Abstract
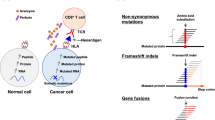
Anti-tumor immunity is driven by self versus non-self discrimination. Many immunotherapeutic approaches to cancer have taken advantage of tumor neoantigens derived from somatic mutations. Here, we demonstrate that gene fusions are a source of immunogenic neoantigens that can mediate responses to immunotherapy. We identified an exceptional responder with metastatic head and neck cancer who experienced a complete response to immune checkpoint inhibitor therapy, despite a low mutational load and minimal pre-treatment immune infiltration in the tumor. Using whole-genome sequencing and RNA sequencing, we identified a novel gene fusion and demonstrated that it produces a neoantigen that can specifically elicit a host cytotoxic T cell response. In a cohort of head and neck tumors with low mutation burden, minimal immune infiltration and prevalent gene fusions, we also identified gene fusion-derived neoantigens that generate cytotoxic T cell responses. Finally, analyzing additional datasets of fusion-positive cancers, including checkpoint-inhibitor-treated tumors, we found evidence of immune surveillance resulting in negative selective pressure against gene fusion-derived neoantigens. These findings highlight an important class of tumor-specific antigens and have implications for targeting gene fusion events in cancers that would otherwise be less poised for response to immunotherapy, including cancers with low mutational load and minimal immune infiltration.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Whole-exome and RNA-sequencing data have been deposited in SRA and are available under project number PRJNA527992.
References
Coulie, P. G., Van den Eynde, B. J., van der Bruggen, P. & Boon, T. Tumour antigens recognized by T lymphocytes: at the core of cancer immunotherapy. Nat. Rev. Cancer 14, 135–146 (2014).
Schreiber, R. D., Old, L. J. & Smyth, M. J. Cancer immunoediting: integrating immunity’s roles in cancer suppression and promotion. Science 331, 1565–1570 (2011).
Sharma, P. & Allison, J. P. The future of immune checkpoint therapy. Science 348, 56–61 (2015).
Rosenberg, S. A. & Restifo, N. P. Adoptive cell transfer as personalized immunotherapy for human cancer. Science 348, 62–68 (2015).
Ott, P. A. et al. An immunogenic personal neoantigen vaccine for patients with melanoma. Nature 547, 217–221 (2017).
Snyder, A. et al. Genetic basis for clinical response to CTLA-4 blockade in melanoma. N. Engl. J. Med. 371, 2189–2199 (2014).
McGranahan, N. et al. Clonal neoantigens elicit T cell immunoreactivity and sensitivity to immune checkpoint blockade. Science 351, 1463–1469 (2016).
Samstein, R. M. et al. Tumor mutational load predicts survival after immunotherapy across multiple cancer types. Nat. Genet. 51, 202–206 (2019).
Ilyas, S. & Yang, J. C. Landscape of tumor antigens in T cell immunotherapy. J. Immunol. 195, 5117–5122 (2015).
Shukla, S. A. et al. Cancer-Germline antigen expression discriminates clinical outcome to CTLA-4 blockade. Cell 173, 624–633 e628 (2018).
Stevanovic, S. et al. Landscape of immunogenic tumor antigens in successful immunotherapy of virally induced epithelial cancer. Science 356, 200–205 (2017).
Gao, Q. et al. Driver fusions and their implications in the development and treatment of human cancers. Cell Rep. 23, 227–238 e223 (2018).
Adams, A. K. et al. DEK promotes HPV-positive and -negative head and neck cancer cell proliferation. Oncogene 34, 868–877 (2015).
Qin, H., Malek, S., Cowell, J. K. & Ren, M. Transformation of human CD34+ hematopoietic progenitor cells with DEK-NUP214 induces AML in an immunocompromised mouse model. Oncogene 35, 5686–5691 (2016).
Forbes, S. A. et al. COSMIC: somatic cancer genetics at high-resolution. Nucleic Acids Res. 45, D777–D783 (2017).
Cancer Genome Atlas, N. Comprehensive genomic characterization of head and neck squamous cell carcinomas. Nature 517, 576–582 (2015).
Scholtalbers, J. et al. TCLP: an online cancer cell line catalogue integrating HLA type, predicted neo-epitopes, virus and gene expression. Genome Med. 7, 118 (2015).
Scheper, W. et al. Low and variable tumor reactivity of the intratumoral TCR repertoire in human cancers. Nat. Med. 25, 89–94 (2018).
Persson, M. et al. Recurrent fusion of MYB and NFIB transcription factor genes in carcinomas of the breast and head and neck. Proc. Natl Acad. Sci. USA 106, 18740–18744 (2009).
Stronen, E. et al. Targeting of cancer neoantigens with donor-derived T cell receptor repertoires. Science 352, 1337–1341 (2016).
McGranahan, N. et al. Allele-Specific HLA loss and immune escape in lung cancer evolution. Cell 171, 1259–1271 e1211 (2017).
Marty, R. et al. MHC-I Genotype restricts the oncogenic mutational landscape. Cell 171, 1272–1283 e1215 (2017).
Shukla, S. A. et al. Comprehensive analysis of cancer-associated somatic mutations in class I HLA genes. Nat. Biotechnol. 33, 1152–1158 (2015).
Angelova M, M. B. et al. Evolution of metastases in space and time under immune selection. Cell 175, 751–765 (2018).
Riaz, N. et al. Tumor and microenvironment evolution during immunotherapy with nivolumab. Cell 171, 934–949 e915 (2017).
Gambacorti-Passerini, C. et al. Human CD4 lymphocytes specifically recognize a peptide representing the fusion region of the hybrid protein pml/RAR alpha present in acute promyelocytic leukemia cells. Blood 81, 1369–1375 (1993).
Buzyn, A. et al. Peptides derived from the whole sequence of BCR-ABL bind to several class I molecules allowing specific induction of human cytotoxic T lymphocytes. Eur. J. Immunol. 27, 2066–2072 (1997).
Yotnda, P. et al. Cytotoxic T cell response against the chimeric ETV6-AML1 protein in childhood acute lymphoblastic leukemia. J. Clin. Invest. 102, 455–462 (1998).
Makita, M. et al. Leukemia-associated fusion proteins, dek-can and bcr-abl, represent immunogenic HLA-DR-restricted epitopes recognized by fusion peptide-specific CD4+ T lymphocytes. Leukemia 16, 2400–2407 (2002).
Sato, Y. et al. Detection and induction of CTLs specific for SYT-SSX-derived peptides in HLA-A24(+) patients with synovial sarcoma. J. Immunol. 169, 1611–1618 (2002).
van den Broeke, L. T., Pendleton, C. D., Mackall, C., Helman, L. J. & Berzofsky, J. A. Identification and epitope enhancement of a PAX-FKHR fusion protein breakpoint epitope in alveolar rhabdomyosarcoma cells created by a tumorigenic chromosomal translocation inducing CTL capable of lysing human tumors. Cancer Res. 66, 1818–1823 (2006).
Lawrence, M. S. et al. Discovery and saturation analysis of cancer genes across 21 tumour types. Nature 505, 495–501 (2014).
Ho, A. S. et al. The mutational landscape of adenoid cystic carcinoma. Nat. Genet. 45, 791–798 (2013).
Dalin, M. G. et al. Multi-dimensional genomic analysis of myoepithelial carcinoma identifies prevalent oncogenic gene fusions. Nat. Commun. 8, 1197 (2017).
Lawrence, M. et al. Software for computing and annotating genomic ranges. PLoS Comput. Biol. 9, e1003118 (2013).
Senbabaoglu, Y. et al. Tumor immune microenvironment characterization in clear cell renal cell carcinoma identifies prognostic and immunotherapeutically relevant messenger RNA signatures. Genome Biol. 17, 231 (2016).
Yoshihara, K. et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat. Commun. 4, 2612 (2013).
Rooney, M. S., Shukla, S. A., Wu, C. J., Getz, G. & Hacohen, N. Molecular and genetic properties of tumors associated with local immune cytolytic activity. Cell 160, 48–61 (2015).
Mandal, R. et al. The head and neck cancer immune landscape and its immunotherapeutic implications. JCI Insight 1, e89829 (2016).
Thorsson, V. et al. The immune landscape of cancer. Immunity 48, 812–830 e814 (2018).
Shen, R. & Seshan, V. E. FACETS: allele-specific copy number and clonal heterogeneity analysis tool for high-throughput DNA sequencing. Nucleic Acids Res. 44, e131 (2016).
Chowell, D. et al. Patient HLA class I genotype influences cancer response to checkpoint blockade immunotherapy. Science 359, 582–587 (2018).
Tran, E. et al. Immunogenicity of somatic mutations in human gastrointestinal cancers. Science 350, 1387–1390 (2015).
Jacob Glanville, H. H. et al. Identifying specificity groups in the T cell receptor repertoire. Nature 547, 94–98 (2017).
Acknowledgements
The authors wish to acknowledge our patients and their families who selflessly contributed time and samples to support this research, the patients and investigators who contributed to the TCGA studies analyzed here, McKenzy and Beth Hupke, members of the Timothy Chan Laboratory for insightful discussions, members of the MSKCC Immunogenomics and Precision Oncology Platform and the Molecular Cytology Core Facility of MSKCC. This work was supported by the NIH/NCI Cancer Center Support Grant no. P30 CA008748 (to MSKCC), Cycle for Survival (R.J.W., L.G.T.M. and T.A.C.), the Frederick Adler Chair at MSKCC, the Jayme Flowers Fund, the Sebastian Nativo Fund, the Damon Runyon Cancer Research Foundation, NIH grant nos. K08 DE024774 and NIH R01 DE027738 (L.G.T.M.), the Adenoid Cystic Carcinoma Cancer Research Foundation, (T.A.C., A.L.H. and L.G.T.M.), The Geoffrey Beene Junior Faculty Chair (A.L.H.), NIH grant nos. R01 CA205426 and NIH R35 CA232097, the Pershing Square Sohn Cancer Research Alliance, the STARR Cancer Consortium and the PaineWebber Chair at MSKCC (T.A.C.).
Author information
Authors and Affiliations
Contributions
W.Y., R. M. Srivastava, K.-W.L., T.A.C. and L.G.T.M. contributed to the study conception and design. L.W., M.A.C., J.T., N.S., A.L.H. and L.G.T.M. contributed to clinical treatment and clinical research coordination. W.Y., R. M. Srivastava, M.G.D., Z.N., J.T. and L.G.T.M. contributed to biospecimen processing. R.G. and N.K. contributed to pathologic analysis. The New York Genome Center (S.K.T., N.R., K.A., H.G. and P.A.) and MSKCC IGO core facility (K.H., N.B. and K.V.) contributed to DNA and RNA sequencing and analyses. V.M., F.K., C.K., D.H., D.C., J.S.S. and L.G.T.M. contributed to the bioinformatics, computational and statistical analyses. W.Y., R. M. Srivastava, K.-W.L. and L.G.T.M. contributed to the experimental design and execution. M.G.D., J.J.H., R.M., R. M. Samstein, N.R, T.A.C., I.G., A.L.H., R.J.W. and L.G.T.M. contributed to the interpretation of data. W.Y., K.-W.L., T.A.C. and L.G.T.M. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
K.-W.L. and J.S.S. are now full-time employees of Regeneron Pharmaceuticals. R. M. Srivastava received speaker fees and travel reimbursement from Innovent Biologics, Inc. A.L.H. receives research funding from Eli Lilly, Genentech, AstraZeneca, Bayer, Kura Oncology, Kotan Pharmaceuticals, Eisai, Bristol-Myers Squibb, Astellas Pharma, Novartis, Merck, Pfizer, Ayla Pharmaceuticals, Allos Therapuetics, Daiichi Sankyo, consulting fees from Bristol-Myers Squibb, Eisai, Genzyme, Merck, Novartis, Sun Pharma, Regeneron, TRM Oncology, Ayala Pharmaceuticals, AstraZeneca, Sanofi Aventis, and travel reimbursement from Janssen Oncology, Merck, Kura Oncology, Ignyta, Ayala Pharmacueticals. J.J.H’s spouse is a full-time employee of Regeneron Pharmaceuticals. R. M. Samstein, T.A.C. and L.G.T.M. are inventors on a provisional patent application (62/569,053) filed by Memorial Sloan Kettering (MSK) relating to the use of TMB in cancer immunotherapy. D.C. and T.A.C. are inventors on a PCT patent application (PCT/US2015/062208) filed by MSK relating to the use of TMB in cancer immunotherapy. MSK and the inventors may receive a share of commercialization revenue from license agreements relating to these patent applications. T.A.C. is a co-founder of Gritstone Oncology and holds equity. He acknowledges grant funding from Bristol-Myers Squibb, AstraZeneca, Illumina, Pfizer, An2H, and Eisai, and has served as an advisor for Bristol-Myers Squibb, Illumina, Eisai and An2H. L.G.T.M. received consulting fees from Rakuten Aspyrian and speaker fees from Physician Educational Resources.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
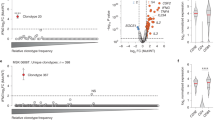
Extended Data Fig. 1 Visualization of DEK–AFF2 and AFF2–DEK gene fusions.
a,b, Visualization of the DEK–AFF2 (a) and AFF2–DEK (b) gene fusions in the primary tumor of patient MSK-HN1 shown on IGV plots of RNA-seq data.
Extended Data Fig. 2 Screen of SNV-derived and alternative splicing-derived nonamer peptides.
a, A screen for binding of SNV-derived nonamer peptides (10 µM) to HLA-A*02:01 on T2 cells reveals no peptides with significant binding affinity. The MFI values are normalized to DMSO. NY-ESO-1 was used as a positive control. b, IFN-γ ELISpot assay of patient MSK-HN1 T cells after 18 h co-culture with autologous PBMCs (n = 3) pulsed with 10 µM of indicated peptides derived from mutations. Due to limited numbers of autologous PBMCs, in several samples (<1>, <2>, <3>, <4>, <5> and <6>), multiple mutant-derived peptides corresponding to a single wild-type peptide were pulsed together. The grouped peptides are indicated. c, IFN-γ ELISpot assay of patient MSK-HN1 T cells after 18 h co-culture with autologous PBMCs (n = 3) pulsed with 10 µM of indicated peptides derived from potential alternative splicing events. Means ± s.e.m. are plotted, with sample n representing the number of independently treated samples.
Extended Data Fig. 3 DEK–AFF2 generates an immunostimulatory peptide recognized by autologous T cells.
a, Flow cytometry analysis of CD137 expression on CD8+ T cells after 18 h co-culture with patient MSK-HN1 PBMCs pulsed with indicated peptides. Data are representative of two independent experiments. b, IFN-γ ELISpot assay of patient MSK-HN1’s T cells after 18 h co-culture with autologous PBMCs (n = 3) which have been pulsed with DMSO or DEKSEEEVS peptide, co-treated with either immunoglobulin G (IgG) control or anti-MHC-I antibody overnight (two-tailed t-tests, 95% CI = −2.187 to +19.85, effect size η2 = 0.685, P = 0.042). c, IFN-γ ELISpot assay of patient MSK-HN1’s T cells after 18 h co-culture with SCC-9 expressing DEK-N-term or DEK–AFF2 fusion (n = 3). Cells are treated with either IgG control or anti-MHC-I antibody (two-tailed t-tests, 95% CI = −278.3 to −147.7, η2 = 0.954, P = 0.0008). d, IFN-γ ELISpot assay of patient MSK-HN1’s T cells after 18 h co-culture with COS-7 cells (n = 3) co-transfected with HLA-C*04:01 plasmid and pLVX–DEK-N-term or pLVX–DEK–AFF2. T cells and COS-7 cells were used at a 6:1 ratio (two-tailed t-test, 95% CI = 48.73–571.9, η2 = 0.731, P = 0.0301). e, Active caspase-3 staining of SCC-9 target cells (n = 2) expressing either DEK-N-term or DEK–AFF2 fusion after 3 h incubation with MSK-HN1 CD8 + TEM cells (CCR7-CD45RA-) at the indicated ratios. Means ± s.e.m. are plotted, with sample n representing the number of independently treated samples.
Extended Data Fig. 4 Screen of HLA binding by fusion peptides predicted to bind to members of HLA-A2 alleles.
a,b, The graphs show stabilization of HLA-A*02:01 on the surface of T2 cells by MYB–NFIB and MYBL1–NFIB peptides (a) and NFIB–MYB peptides (b). The peptides in which we further tested are indicated by arrows. Data are representative of two independent experiments.
Extended Data Fig. 5 Validation of MYB–NFIB and NFIB–MYB fusions.
Validation of MYB–NFIB and NFIB–MYB fusions expressed in patient ACC_M9 by visualization on IGV plots of RNA-seq data.
Extended Data Fig. 6 Schematic of ACC_M9-, ACC_M1-, ACC_P11-, ACC_P14-derived MYB–NFIB fusion constructs that were cloned into pcRNA6SL.
The amino acid sequences surrounding the junctions are shown. Predicted HLA-A2-binding peptides derived from each fusion are indicated.
Extended Data Fig. 7 MYB–NFIB generates an immunostimulatory peptide recognized by patient ACC_M9’s T cells.
a, IFN-γ ELISpot assay of patient ACC_M9’s T cells after 18 h co-culture with healthy donor HD1 dendritic cells electroporated with 2 µg of in vitro transcribed mRNA as described in the methods. b, IFN-γ ELISpot assay of patientACC_M9’s T cells after 18 h co-culture with T2 cells pulsed with 10 µM of indicated peptides. Data are representative of two independent experiments.
Extended Data Fig. 8 PD-1, CD40L and CD137 expression on patient ACC_M9’s T cells after 18 h co-culture with autologous PBMCs pulsed with the indicated peptides.
Flow cytometry analysis of patient ACC_M9’s PBMCs after 18 h pulse with the indicated peptides (n = 3). The immunogenic peptide QFIDSSWYL led to an increased fraction of CD8+ T cells that are CD137+, CD40L+ or PD-1+. Data are representative of three independent experiments.
Extended Data Fig. 9 Patient ACC_M9’s CD8+ T cells that specifically bind to HLA-A*02:01-presented QFIDSSWYL peptide proliferate over 21 days during co-culture with irradiated peptide-pulsed T2 cells.
Patient ACC_M9’s T cells were expanded on irradiated T2 pulsed with 10 µM of indicated peptides over 21 days and stained with QFIDSSWYL–dextramer–PE. A population of QFIDSSWYL-specific T cells is selectively expanded. Data are representative of two independent experiments.
Extended Data Fig. 10 Healthy donor T cells are stimulated by MYB-NFIB-derived and NFIB-MYB-derived peptides.
IFN-γ ELISpot assay of healthy donors HD2 and HD3 T cells after 18 h co-culture with T2 cells pulsed with 10 µM of indicated peptides. Data are representative of two independent experiments.
Supplementary Information
Supplementary Information
Supplementary Figs. 1–8, Supplementary Tables 1–5, 7–9
Supplementary Tables
Supplementary Tables 6, 10–12
Source data
Rights and permissions
About this article
Cite this article
Yang, W., Lee, KW., Srivastava, R.M. et al. Immunogenic neoantigens derived from gene fusions stimulate T cell responses. Nat Med 25, 767–775 (2019). https://doi.org/10.1038/s41591-019-0434-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-019-0434-2
This article is cited by
-
Tumour circular RNAs elicit anti-tumour immunity by encoding cryptic peptides
Nature (2024)
-
Discovery of lung adenocarcinoma tumor antigens and ferroptosis subtypes for developing mRNA vaccines
Scientific Reports (2024)
-
Computational immunogenomic approaches to predict response to cancer immunotherapies
Nature Reviews Clinical Oncology (2024)
-
Molecularly defined sinonasal malignancies: an overview with focus on the current WHO classification and recently described provisional entities
Virchows Archiv (2024)
-
Molecular pathology in diagnosis and prognostication of head and neck tumors
Virchows Archiv (2024)