tailed unpaired t-test with Welch’s correction; * P < 0.05, ** P < 0.01. e–g, Representative striatal immunostaining images from 2–4- (e) or 6–10- (f,g) month-old zQ175 het cohorts injected with AAV2/1 + 2 GFP (e,f) or ΔDBD.T2A.GFP (g); scale bar, 200 μm. HTT inclusions (mEM48), yellow; GFP, green; DARPP32, red; DAPI, blue. Expanded GFP+ and GFP– regions are shown at right; scale bar, 20 μm. Similar results were replicated in at least one independent experiment. h, Representative striatal GFAP and IBA1 immunostaining images from 6–10-month-old zQ175 het cohorts injected with AAV2/1 + 2 GFP; scale bar, 200 μm. GFP, green; DAPI, blue; GFAP, red; IBA1, yellow. Expanded GFP+ and GFP– regions are shown at right; scale bar, 50 μm. Similar results were replicated in at least one independent experiment. i–n, Quantitation of IBA1+ cells (i,l), GFAP+ cells (j,m) and GFAP intensity (k,n) in WT (left panels) or zQ175 (right panels) striata injected with AAV2/1 + 2 ΔDBD or ZFP-D for 2–6- (i–k) or 6–12- (l–n) month cohorts. n = 4 mice; mean ± s.d.; two-tailed unpaired t-test with Welch’s correction.
Extended Data Fig. 7 Assessment of NEUN+ cells and body weight in ZFP-treated mice.
a–d, Representative whole-brain NEUN and ZFP immunostaining images from 6–12-month-old WT (a,b) and zQ175 het (c,d) cohorts injected with AAV2/1 + 2 ΔDBD in the left hemisphere and either AAV2/1 + 2 ZFP-B (a,c) or ZFP-D (b,d) in the right hemisphere; scale bar, 200 μm. ZFP, green; DAPI, blue; NEUN, red. Expanded regions are shown at left (labeled 1) and right (labeled 2); scale bar, 50 μm. Similar results were replicated in at least one independent experiment. e–h, Quantitation of NEUN+ cells (e,f) and NEUN intensity (g,h) in WT (left panels) or zQ175 het (right panels) mice injected with AAV2/1 + 2 ZFP-B (e,g) or ZFP-D (f,h). e–h. n = 4 mice; mean ± s.d.; two-sided unpaired t-test with Welch’s correction. i,j, Weekly body weight assessments for ZFP-B- (i) and ZFP-D- (j) treated 6-month-old WT and zQ175 het mice until 12 months of age. WT female (red circles, n = 6); WT male (navy triangles, n = 6); zQ175 het female (pink squares, n = 6); zQ175 het male (blue inverted triangles, n = 6); mean ± s.d.
Extended Data Fig. 8 Analysis of CAG repeat content in the human and mouse genomes.
a–d, Distribution of all ≥6× CAG arrays with 0 (a,c) or up to three (b,d) mismatches per CAG hexamer subsite to capture longer, discontiguous repeat tracts (see Methods and Supplementary Tables 8–15) in the human (hg38, orange) and mouse (mm10, blue) genomes. a,b, All CAG arrays represented. c,d, CAG arrays located within 1 kb of a TSS. e,f, Ortholog mapping and corresponding CAG counts for the top 100 genes with the largest CAG repeats within 1 kb of a TSS in the mouse (e) or human (f) genome. A line connects the total CAG count for the mouse gene and its human ortholog (e), or the human gene and its mouse ortholog (f). Search criteria were ≥6× CAG repeats within 1 kb of the TSS with zero (left panel) or up to three (right panel) mismatches per 6× CAG subsite. The number of genes with either no CAG array in the orthologous gene or no ortholog in the other genome is shown at the bottom right of each panel. For clarity, only the top 100 genes are shown; lists were sorted on CAG length, then absolute distance from the TSS. Overlaps may be present for some entries due to multiple ortholog relationships sharing the same CAG counts. See Supplementary Tables 16–19 for complete ortholog lists. g, As in (f) except that only the 12 genes that were selected for monitoring by RT–qPCR are shown (see Extended Data Fig. 9). Only genes with a human ortholog are included.
Extended Data Fig. 9 In vivo examination of neuronal biomarker and CAG repeat gene regulation in treated WT and zQ175 het mice.
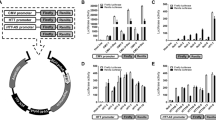
a, Timeline overview for the 1-month in vivo RT–qPCR study. Six-month-old WT or zQ175 het mice were treated with unilateral injections of AAV2/1+2 encoding either ΔDBD (n = 8 WT, n = 9 zQ175), ZFP-B (n = 6 WT, n = 9 zQ175) or ZFP-D (n = 6 WT, n = 8 zQ175) in the right hemisphere. PBS was injected into the left hemisphere. b,c, Normalized allele-specific transcript levels for Q175 (b) and WT (c) Htt. d–u, Normalized RT–qPCR levels for total Htt (d), 12 mouse genes with TSS-adjacent CAG repeats (e–p) and five neuronal biomarker transcripts (q–u). b–u, The ratio of the normalized transcript level for the AAV-treated hemisphere to that of the PBS-treated hemisphere is shown for each mouse in the WT (left panel) and zQ175 het (right panel) treatment groups. Mean ± s.d.; unpaired two-tailed t-test with Welch’s correction. * P < 0.05, ** P < 0.01, *** P < 0.001, **** P < 0.0001. e–p, The length of the closest CAG tract (≤ three mismatches per 6× CAG site; see Extended Data Fig. 8g) and its distance to the TSS are shown for the mouse and human ortholog of each profiled gene.
Extended Data Fig. 10 Tolerability assessment of ZFP-treated WT and zQ175 het mice.
a, Six-month-old zQ175 mice were bilaterally injected with ZFP-D (green, n = 17), eGFP (brown, n = 21) or PBS (blue, n = 21) and monitored for 9 months alongside age-matched WT (black, n = 19) or non-injected zQ175 (red, n = 14) controls. b–e, Open field testing of total (b) or center (c) distance traveled, average velocity (d) and rearing frequency (e). f, Neurological index scores represent the average of 33 metrics (see Methods). g, Average latency to fall using the rotarod test. h, Body weight measurements. i, MRI-based volumetric measurements of whole brain (left), striatum (middle) and cortex (right). Measurements were taken at 5–6 months (baseline, before injection), 9, 12 and 15 months of age (b–g,i) or weekly (h). Repeated measures (b–e,g,i) or ordinary (f,h) two-way ANOVA with Dunnett’s multiple comparison test. b–e,g,i, mean ± s.d. f, Center line, mean; limits, 25–75th percentiles; whiskers, 5–95% confidence interval; points, outliers. h, mean ± s.e.m. b–i, P values are indicated only for comparisons achieving significance; in all cases these correspond to genotype effects. b, * P = 0.0146; zQ175 het versus WT. c, * P = 0.0212; zQ175 het versus WT. g, * P = 0.0257, *** P = 0.0008; zQ175 het versus WT. h, * P < 0.05; zQ175 het versus WT. i, * P < 0.05, ** P < 0.01, *** P < 0.001, **** P < 0.0001; zQ175 het versus WT.
Supplementary information
Supplementary Information
Supplementary Tables 1–3, 6, 7, 20, 21 and Supplementary Data Fig. 1
Supplementary Tables
Supplementary Tables 4, 5, 8–19
Source Data
Source Data Fig. 1
Uncropped western blots
Source Data Fig. 2
Uncropped western blots
Rights and permissions
About this article
Cite this article
Zeitler, B., Froelich, S., Marlen, K. et al. Allele-selective transcriptional repression of mutant HTT for the treatment of Huntington’s disease. Nat Med 25, 1131–1142 (2019). https://doi.org/10.1038/s41591-019-0478-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-019-0478-3
This article is cited by
-
Durable and efficient gene silencing in vivo by hit-and-run epigenome editing
Nature (2024)
-
Decoding the Cellular Trafficking of Prion-like Proteins in Neurodegenerative Diseases
Neuroscience Bulletin (2024)
-
Compact zinc finger architecture utilizing toxin-derived cytidine deaminases for highly efficient base editing in human cells
Nature Communications (2024)
-
Development of super-specific epigenome editing by targeted allele-specific DNA methylation
Epigenetics & Chromatin (2023)
-
hESC-derived striatal progenitors grafted into a Huntington’s disease rat model support long-term functional motor recovery by differentiating, self-organizing and connecting into the lesioned striatum
Stem Cell Research & Therapy (2023)