Abstract
Chronic inflammation is postulated to be involved in the development of end-stage renal disease in diabetes, but which specific circulating inflammatory proteins contribute to this risk remain unknown. To study this, we examined 194 circulating inflammatory proteins in subjects from three independent cohorts with type 1 and type 2 diabetes. In each cohort, we identified an extremely robust kidney risk inflammatory signature (KRIS), consisting of 17 proteins enriched in tumor necrosis factor-receptor superfamily members, that was associated with a 10-year risk of end-stage renal disease. All these proteins had a systemic, non-kidney source. Our prospective study findings provide strong evidence that KRIS proteins contribute to the inflammatory process underlying end-stage renal disease development in both types of diabetes. These proteins point to new therapeutic targets and new prognostic tests to identify subjects at risk of end-stage renal disease, as well as biomarkers to measure responses to treatment of diabetic kidney disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Global proteomic profiling coming from the prospective study followed for ESRD risk are provided in the Supplementary Information of this article. The datasets that support the findings of this study are available in an anonymous manner from the corresponding authors upon request.
References
Gregg, E. W. et al. Changes in diabetes-related complications in the United States, 1990-2010. N. Engl. J. Med. 370, 1514–1523 (2014).
Macisaac, R. J., Ekinci, E. I. & Jerums, G. Markers of and risk factors for the development and progression of diabetic kidney disease. Am. J. Kidney. Dis. 63, S39–S62 (2014).
Pichler, R., Afkarian, M., Dieter, B. P. & Tuttle, K. R. Immunity and inflammation in diabetic kidney disease: translating mechanisms to biomarkers and treatment targets. Am. J. Physiol. Renal Physiol. 312, F716–F731 (2017).
Niewczas, M. A. et al. Circulating TNF receptors 1 and 2 predict ESRD in type 2 diabetes. J. Am. Soc. Nephrol. 23, 507–515 (2012).
Skupien, J. et al. Synergism between circulating tumor necrosis factor receptor 2 and HbA(1c) in determining renal decline during 5-18 years of follow-up in patients with type 1 diabetes and proteinuria. Diabetes Care 37, 2601–2608 (2014).
Gohda, T. et al. Circulating TNF receptors 1 and 2 predict stage 3 CKD in type 1 diabetes. J. Am. Soc. Nephrol. 23, 516–524 (2012).
Pavkov, M. E. et al. Elevation of circulating TNF receptors 1 and 2 increases the risk of end-stage renal disease in American Indians with type 2 diabetes. Kidney Int. 87, 812–819 (2015).
Forsblom, C. et al. Added value of soluble tumor necrosis factor-alpha receptor 1 as a biomarker of ESRD risk in patients with type 1 diabetes. Diabetes Care 37, 2334–2342 (2014).
Saulnier, P. J. et al. Association of serum concentration of TNFR1 with all-cause mortality in patients with type 2 diabetes and chronic kidney disease: follow-up of the SURDIAGENE Cohort. Diabetes Care 37, 1425–1431 (2014).
Coca, S. G. et al. Plasma biomarkers and kidney function decline in early and established diabetic kidney disease. J. Am. Soc. Nephrol. 28, 2786–2793 (2017).
Barr, E. L. M. et al. High baseline levels of tumor necrosis factor receptor 1 are associated with progression of kidney disease in indigenous Australians with diabetes: the eGFR follow-up study. Diabetes Care 41, 739–747 (2018).
Yamanouchi, M. et al. Improved clinical trial enrollment criterion to identify patients with diabetes at risk of end-stage renal disease. Kidney Int. 92, 258–266 (2017).
Gold, L. et al. Aptamer-based multiplexed proteomic technology for biomarker discovery. PLoS ONE 5, e15004 (2010).
Tuerk, C. & Gold, L. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science 249, 505–510 (1990).
Krolewski, A. S. Progressive renal decline: the new paradigm of diabetic nephropathy in type 1 diabetes. Diabetes Care 38, 954–962 (2015).
Nelson, R. G. et al. Development and progression of renal disease in Pima Indians with non-insulin-dependent diabetes mellitus. diabetic renal disease study group. N. Engl. J. Med. 335, 1636–1642 (1996).
Nair, V. et al. A molecular morphometric approach to diabetic kidney disease can link structure to function and outcome. Kidney Int. 93, 439–449 (2018).
Saito, R. et al. Systems biology analysis reveals role of MDM2 in diabetic nephropathy. JCI Insight 1, e87877 (2016).
Beckerman, P. et al. Human kidney tubule-specific gene expression based dissection of chronic kidney disease traits. EBioMedicine 24, 267–276 (2017).
Bossen, C. et al. Interactions of tumor necrosis factor (TNF) and TNF receptor family members in the mouse and human. J. Biol. Chem. 281, 13964–13971 (2006).
Kanehisa, M. et al. Data, information, knowledge and principle: back to metabolism in KEGG. Nucleic Acids Res. 42, D199–D205 (2014).
Locksley, R. M., Killeen, N. & Lenardo, M. J. The TNF and TNF receptor superfamilies: integrating mammalian biology. Cell 104, 487–501 (2001).
Tanamas, S. K. et al. Long-term effect of losartan on kidney disease in American Indians with type 2 diabetes: a follow-up analysis of a randomized clinical trial. Diabetes Care 39, 2004–2010 (2016).
Tuttle, K. R. et al. JAK1/JAK2 inhibition by baricitinib in diabetic kidney disease: results from a Phase 2 randomized controlled clinical trial. Nephrol. Dial. Transplant. 33, 1950–1959 (2018).
Gohda, T. et al. Clinical predictive biomarkers for normoalbuminuric diabetic kidney disease. Diabetes Res. Clin. Pract. 141, 62–68 (2018).
Al-Lamki, R. S. & Mayadas, T. N. TNF receptors: signaling pathways and contribution to renal dysfunction. Kidney Int. 87, 281–296 (2015).
Cui, C. Y. & Schlessinger, D. EDA signaling and skin appendage development. Cell Cycle 5, 2477–2483 (2006).
Sica, G. L. et al. RELT, a new member of the tumor necrosis factor receptor superfamily, is selectively expressed in hematopoietic tissues and activates transcription factor NF-kappaB. Blood 97, 2702–2707 (2001).
Tam, S. J. et al. Death receptors DR6 and TROY regulate brain vascular development. Dev. Cell 22, 403–417 (2012).
McInnes, I. B. & Gracie, J. A. Interleukin-15: a new cytokine target for the treatment of inflammatory diseases. Curr. Opin. Pharmacol. 4, 392–397 (2004).
Araki, S. et al. Predictive impact of elevated serum level of IL-18 for early renal dysfunction in type 2 diabetes: an observational follow-up study. Diabetologia 50, 867–873 (2007).
Cortvrindt, C., Speeckaert, R., Moerman, A., Delanghe, J. R. & Speeckaert, M. M. The role of interleukin-17A in the pathogenesis of kidney diseases. Pathology 49, 247–258 (2017).
Kitching, A. R. & Holdsworth, S. R. The emergence of TH17 cells as effectors of renal injury. J. Am. Soc. Nephrol. 22, 235–238 (2011).
Tesch, G. H. MCP-1/CCL2: a new diagnostic marker and therapeutic target for progressive renal injury in diabetic nephropathy. Am. J. Physiol. Renal Physiol. 294, F697–F701 (2008).
Forssmann, U. et al. chemokines with unique biochemical properties: HCC-1/CCL14a and HCC-2/CCL15. J. Leukoc. Biol. 70, 357–366 (2001).
Schulz-Knappe, P. et al. HCC-1, a novel chemokine from human plasma. J. Exp. Med. 183, 295–299 (1996).
Sharma, K., Susztak, K. & Pennathur, S. Introduction: Systems biology of kidney disease. Semin. Nephrol. 38, 99–100 (2018).
Bohle, A. et al. The pathogenesis of chronic renal failure in diabetic nephropathy. Investigation of 488 cases of diabetic glomerulosclerosis. Pathol. Res. Pract. 187, 251–259 (1991).
Kojima, H., Kim, J. & Chan, L. Emerging roles of hematopoietic cells in the pathobiology of diabetic complications. Trends Endocrinol. Metab. 25, 178–187 (2014).
Chow, F., Ozols, E., Nikolic-Paterson, D. J., Atkins, R. C. & Tesch, G. H. Macrophages in mouse type 2 diabetic nephropathy: correlation with diabetic state and progressive renal injury. Kidney Int. 65, 116–128 (2004).
Wheelock, K. M. et al. White blood cell fractions correlate with lesions of diabetic kidney disease and predict loss of kidney function in Type 2 diabetes. Nephrol. Dial. Transplant. 32, 2145 (2017).
Omote, K. et al. Role of the TNF pathway in the progression of diabetic nephropathy in KK-A(y) mice. Am. J. Physiol. Renal Physiol. 306, F1335–F1347 (2014).
Navarro, J. F., et al. Tumor necrosis factor-α gene expression in diabetic nephropathy: relationship with urinary albumin excretion and effect of angiotensin-converting enzyme inhibition. Kidney Int. Suppl. 68, S98–S102 (2005).
Menne, J. et al. C-C motif-ligand 2 inhibition with emapticap pegol (NOX-E36) in type 2 diabetic patients with albuminuria. Nephrol. Dial. Transplant. 32, 307–315 (2017).
Navarro-Gonzalez, J. F. et al. Effect of pentoxifylline on renal function and urinary albumin excretion in patients with diabetic kidney disease: the PREDIAN trial. J. Am. Soc. Nephrol. 26, 220–229 (2015).
Moriwaki, Y. et al. Effect of TNF-alpha inhibition on urinary albumin excretion in experimental diabetic rats. Acta Diabetol. 44, 215–218 (2007).
Croft, M., Benedict, C. A. & Ware, C. F. Clinical targeting of the TNF and TNFR superfamilies. Nat. Rev. Drug. Discov. 12, 147–168 (2013).
Croft, M. & Siegel, R. M. Beyond TNF: TNF superfamily cytokines as targets for the treatment of rheumatic diseases. Nat. Rev. Rheumatol. 13, 217–233 (2017).
O’Shea, J. J., Kanno, Y. & Chan, A. C. In search of magic bullets: the golden age of immunotherapeutics. Cell 157, 227–240 (2014).
Ridker, P. M. et al. Antiinflammatory therapy with canakinumab for atherosclerotic disease. N. Engl. J. Med. 377, 1119–1131 (2017).
Mehrotra, P. et al. IL-17 mediates neutrophil infiltration and renal fibrosis following recovery from ischemia reperfusion: compensatory role of natural killer cells in athymic rats. Am. J. Physiol. Renal Physiol. 312, F385–F397 (2017).
Anders, H. J. et al. A chemokine receptor CCR-1 antagonist reduces renal fibrosis after unilateral ureter ligation. J. Clin. Invest. 109, 251–259 (2002).
Vielhauer, V. et al. CCR1 blockade reduces interstitial inflammation and fibrosis in mice with glomerulosclerosis and nephrotic syndrome. Kidney Int. 66, 2264–2278 (2004).
Krolewski, A. S., Skupien, J., Rossing, P. & Warram, J. H. Fast renal decline to end-stage renal disease: an unrecognized feature of nephropathy in diabetes. Kidney Int. 91, 1300–1311 (2017).
Krolewski, A. S. et al. Early progressive renal decline precedes the onset of microalbuminuria and its progression to macroalbuminuria. Diabetes Care 37, 226–234 (2014).
Nowak, N. et al. Markers of early progressive renal decline in type 2 diabetes suggest different implications for etiological studies and prognostic tests development. Kidney Int. 93, 1198–1206 (2018).
Nelson, R. G. et al. Incidence of end-stage renal disease in type 2 (non-insulin-dependent) diabetes mellitus in Pima Indians. Diabetologia 31, 730–736 (1988).
Ganz, P. et al. Development and validation of a protein-based risk score for cardiovascular outcomes among patients with stable coronary heart disease. JAMA 315, 2532–2541 (2016).
Ngo, D. et al. Aptamer-based proteomic profiling reveals novel candidate biomarkers and pathways in cardiovascular disease. Circulation 134, 270–285 (2016).
Williams, S. A. et al. Improving assessment of drug safety through proteomics: early detection and mechanistic characterization of the unforeseen harmful effects of torcetrapib. Circulation 137, 999–1010 (2018).
Assarsson, E. et al. Homogenous 96-plex PEA immunoassay exhibiting high sensitivity, specificity, and excellent scalability. PLoS ONE 9, e95192 (2014).
Gray, K. A., Yates, B., Seal, R. L., Wright, M. W. & Bruford, E. A. Genenames.org: the HGNC resources in 2015. Nucleic Acids Res. 43, D1079–D1085 (2015).
Afshinnia, F. et al. Lipidomic signature of progression of chronic kidney disease in the chronic renal insufficiency cohort. Kidney Int. Rep. 1, 256–268 (2016).
Valeri, L. & Vanderweele, T. J. Mediation analysis allowing for exposure-mediator interactions and causal interpretation: theoretical assumptions and implementation with SAS and SPSS macros. Psychol. Methods 18, 137–150 (2013).
Acknowledgments
This study was supported by grants: from the National Institutes of Health to A.S.K. (DK41526 and DP3DK112177), K.S. (DK 087635 and DK108220) and the Joslin Diabetes Center (P30 DK036836); from the Novo Nordisk Foundation to A.S.K. (NNF OC0013659); and from the JDRF to M.A.N. (5-CDA-2015-89-A-B). The study was also supported by the Intramural Research Program of the National Institute of Diabetes and Digestive and Kidney Diseases. P.F. is supported by the Romeo ed Enrica Invernizzi Foundation. We would like to acknowledge E. Mills and N. Rashidi from Olink Proteomics Inc. for their assistance with protein measurements.
Author information
Authors and Affiliations
Contributions
M.A.N. contributed to the design of the study, supervised proteomics data collection, conducted the data analysis, interpreted the results and wrote the manuscript. J.S, A. Smiles, A. Schlafly, E.S. and C.A.S. were involved in data collection and data management of the Joslin Kidney Study, performed preliminary data analyses and reviewed the manuscript. A.D., Z.I.M.D, H.S. and J.K.S. designed the Joslin study on retinopathy, contributed to eye data collection and analysis, and edited and reviewed the manuscript. R.G.N., M.E.P., P.-J.S. and H.C.L. were responsible for design and implementation of the Pima Indian Study, contributed to the proteomic data collection in the Pima Indian Study, performed preliminary data analysis, and reviewed and edited the manuscript. M.K., V.N. and R.G.N. designed the expression study in Pima Indians, provided data analysis and reviewed the manuscript. J.P., C.Q. and K.S. designed the expression study (1KGP) used in the present study, performed the data analyses, and interpreted the results and edited the manuscript. P.F. and C.F.W. were involved in the interpretation of the results of the study and edited the manuscript. K.L.D. and J.M.W. provided the samples from the baricitinib study, facilitated measurements on the Olink platform, reviewed the analysis, and reviewed and edited the manuscript. A.S.K designed the whole study, supervised all aspects of the study implementation, planned and contributed the data analysis, interpreted the data, and contributed to writing the manuscript. M.A.N. and A.S.K. are the guarantors of this work and, as such, had full access to all the data in the study and take responsibility for the integrity of the data and the accuracy of the data analysis.
Corresponding authors
Ethics declarations
Competing interests
A.S.K. and M.A.N. are co-inventors of the TNF-R1 and TNF-R2 patent for predicting risk of ESRD. This patent was licensed by the Joslin Diabetes Center to EKF Diagnostics. The other authors of this report declare no competing conflicts of interest. The findings and conclusions in this report are those of the authors and do not necessarily represent the official position of the Centers for Disease Control and Prevention.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Circulating KRIS proteins are enriched in TNF-RSF members.
a, Distribution of the inflammatory classes among a proteomic platform (SOMAscan) array; and b, within the KRIS in the Joslin cohorts (n = 363). A two-sided Fisher’s exact test detected enrichment in TNF-RSF members (P = 0.007, the class marked in dark red), but not in other inflammatory classes (shades of gray). Results extracted from the multivariate screen – volcano plot (Fig. 1a). ILEU, interleukins; ILEUR, interleukin receptors; CHK, chemokines; CHKR, chemokine receptors; CPL, complement proteins; IFN, interferons; TNFL, TNF superfamily ligands; TNF-RSF, TNF-receptor superfamily members, VR, varia or other inflammatory proteins.
Extended Data Fig. 2 Circulating KRIS proteins and progressive renal function decline rate over time in three cohorts.
Spearman’s rank correlation coefficients (r) between baseline concentration of KRIS proteins and renal function decline rate (eGFR loss – Joslin; GFR loss – Pima) over 8–11 years of follow-up in the Joslin cohorts (n = 363) and the Pima cohort (n = 162). Red bars are a graphic representation of the effect size. Corresponding two-sided P values have been transformed into their base 10 logarithms.
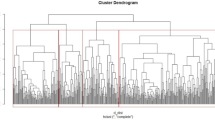
Extended Data Fig. 3 Orthogonal relationships among circulating KRIS proteins.
a, Hierarchical cluster analysis in the Joslin cohort with T1D, n = 219. b, Joslin cohort with T2D, n = 144. c, Spearman’s rank correlation matrix in the two Joslin cohorts, n = 363. a, b, Distances are shown (Ward’s method). TNF-RSF members are marked in red. c, Spearman’s correlation matrix represents relationships among proteins in the two cohorts in the analysis adjusted for the cohort indicator. Coefficients (r) are presented. Color intensity corresponds to the effect size (r).
Extended Data Fig. 4 KRIS proteins in the urine and development of ESRD.
Aptamer-based, urinary creatinine-adjusted KRIS protein profiles and development of ESRD in nested case–control studies selected from a, Joslin cohort with T1D (n = 60) and b, Joslin cohort with T2D (n = 52). Effect size (fold-change) is presented. Urinary proteins significant for corresponding thresholds used in multivariate screening for plasma proteomics study (two-sided) are marked with dark red bars.
Extended Data Fig. 5 Circulating KRIS proteins and renal mRNA expression of KRIS-encoding genes.
Plasma KRIS proteins and Affymetrix-based mRNA expression of the corresponding KRIS genes in kidney tissue derived from the Pima Indian Study (n = 56). Spearman’s rank correlation coefficients (r) are presented between glomerular and tubular gene expressions and circulating KRIS protein levels. The nominally significant correlations (two-sided P values) are marked with an asterisk. Shades of red, white and blue correspond to the magnitude of the effect size (r).
Extended Data Fig. 6 Renal mRNA expression of KRIS-encoding genes and histology of the diabetic kidney tissue.
Affymetrix-based mRNA expression data in kidney tissue specimens from subjects with T2D (1KGP). Glomerular (n = 23) and tubular (n = 37) expressions of candidate genes are correlated with histological indices of the diabetic kidney: glomerular sclerosis (Glom scl), tubulointerstitial fibrosis (Tub-int fibrosis) and lymphocytic infiltrate in the tubulointerstitium (Tub-int l-infiltr). Spearman’s rank correlation coefficients (r) are presented. The asterisk marks significant correlations (threshold for α = 0.05). Light-red fields mark proteins for which at least two coefficients were significant and positive.
Extended Data Fig. 7 Circulating KRIS proteins and prevalent diabetic eye complications.
Circulating KRIS proteins and prevalent PDR in the Joslin Kidney Study subjects with T1D (n = 180). OR and 95% CIs are adjusted for HbA1c, ACR and eGFR in the logistic analysis. Effect of KRIS proteins is per one tertile change; the two-sided P value was examined.
Supplementary information
Supplementary Information
Supplementary Methods and Supplementary Tables 1–8
Rights and permissions
About this article
Cite this article
Niewczas, M.A., Pavkov, M.E., Skupien, J. et al. A signature of circulating inflammatory proteins and development of end-stage renal disease in diabetes. Nat Med 25, 805–813 (2019). https://doi.org/10.1038/s41591-019-0415-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-019-0415-5
This article is cited by
-
Endothelial CXCR2 deficiency attenuates renal inflammation and glycocalyx shedding through NF-κB signaling in diabetic kidney disease
Cell Communication and Signaling (2024)
-
Circulating TNF receptor levels are associated with estimated glomerular filtration rate even in healthy individuals with normal kidney function
Scientific Reports (2024)
-
NF-κB in biology and targeted therapy: new insights and translational implications
Signal Transduction and Targeted Therapy (2024)
-
Albuminuria and Serum Tumor Necrosis Factor Receptor Levels in Patients with Type 2 Diabetes on SGLT2 Inhibitors: A Prospective Study
Diabetes Therapy (2024)
-
Mechanistic Insights and Potential Therapeutic Implications of NRF2 in Diabetic Encephalopathy
Molecular Neurobiology (2024)