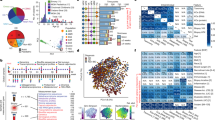
Abstract
Inflammatory bowel diseases (IBD) can be broadly divided into Crohn’s disease (CD) and ulcerative colitis (UC) from their clinical phenotypes. Over 150 host susceptibility genes have been described, although most overlap between CD, UC and their subtypes, and they do not adequately account for the overall incidence or the highly variable severity of disease. Replicating key findings between two long-term IBD cohorts, we have defined distinct networks of taxa associations within intestinal biopsies of CD and UC patients. Disturbances in an association network containing taxa of the Lachnospiraceae and Ruminococcaceae families, typically producing short chain fatty acids, characterize frequently relapsing disease and poor responses to treatment with anti-TNF-α therapeutic antibodies. Alterations of taxa within this network also characterize risk of later disease recurrence of patients in remission after the active inflamed segment of CD has been surgically removed.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All sequencing datasets from the current study have been deposited in a figshare repository and are publicly available. The Cohort 1 fasta file withthe mapping file are available at https://figshare.com/s/e9f2cffd0f0328ca5811 (https://doi.org/10.6084/m9.figshare.7335068) and the Cohort 2 fasta file with the mapping file are available at https://figshare.com/s/bbdd5dfb01e29484efa1 (https://doi.org/10.6084/m9.figshare.7335071). Associated codes for the analysis using R packages and QIIME can be found in these depositories. The genome-scale metabolic model script and dataset are available at https://figshare.com/s/a34f96698ca6fcd36ac2.
Change history
07 March 2019
Owing to an error during typesetting, a number of references were deleted from the Methods reference list. This altered all of the references in the Methods section and some of the references in Extended Data Fig. 5, making them inaccurate. References 121–134 were added back into the Methods reference list, and the references in the Methods section and in Extended Data Fig. 5 were renumbered accordingly. The error has been corrected in the PDF and HTML versions of this article.
References
Strober, W., Fuss, I. J. & Blumberg, R. S. The immunology of mucosal models of inflammation. Annu. Rev. Immunol. 20, 495–549 (2002).
Blumberg, R. & Powrie, F. Microbiota, disease, and back to health: a metastable journey. Sci Transl Med 4, 137rv137 (2012).
Hegazy, A. N. et al. Circulating and tissue-resident CD4(+) T cells with reactivity to intestinal microbiota are abundant in healthy individuals and function is altered during inflammation. Gastroenterology 153, 1320–1337 e1316 (2017).
Maloy, K. J. & Powrie, F. Intestinal homeostasis and its breakdown in inflammatory bowel disease. Nature 474, 298–306 (2011).
Rutgeerts, P. et al. Effect of faecal stream diversion on recurrence of Crohn’s disease in the neoterminal ileum. Lancet 338, 771–774 (1991).
Zachos, M., Tondeur, M. & Griffiths, A. M. Enteral nutritional therapy for induction of remission in Crohn’s disease.Cochrane Database Syst. Rev. 4, CD000542 (2007).
Cummings, J. H. & Kong, S. C. Probiotics, prebiotics and antibiotics in inflammatory bowel disease. Novartis. Found. Symp. 263, 99–111 (2004). discussion 111–114, 211–118.
Gecse, K. B. et al. A global consensus on the classification, diagnosis and multidisciplinary treatment of perianal fistulising Crohn’s disease. Gut 63, 1381–1392 (2014).
Faith, J. J. et al. The long-term stability of the human gut microbiota. Science 341, 1237439 (2013).
Arumugam, M. et al. Enterotypes of the human gut microbiome. Nature 473, 174–180 (2011).
Schloissnig, S. et al. Genomic variation landscape of the human gut microbiome. Nature 493, 45–50 (2013).
Flint, H. J., Duncan, S. H. & Louis, P. The impact of nutrition on intestinal bacterial communities. Curr. Opin. Microbiol. 38, 59–65 (2017).
Falony, G. et al. Population-level analysis of gut microbiome variation. Science 352, 560–564 (2016).
Vandeputte, D. et al. Stool consistency is strongly associated with gut microbiota richness and composition, enterotypes and bacterial growth rates. Gut 65, 57–62 (2016).
Jostins, L. et al. Host-microbe interactions have shaped the genetic architecture of inflammatory bowel disease. Nature 491, 119–124 (2012).
Zuk, O., Hechter, E., Sunyaev, S. R. & Lander, E. S. The mystery of missing heritability: genetic interactions create phantom heritability. Proc. Natl Acad. Sci. USA 109, 1193–1198 (2012).
Uhlig, H. H. Monogenic diseases associated with intestinal inflammation: implications for the understanding of inflammatory bowel disease. Gut 62, 1795–1805 (2013).
Lee, J. C. & Lennard-Jones, J. E. Inflammatory bowel disease in 67 families each with three or more affected first-degree relatives. Gastroenterology 111, 587–596 (1996).
Lee, J. C. et al. Gene expression profiling of CD8+ T cells predicts prognosis in patients with Crohn disease and ulcerative colitis. J. Clin. Invest. 121, 4170–4179 (2011).
Lee, J. C. et al. Genome-wide association study identifies distinct genetic contributions to prognosis and susceptibility in Crohn’s disease. Nat. Genet. 49, 262–268 (2017).
Park, S. H., Aniwan, S. & Loftus, E. V. Jr. Advances in the use of biologics and other novel drugs for managing inflammatory bowel disease. Curr. Opin. Pharmacol. 37, 65–71 (2017).
Pillai, N., Dusheiko, M., Burnand, B. & Pittet, V. A systematic review of cost-effectiveness studies comparing conventional, biological and surgical interventions for inflammatory bowel disease. PLoS ONE 12, e0185500 (2017).
Engels, M., Cross, R. K. & Long, M. D. Exercise in patients with inflammatory bowel diseases: current perspectives. Clin. Exp. Gastroenterol. 11, 1–11 (2018).
Eckburg, P. B. et al. Diversity of the human intestinal microbial flora. Science 308, 1635–1638 (2005).
Sartor, R. B. & Wu, G. D. Roles for intestinal bacteria, viruses, and fungi in pathogenesis of inflammatory bowel diseases and therapeutic approaches.Gastroenterology 152, 327–339.e4 (2017).
Png, C. W. et al. Mucolytic bacteria with increased prevalence in IBD mucosa augment in vitro utilization of mucin by other bacteria. Am. J. Gastroenterol. 105, 2420–2428 (2010).
Sokol, H. et al. Faecalibacterium prausnitzii is an anti-inflammatory commensal bacterium identified by gut microbiota analysis of Crohn disease patients. Proc. Natl Acad. Sci. USA 105, 16731–16736 (2008).
Magnusdottir, S. et al. Generation of genome-scale metabolic reconstructions for 773 members of the human gut microbiota. Nat. Biotechnol. 35, 81–89 (2017).
Fujimoto, T. et al. Decreased abundance of Faecalibacterium prausnitzii in the gut microbiota of Crohn’s disease. J. Gastroenterol. Hepatol. 28, 613–619 (2013).
Lopez-Siles, M. et al. Mucosa-associated Faecalibacterium prausnitzii phylotype richness is reduced in patients with inflammatory bowel disease. Appl. Environ. Microbiol. 81, 7582–7592 (2015).
Imhann, F. et al. Interplay of host genetics and gut microbiota underlying the onset and clinical presentation of inflammatory bowel disease. Gut 67, 108–119 (2018).
Goodman, A. L. & Gordon, J. I. Our unindicted coconspirators: human metabolism from a microbial perspective. Cell. Metab. 12, 111–116 (2010).
Muegge, B. D. et al. Diet drives convergence in gut microbiome functions across mammalian phylogeny and within humans. Science 332, 970–974 (2011).
Mainali, K. P. et al. Statistical analysis of co-occurrence patterns in microbial presence-absence datasets. PLoS ONE 12, e0187132 (2017).
Williams, R. J., Howe, A. & Hofmockel, K. S. Demonstrating microbial co-occurrence pattern analyses within and between ecosystems. Front. Microbiol. 5, 358 (2014).
Rahnavard, G. F. et al. HAllA: Hierarchical All-against-All significance testing. http://huttenhower.sph.harvard.edu/halla (2017).
Plevinsky, J. M., Wojtowicz, A. A., Poulopoulos, N., Schneider, K. L. & Greenley, R. N. Perceived impairment in sports participation in adolescents with inflammatory bowel diseases: a preliminary examination. J. Pediatr. Gastroenterol. Nutr. 66, 79–83 (2018).
Torres, J., Mehandru, S., Colombel, J. F. & Peyrin-Biroulet, L. Crohn’s disease. Lancet 389, 1741–1755 (2017).
Khanna, R. et al. Early combined immunosuppression for the management of Crohn’s disease (REACT): a cluster randomised controlled trial. Lancet 386, 1825–1834 (2015).
Ungaro, R., Mehandru, S., Allen, P. B., Peyrin-Biroulet, L. & Colombel, J. F. Ulcerative colitis. Lancet 389, 1756–1770 (2017).
Magnusson, M. K. et al. Anti-TNF therapy response in patients with ulcerative colitis is associated with colonic antimicrobial peptide expression and microbiota composition. J. Crohns Colitis 10, 943–952 (2016).
Ghouri, Y. A. et al. Systematic review of randomized controlled trials of probiotics, prebiotics, and synbiotics in inflammatory bowel disease. Clin. Exp. Gastroenterol. 7, 473–487 (2014).
Topping, D. L. & Clifton, P. M. Short-chain fatty acids and human colonic function: roles of resistant starch and nonstarch polysaccharides. Physiol. Rev. 81, 1031–1064 (2001).
Arpaia, N. et al. Metabolites produced by commensal bacteria promote peripheral regulatory T-cell generation. Nature 504, 451–455 (2013).
Wu, F. et al. Phascolarctobacterium faecium abundant colonization in human gastrointestinal tract. Exp. Ther. Med. 14, 3122–3126 (2017).
Chen, J. et al. Dysbiosis of intestinal microbiota and decrease in paneth cell antimicrobial peptide level during acute necrotizing pancreatitis in rats. PLoS ONE 12, e0176583 (2017).
Hancock, L. & Mortensen, N. J. How often do IBD patients require resection of their intestine? Inflamm. Bowel Dis. 14, S68–S69 (2008).
Olivera, P., Spinelli, A., Gower-Rousseau, C., Danese, S. & Peyrin-Biroulet, L. Surgical rates in the era of biological therapy: up, down or unchanged? Curr. Opin. Gastroenterol. 33, 246–253 (2017).
Rutgeerts, P. et al. Predictability of the postoperative course of Crohn’s disease. Gastroenterology 99, 956–963 (1990).
Patterson, A. M. et al. Human gut symbiont roseburia hominis promotes and regulates innate immunity. Front. Immunol. 8, 1166 (2017).
Machiels, K. et al. Specific members of the predominant gut microbiota predict pouchitis following colectomy and IPAA in UC. Gut 66, 79–88 (2017).
Machiels, K. et al. A decrease of the butyrate-producing species Roseburia hominis and Faecalibacterium prausnitzii defines dysbiosis in patients with ulcerative colitis. Gut 63, 1275–1283 (2014).
Flint, H. J., Duncan, S. H., Scott, K. P. & Louis, P. Links between diet, gut microbiota composition and gut metabolism. Proc. Nutr. Soc. 74, 13–22 (2015).
Byndloss, M. X. et al. Microbiota-activated PPAR-gamma signaling inhibits dysbiotic Enterobacteriaceae expansion. Science 357, 570–575 (2017).
Liguori, G. et al. Fungal dysbiosis in mucosa-associated microbiota of Crohn’s disease patients. J. Crohns Colitis 10, 296–305 (2016).
Papa, E. et al. Non-invasive mapping of the gastrointestinal microbiota identifies children with inflammatory bowel disease. PLoS ONE 7, e39242 (2012).
Santoru, M. L. et al. Cross-sectional evaluation of the gut-microbiome metabolome axis in an Italian cohort of IBD patients. Sci. Rep. 7, 9523 (2017).
Pedamallu, C. S. et al. Metagenomic characterization of microbial communities in situ within the deeper layers of the ileum in crohn’s disease. Cell. Mol. Gastroenterol. Hepatol. 2, 563–566 e565 (2016).
Kellermayer, R. et al. Microbiota separation and C-reactive protein elevation in treatment-naive pediatric granulomatous Crohn disease. J. Pediatr. Gastroenterol. Nutr. 55, 243–250 (2012).
Frank, D. N. et al. Molecular-phylogenetic characterization of microbial community imbalances in human inflammatory bowel diseases. Proc. Natl Acad. Sci. USA 104, 13780–13785 (2007).
Willing, B. P. et al. A pyrosequencing study in twins shows that gastrointestinal microbial profiles vary with inflammatory bowel disease phenotypes. Gastroenterology 139, 1844–1854 e1841 (2010).
Wang, W. et al. Increased proportions of Bifidobacterium and the Lactobacillus group and loss of butyrate-producing bacteria in inflammatory bowel disease. J. Clin. Microbiol. 52, 398–406 (2014).
Rajilic-Stojanovic, M., Shanahan, F., Guarner, F. & de Vos, W. M. Phylogenetic analysis of dysbiosis in ulcerative colitis during remission. Inflamm. Bowel Dis. 19, 481–488 (2013).
Seksik, P. et al. Alterations of the dominant faecal bacterial groups in patients with Crohn’s disease of the colon. Gut 52, 237–242 (2003).
Halfvarson, J. et al. Dynamics of the human gut microbiome in inflammatory bowel disease. Nat. Microbiol. 2, 17004 (2017).
Erickson, A. R. et al. Integrated metagenomics/metaproteomics reveals human host-microbiota signatures of Crohn’s disease. PLoS ONE 7, e49138 (2012).
Eun, C. S. et al. Does the intestinal microbial community of Korean Crohn’s disease patients differ from that of western patients? BMC. Gastroenterol. 16, 28 (2016).
Forbes, J. D., Van Domselaar, G. & Bernstein, C. N. Microbiome survey of the inflamed and noninflamed gut at different compartments within the gastrointestinal tract of inflammatory bowel disease patients. Inflamm. Bowel Dis. 22, 817–825 (2016).
Andoh, A. et al. Multicenter analysis of fecal microbiota profiles in Japanese patients with Crohn’s disease. J. Gastroenterol. 47, 1298–1307 (2012).
Rehman, A. et al. Transcriptional activity of the dominant gut mucosal microbiota in chronic inflammatory bowel disease patients. J. Med. Microbiol. 59, 1114–1122 (2010).
Martinez-Medina, M., Aldeguer, X., Gonzalez-Huix, F., Acero, D. & Garcia-Gil, L. J. Abnormal microbiota composition in the ileocolonic mucosa of Crohn’s disease patients as revealed by polymerase chain reaction-denaturing gradient gel electrophoresis. Inflamm. Bowel Dis. 12, 1136–1145 (2006).
Norman, J. M. et al. Disease-specific alterations in the enteric virome in inflammatory bowel disease. Cell 160, 447–460 (2015).
Ashton, J. J. et al. 16S sequencing and functional analysis of the fecal microbiome during treatment of newly diagnosed pediatric inflammatory bowel disease. Medicine. 96, e7347 (2017).
Kabeerdoss, J., Jayakanthan, P., Pugazhendhi, S. & Ramakrishna, B. S. Alterations of mucosal microbiota in the colon of patients with inflammatory bowel disease revealed by real time polymerase chain reaction amplification of 16 S ribosomal ribonucleic acid. Indian J. Med. Res. 142, 23–32 (2015).
Kolho, K. L. et al. Fecal microbiota in pediatric inflammatory bowel disease and its relation to inflammation. Am. J. Gastroenterol. 110, 921–930 (2015).
Martinez-Medina, M. et al. Molecular diversity of Escherichia coli in the human gut: new ecological evidence supporting the role of adherent-invasive E. coli (AIEC) in Crohn’s disease. Inflamm. Bowel Dis. 15, 872–882 (2009).
Duboc, H. et al. Connecting dysbiosis, bile-acid dysmetabolism and gut inflammation in inflammatory bowel diseases. Gut 62, 531–539 (2013).
Schwiertz, A. et al. Microbiota in pediatric inflammatory bowel disease. J. Pediatr. 157, 240–244 e241 (2010).
Kaakoush, N. O. et al. Microbial dysbiosis in pediatric patients with Crohn’s disease. J. Clin. Microbiol. 50, 3258–3266 (2012).
Walker, A. W. et al. High-throughput clone library analysis of the mucosa-associated microbiota reveals dysbiosis and differences between inflamed and non-inflamed regions of the intestine in inflammatory bowel disease. BMC. Microbiol. 11, 7 (2011).
Gophna, U., Sommerfeld, K., Gophna, S., Doolittle, W. F. & van Zanten, S. J. O. V. Differences between tissue-associated intestinal microfloras of patients with Crohn’s disease and ulcerative colitis. J. Clin. Microbiol. 44, 4136–4141 (2006).
Knoll, R. L. et al. Gut microbiota differs between children with Inflammatory Bowel Disease and healthy siblings in taxonomic and functional composition: a metagenomic analysis. Am. J. Physiol-Gastr. L 312, G327–G339 (2017).
Hansen, R. et al. Microbiota of de-novo pediatric IBD: increased Faecalibacterium prausnitzii and reduced bacterial diversity in Crohn’s but not in ulcerative colitis. Am. J. Gastr. 107, 1913–1922 (2012).
Jacobs, J. P. et al. A disease-associated microbial and metabolomics state in relatives of pediatric inflammatory bowel disease patients. Cell. Mol. Gastroenterol. Hepatol. 2, 750–766 (2016).
Bajer, L. et al. Distinct gut microbiota profiles in patients with primary sclerosing cholangitis and ulcerative colitis. World J. Gastr. 23, 4548–4558 (2017).
Baumgart, M. et al. Culture independent analysis of ileal mucosa reveals a selective increase in invasive Escherichia coli of novel phylogeny relative to depletion of Clostridiales in Crohn’s disease involving the ileum. ISME. J. 1, 403–418 (2007).
Gevers, D. et al. The treatment-naive microbiome in new-onset Crohn’s disease. Cell. Host. Microbe. 15, 382–392 (2014).
Tong, M. et al. A modular organization of the human intestinal mucosal microbiota and its association with inflammatory bowel disease. PLoS ONE 8, e80702 (2013).
Pascal, V. et al. A microbial signature for Crohn’s disease. Gut 66, 813–822 (2017).
Mottawea, W. et al. Altered intestinal microbiota-host mitochondria crosstalk in new onset Crohn’s disease. Nat. Commun. 7, 13419 (2016).
Morgan, X. C. et al. Dysfunction of the intestinal microbiome in inflammatory bowel disease and treatment. Genome. Biol. 13, R79 (2012).
Verma, R., Verma, A. K., Ahuja, V. & Paul, J. Real-time analysis of mucosal flora in patients with inflammatory bowel disease in India. J. Clin. Microbiol. 48, 4279–4282 (2010).
Arun Gupta, S. K., Wagner, Josef, Kirkwood, Carl, Morrison, Mark & McSweeney, Chris and finlay macrae. analysis of mucosal microbiota in in ammatory bowel disease using a custom phylogenetic microarray. Austin J. Gastr. 1, 1–6 (2014).
Bibiloni, R., Mangold, M., Madsen, K. L., Fedorak, R. N. & Tannock, G. W. The bacteriology of biopsies differs between newly diagnosed, untreated, Crohn’s disease and ulcerative colitis patients. J. Med. Microbiol. 55, 1141–1149 (2006).
Sokol, H. et al. Low counts of Faecalibacterium prausnitzii in colitis microbiota. Inflamm. Bowel Dis. 15, 1183–1189 (2009).
Andoh, A. et al. Comparison of the fecal microbiota profiles between ulcerative colitis and Crohn’s disease using terminal restriction fragment length polymorphism analysis. J. Gastroenterol. 46, 479–486 (2011).
Swidsinski, A., Loening-Baucke, V., Vaneechoutte, M. & Doerffel, Y. Active Crohn’s disease and ulcerative colitis can be specifically diagnosed and monitored based on the biostructure of the fecal flora. Inflamm. Bowel Dis. 14, 147–161 (2008).
Nishino, K. et al. Analysis of endoscopic brush samples identified mucosa-associated dysbiosis in inflammatory bowel disease. J. Gastroenterol. 53, 95–106 (2017).
Chen, L. et al. Characteristics of fecal and mucosa-associated microbiota in Chinese patients with inflammatory bowel disease. Medicine (Baltimore) 93, e51 (2014).
Suchodolski, J. S., Dowd, S. E., Wilke, V., Steiner, J. M. & Jergens, A. E. 16S rRNA gene pyrosequencing reveals bacterial dysbiosis in the duodenum of dogs with idiopathic inflammatory bowel disease. PLoS ONE 7, e39333 (2012).
Xenoulis, P. G. et al. Molecular-phylogenetic characterization of microbial communities imbalances in the small intestine of dogs with inflammatory bowel disease. FEMS Microbiol. Ecol. 66, 579–589 (2008).
Omori, M. et al. Fecal microbiome in dogs with inflammatory bowel disease and intestinal lymphoma. J. Vet. Med. Sci. 79, 1840–1847 (2017).
Allenspach, K. et al. Evaluation of mucosal bacteria and histopathology, clinical disease activity and expression of Toll-like receptors in German shepherd dogs with chronic enteropathies. Vet. Microbiol. 146, 326–335 (2010).
Suchodolski, J. S., Xenoulis, P. G., Paddock, C. G., Steiner, J. M. & Jergens, A. E. Molecular analysis of the bacterial microbiota in duodenal biopsies from dogs with idiopathic inflammatory bowel disease. Vet. Microbiol. 142, 394–400 (2010).
Suchodolski, J. S. et al. The fecal microbiome in dogs with acute diarrhea and idiopathic inflammatory bowel disease. PLoS ONE 7, e51907 (2012).
Rossi, G. et al. Comparison of microbiological, histological, and immunomodulatory parameters in response to treatment with either combination therapy with prednisone and metronidazole or probiotic VSL#3 strains in dogs with idiopathic inflammatory bowel disease. PLoS ONE 9, e94699 (2014).
Minamoto, Y. et al. Alteration of the fecal microbiota and serum metabolite profiles in dogs with idiopathic inflammatory bowel disease. Gut Microbes 6, 33–47 (2015).
Jergens, A. E. Feline idiopathic inflammatory bowel disease: what we know and what remains to be unraveled. J. Feline. Med. Surg. 14, 445–458 (2012).
Vazquez-Baeza, Y., Hyde, E. R., Suchodolski, J. S. & Knight, R. Dog and human inflammatory bowel disease rely on overlapping yet distinct dysbiosis networks. Nat. Microbiol. 1, 16177 (2016).
Inness, V. L., McCartney, A. L., Khoo, C., Gross, K. L. & Gibson, G. R. Molecular characterisation of the gut microflora of healthy and inflammatory bowel disease cats using fluorescence in situ hybridisation with special reference to Desulfovibrio spp. J. Anim. Physiol. Anim. Nutr. 91, 48–53 (2007).
Elinav, E. et al. NLRP6 inflammasome regulates colonic microbial ecology and risk for colitis. Cell 145, 745–757 (2011).
Li, M. et al. Upregulation of intestinal barrier function in mice with DSS-induced colitis by a defined bacterial consortium is associated with expansion of IL-17A producing gamma delta T cells. Front. Immunol. 8, 824 (2017).
Rooks, M. G. et al. Gut microbiome composition and function in experimental colitis during active disease and treatment-induced remission. ISME. J. 8, 1403–1417 (2014).
Simpson, K. W. et al. Adherent and invasive Escherichia coli is associated with granulomatous colitis in boxer dogs. Infect. Immun. 74, 4778–4792 (2006).
Janeczko, S. et al. The relationship of mucosal bacteria to duodenal histopathology, cytokine mRNA, and clinical disease activity in cats with inflammatory bowel disease. Vet. Microbiol. 128, 178–193 (2008).
Larmonier, C. B. et al. Reduced colonic microbial diversity is associated with colitis in NHE3-deficient mice. Am. J. Physiol. Gastrointest. Liver Physiol. 305, G667–G677 (2013).
Spalinger, M. R. et al. PTPN2 controls differentiation of CD4(+) T cells and limits intestinal inflammation and intestinal dysbiosis. Mucosal Immunol 8, 918–929 (2015).
Munyaka, P. M., Rabbi, M. F., Khafipour, E. & Ghia, J. E. Acute dextran sulfate sodium (DSS)-induced colitis promotes gut microbial dysbiosis in mice. J. Basic Microbiol. 56, 986–998 (2016).
Okayasu, I. et al. A novel method in the induction of reliable experimental acute and chronic ulcerative colitis in mice. Gastroenterology 98, 694–702 (1990).
Robinson, A. M. et al. Alterations of colonic function in the Winnie mouse model of spontaneous chronic colitis. Am. J. Physiol. Gastrointest. Liver Physiol. 312, G85–G102 (2017).
Pittet, V. et al. Cohort profile: the Swiss Inflammatory Bowel Disease Cohort Study (SIBDCS). Int. J. Epidemiol. 38, 922–931 (2009).
Lennard-Jones, J. E. Classification of inflammatory bowel disease. Scand. J. Gastroenterol. Suppl. 170, 2–6, discussion 16–19 (1989).
Harris, P. A. et al. Research Electronic Data Capture (REDCap)—a metadata-driven methodology and workflow process for providing translational research informatics support. J. Biomed. Inform. 42, 377–381 (2009).
Sundquist, A. et al. Bacterial flora-typing with targeted, chip-based Pyrosequencing. BMC Microbiol. 7, 108 (2007).
Yilmaz, B. et al. The presence of genetic risk variants within PTPN2 and PTPN22 is associated with intestinal microbiota alterations in Swiss IBD cohort patients. PLoS One 13, e0199664 (2018).
Whiteley, A. S. et al. Microbial 16S rRNA Ion Tag and community metagenome sequencing using the Ion Torrent (PGM) Platform. J. Microbiol. Methods 91, 80–88 (2012).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336 (2010).
McMurdie, P. J. & Holmes, S. phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS One 8, e61217 (2013).
Callahan, B. J., Sankaran, K., Fukuyama, J. A., McMurdie, P. J. & Holmes, S. P. Bioconductor Workflow for Microbiome Data Analysis: from raw reads to community analyses. F1000Res 5, 1492 (2016).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Good, I. J. The population frequencies of species and the estimation of population parameters. Biometrika 40, 237–264 (1953).
Su, G., Kuchinsky, A., Morris, J. H., States, D. J. & Meng, F. GLay: community structure analysis of biological networks. Bioinformatics 26, 3135–3137 (2010).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate - a practical and powerful approach to multiple testing. J. R. Stat. Soc. Series B Stat. Methodol. 57, 289–300 (1995).
Yang, I. et al. Intestinal microbiota composition of interleukin-10 deficient C57BL/6J mice and susceptibility to Helicobacter hepaticus-induced colitis. PLoS ONE 8, e70783 (2013).
Ren, Y. et al. Polysaccharide of Hericium erinaceus attenuates colitis in C57BL/6 mice via regulation of oxidative stress, inflammation-related signaling pathways and modulating the composition of the gut microbiota. J. Nutr. Biochem. 57, 67–76 (2018).
Nunberg, M. et al. Interleukin 1alpha-deficient mice have an altered gut microbiota leading to protection from dextran sodium sulfate-induced colitis. mSystems 3, e00213–17 (2018).
Schaubeck, M. et al. Dysbiotic gut microbiota causes transmissible Crohn’s disease-like ileitis independent of failure in antimicrobial defence. Gut 65, 225–237 (2016).
Lupp, C. et al. Host-mediated inflammation disrupts the intestinal microbiota and promotes the overgrowth of Enterobacteriaceae. Cell. Host. Microbe. 2, 204 (2007).
Nones, K. et al. Multidrug resistance gene deficient (mdr1a−/−) mice have an altered caecal microbiota that precedes the onset of intestinal inflammation. J. Appl. Microbiol. 107, 557–566 (2009).
Manchester, A. C. et al. Association between granulomatous colitis in french bulldogs and invasive Escherichia coli and response to fluoroquinolone antimicrobials. J. Vet. Intern. Med. 27, 56–61 (2013).
Roy, U. et al. Distinct microbial communities trigger colitis development upon intestinal barrier damage via innate or adaptive immune cells. Cell Rep 21, 994–1008 (2017).
Johnston, D. G. W. et al. Loss of microRNA-21 influences the gut microbiota causing reduced susceptibility in a murine model of colitis. J. Crohns Colitis 12, 835–848 (2018).
Berry, D. et al. Phylotype-level 16S rRNA analysis reveals new bacterial indicators of health state in acute murine colitis. ISME. J. 6, 2091–2106 (2012).
Zhang, Q. et al. Accelerated dysbiosis of gut microbiota during aggravation of DSS-induced colitis by a butyrate-producing bacterium. Sci. Rep. 6, 27572 (2016).
Schwab, C. et al. Longitudinal study of murine microbiota activity and interactions with the host during acute inflammation and recovery. ISME. J. 8, 1101–1114 (2014).
Zimmermann, J. et al. The intestinal microbiota determines the colitis-inducing potential of T-bet-deficient Th cells in mice. Eur. J. Immunol. 48, 161–167 (2018).
Garrett, W. S. et al. Enterobacteriaceae act in concert with the gut microbiota to induce spontaneous and maternally transmitted colitis. Cell. Host. Microbe. 8, 292–300 (2010).
Palm, N. W. et al. Immunoglobulin A coating identifies colitogenic bacteria in inflammatory bowel disease. Cell 158, 1000–1010 (2014).
Maharshak, N. et al. Altered enteric microbiota ecology in interleukin 10-deficient mice during development and progression of intestinal inflammation. Gut Microbes 4, 316–324 (2013).
Chassaing, B. et al. Dietary emulsifiers impact the mouse gut microbiota promoting colitis and metabolic syndrome. Nature 519, 92–96 (2015).
Osaka, T. et al. Meta-analysis of fecal microbiota and metabolites in experimental colitic mice during the inflammatory and healing phases. Nutrients 9, 1329 (2017).
Perez-Munoz, M. E. et al. Discordance between changes in the gut microbiota and pathogenicity in a mouse model of spontaneous colitis. Gut Microbes 5, 286–295 (2014).
Nagalingam, N. A., Kao, J. Y. & Young, V. B. Microbial ecology of the murine gut associated with the development of dextran sodium sulfate-induced colitis. Inflamm. Bowel Dis. 17, 917–926 (2011).
Alkadhi, S., Kunde, D., Cheluvappa, R., Randall-Demllo, S. & Eri, R. The murine appendiceal microbiome is altered in spontaneous colitis and its pathological progression. Gut Pathog. 6, 25 (2014).
Carvalho, F. A. et al. Interleukin-1beta (IL-1beta) promotes susceptibility of Toll-like receptor 5 (TLR5) deficient mice to colitis. Gut. 61, 373–384 (2012).
Seregin, S. S. et al. NLRP6 protects Il10(−/−) mice from colitis by limiting colonization of Akkermansia muciniphila. Cell Rep. 19, 2174 (2017).
Selvanantham, T. et al. NKT cell-deficient mice harbor an altered microbiota that fuels intestinal inflammation during chemically induced colitis. J. Immunol. 197, 4464–4472 (2016).
Lamas, B. et al. CARD9 impacts colitis by altering gut microbiota metabolism of tryptophan into aryl hydrocarbon receptor ligands. Nat. Med. 22, 598 (2016).
Madsen, K. L. et al. Antibiotic therapy attenuates colitis in interleukin 10 gene-deficient mice. Gastroenterology 118, 1094–1105 (2000).
Mitchell, J. et al. Colonic inhibition of phosphatase and tensin homolog increases colitogenic bacteria, causing development of colitis in Il10−/− mice. Inflamm. Bowel Dis. 24, 1718–1732 (2018).
Hakansson, A. et al. Immunological alteration and changes of gut microbiota after dextran sulfate sodium (DSS) administration in mice. Clin. Exp. Med. 15, 107–120 (2015).
Heimesaat, M. M. et al. Shift towards pro-inflammatory intestinal bacteria aggravates acute murine colitis via Toll-like receptors 2 and 4. PLoS ONE 2, e662 (2007).
Wohlgemuth, S., Haller, D., Blaut, M. & Loh, G. Reduced microbial diversity and high numbers of one single Escherichia coli strain in the intestine of colitic mice. Environ. Microbiol. 11, 1562–1571 (2009).
Arthur, J. C. et al. Intestinal inflammation targets cancer-inducing activity of the microbiota. Science 338, 120–123 (2012).
Schuppler, M., Lotzsch, K., Waidmann, M. & Autenrieth, I. B. An abundance of Escherichia coli is harbored by the mucosa-associated bacterial flora of interleukin-2-deficient mice. Infect. Immun. 72, 1983–1990 (2004).
Hoentjen, F. et al. Antibiotics with a selective aerobic or anaerobic spectrum have different therapeutic activities in various regions of the colon in interleukin 10 gene deficient mice. Gut. 52, 1721–1727 (2003).
Zenewicz, L. A. et al. IL-22 deficiency alters colonic microbiota to be transmissible and colitogenic. J. Immunol. 190, 5306–5312 (2013).
Dennis, K. L. et al. Adenomatous polyps are driven by microbe-instigated focal inflammation and are controlled by IL-10-producing T cells. Cancer Res. 73, 5905–5913 (2013).
Abecia, L. H. L., Khoo, C., Frantz, N. & McCartney, A. Effects of a novel galactooligosaccharide on the faecal microbiota of healthy and inflammatory bowel disease. Int. J. Probiotic. Prebiotics 5, 61–68 (2010).
Xu, J. et al. Does canine inflammatory bowel disease influence gut microbial profile and host metabolism? BMC Vet. Res. 12, 114 (2016).
Jones-Hall, Y. L., Kozik, A. & Nakatsu, C. Ablation of tumor necrosis factor is associated with decreased inflammation and alterations of the microbiota in a mouse model of inflammatory bowel disease. PLoS ONE 10, e0119441 (2015).
Bel, S. et al. Reprogrammed and transmissible intestinal microbiota confer diminished susceptibility to induced colitis in TMF−/− mice. Proc. Natl Acad. Sci. USA 111, 4964–4969 (2014).
Bloom, S. M. et al. Commensal Bacteroides species induce colitis in host-genotype-specific fashion in a mouse model of inflammatory bowel disease. Cell. Host. Microbe. 9, 390–403 (2011).
He, Q. et al. Dysbiosis of the fecal microbiota in the TNBS-induced Crohn’s disease mouse model. Appl. Microbiol. Biotechnol. 100, 4485–4494 (2016).
Couturier-Maillard, A. et al. NOD2-mediated dysbiosis predisposes mice to transmissible colitis and colorectal cancer. J. Clin. Invest. 123, 700–711 (2013).
Ettreiki, C. et al. Juvenile ferric iron prevents microbiota dysbiosis and colitis in adult rodents. W. J. Gastr. 18, 2619–2629 (2012).
Devkota, S. et al. Dietary-fat-induced taurocholic acid promotes pathobiont expansion and colitis in Il10−/− mice. Nature 487, 104–108 (2012).
Zhang, Z. et al. Chlorogenic acid ameliorates experimental colitis by promoting growth of akkermansia in mice. Nutrients 9, 677 (2017).
Ghosh, S., Molcan, E., DeCoffe, D., Dai, C. & Gibson, D. L. Diets rich in n-6 PUFA induce intestinal microbial dysbiosis in aged mice. Br. J. Nutr. 110, 515–523 (2013).
Ye, J. et al. Bacteria and bacterial rRNA genes associated with the development of colitis in IL-10(−/−) mice. Inflamm. Bowel Dis. 14, 1041–1050 (2008).
Vijay-Kumar, M. et al. Metabolic syndrome and altered gut microbiota in mice lacking Toll-like receptor 5. Science 328, 228–231 (2010).
Zhu, W. et al. Precision editing of the gut microbiota ameliorates colitis. Nature 553, 208–211 (2018).
Moon, C., Stupp, G. S., Su, A. I. & Wolan, D. W. Metaproteomics of colonic microbiota unveils discrete protein functions among colitic mice and control groups.Proteomics 18, 1700391 (2018).
Du, Z. et al. Development of gut inflammation in mice colonized with mucosa-associated bacteria from patients with ulcerative colitis. Gut Pathog. 7, 32 (2015).
Bibiloni, R., Simon, M. A., Albright, C., Sartor, B. & Tannock, G. W. Analysis of the large bowel microbiota of colitic mice using PCR/DGGE. Lett. Appl. Microbiol. 41, 45–51 (2005).
Kudelka, M. R. et al. Cosmc is an X-linked inflammatory bowel disease risk gene that spatially regulates gut microbiota and contributes to sex-specific risk. Proc. Natl Acad. Sci. USA 113, 14787–14792 (2016).
Vereecke, L. et al. A20 controls intestinal homeostasis through cell-specific activities. Nat. Commun. 5, 5103 (2014).
Acknowledgements
We thank all patients and the members of the Swiss IBD cohort and Bern cohort for their commitment. We also thank the staff of the University Hospital of Bern, Clinic of Visceral Medicine and Surgery, and the Bern City Hospitals led by F. Seibold and R. Tutuian for obtaining samples in Cohort 2. This research was supported by Systems X (GutX) to A.J.M. and J.S., and the Swiss IBD cohort (grant no. 33CS30-148422) to G.R., A.J.M and C.M. The founding institutions had no role in the study design, analysis or interpretation of the results. We thank G. Rahnavard and C. Huttenhower (Department of Biostatistics, Harvard T.H. Chan School of Public Health, USA) for their help in using the HAllA pipeline. We also thank J. Harrell Rieder, A. Suter, S. Brand, C. Mooser, W. Kwong Chung and J. Hugenschmidt for helping B.Y. during the process of sample preparation. We also thank G. Weingart (Department of Biostatistics, Harvard T.H. Chan School of Public Health, USA) for his enormous help in the optimization of MaAsLin running on the MacOS platform using R.
Author information
Authors and Affiliations
Consortia
Contributions
A.J.M. conceived, designed and supervised the study. B.Y. performed all the experiments, analyzed the data and wrote the manuscript with A.J.M. P.J. organized and collected the samples of the second cohort. P.J. F.D.B., Y.F., N.F. and M.G. were involved in data curation. O.O. and C.R. carried out metabolic reaction analysis, and J.S. supervised these analyses. A.J.M., P.J. P.M., C.M., V.E.H.P, M.H.M., G.R., R.W. and Swiss IBD cohort investigators acquired patient samples and detailed structured clinical phenotypes.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Unique microbial taxa identified as IBD signatures using unsupervised meta-analysis of published human IBD studies.
a, The heatmap shows significant microbial changes in CD, UC or IBD (CD + UC were combined as IBD) disease groups compared to non-IBD subjects. Each disease group (CD, UC or CD + UC) was compared independently to non-IBD and each color code reports the direction of microbial changes in each respective disease status against non-IBD subjects. Euclidean clustering was performed for sample annotations (vertical) including race/ethnicity, gender, median age, patient number, sample type, sequencing method and microbial taxa (horizontal) at different taxonomic ranks. Taxa in black bold label with asterisk demonstrate those findings verified by a subset of the findings in Fig. 1. Taxa in gray bold label with gray asterisk demonstrate findings in some of the studies verified by of the findings in Fig. 1 specifically, Lachnospiraceae family, Lachnospira, Coprococcus, Clostridiales order, Faecalibacterium, Ruminococcus, Roseburia and Ruminococcaceae family for Group 1, and Actinobacteria, Proteobacteria phyla and Enterobacteriaceae family for Group 2. 18 significant additional replicated taxa in the heterogeneous Group 3 are Bacteroidetes phylum and genera from this phylum including Bacteroides, Odoribacter, Butyricimonas, Parabacteroides, Sutterella, Prevotella (of Prevotellaceae), Prevotella (of Paraprevotellaceae) and Rikenellaceae family; also Firmicutes and genera from this phylum including Phascolarctobacterium, Dialister, Eubacterium∙∙∙ and Ruminococcus∙∙∙; Blautia, Collinsella and Bifidobacterium; Sutturella from Proteobacteria and Tenericutes. Underlined taxa are matching with Cluster CDA in Fig. 2. b, The heatmap (studies are in order as in a shows clinically relevant information collected through the studies analyzed in a. Recorded clinical phenotyping information in a given study is shown in green color (Identified) and the lack of clinical data is represented in white color (non-identified).
Extended Data Fig. 2 Microbial taxa comparison of CD with UC in published IBD studies.
The heatmap shows the comparison of microbial changes between CD and UC. Taxa higher in CD (lower in UC) are in red, while taxa higher in UC (lower CD) are in blue. Euclidean clustering was performed for sample annotations (vertical) including race/ethnicity, gender, median age, patient number, sample type and sequencing method and microbial taxa (horizontal) at different taxonomic ranks. Taxa names in bold demonstrate those findings verified by a subset of the findings in Fig. 1 and taxa names in gray bold shows the findings that partially match with our findings in Fig. 1.
Extended Data Fig. 3 The dominant bacterial phyla along the gastrointestinal tract of IBD patients and dysbiosis in IBD patients.
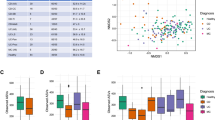
a,b, Distribution of predominant bacterial phylotypes along the cephalocaudal axis of the gut in CD, UC and non-IBD subjects of Cohort 1 (a) and Cohort 2 (b) are depicted after stratification according to the relative abundance of Firmicutes at each sampling site. The dominant bacterial phylotypes are Bacteroidetes (51.6% IBD Cohort 1; 56% IBD Cohort 2 and 56% non-IBD), Firmicutes (34.9, 29 and 25.7%, respectively) and Proteobacteria (9.1, 17 and 14%); with a smaller proportion of Fusobacteria (0.8, 0.4 and 1.1%), Actinobacteria (0.79, 0.46 and 0.33%) and Tenericutes (0.2, 0.04 and 0.09%). c–f, Microbial composition differences between IBD patients and non-IBD subjects were identified by species richness (Observed OTUs, Shannon and Simpson indices) in Cohort 1 (c) and Cohort 2 (d) and microbiome clustering based on unweighted and weighted UniFrac PCoA metrics for Cohort 1 (e) and Cohort 2 (f). Box-and-whisker plots in c and d display first and third quartiles and whiskers are from each quartile to the minimum or maximum. g,h, Beta dispersion statistics were performed by analyzing the sampling distance to centroids for Cohort 1 (g) and Cohort 2 (h) and there is no significant differences between compared groups in g and h. i,j, Only significant taxa associated with CD or UC shown as relative abundance ratio in Cohort 1 (i) or Cohort 2 (j) were identified using MaAsLin pipeline with BH-FDR correction (q value). q < 0.05 was considered significant. Significant differences were determined by either non-parametric two-sided Mann–Whitney U-test (c,d,g,h) or Adonis test for multiple comparisons (e,f) and P < 0.05 was considered significant. Box-and-whisker plots in c,d,g,h display first and third quartiles and whiskers are from each quartile to the minimum or maximum. 494 CD and 447 UC samples in Cohort 1 and 230 CD, 195 UC and 770 non-IBD samples in Cohort 2 were used for analysis (a–j).
Extended Data Fig. 4 Microbial taxa and functional metabolic subsystem differences in IBD patients.
a,b, Significant taxonomic differences are depicted as relative abundance ratios between CD and non-IBD samples (a) and between UC and non-IBD samples (b) identified in MaAsLin pipeline with BH-FDR correction. q < 0.05 was considered significant. c,d, Relative abundance of the most important matching OTUs were identified using machine learning algorithm and are depicted for Cohort 1 (c) and Cohort 2 (d) using notched box whisker showing first and third quartiles with median value. Each dot represents a single sample (c,d). e,f,h,i, After mapping OTUs to metabolic reactions, calculated the metabolic distances between all pairs of patients based on raw reaction counts are shown on PCA plots based on L2 distance of total reaction counts between UC and CD and boxplots show the respective coefficients PC1 and PC2 axis in Cohort 1 (e,f) and Cohort 2 (h,i). PC1 and PC2 are the first two principal components. g,j, The principal component analysis (PCA) analysis illustrates robust data at the metabolic reaction level: (∼60% variance explained by PC1/2 Blue indicates UC and red indicates CD patients and are shown using notched box whisker plots showing first and third quartiles with median value. Metabolic subsystems different between in CD and UC patients were identified in Cohort 1 (g) and in Cohort 2 (j). Similar significant metabolic pathway enrichment was detected in both cohorts. Red color represents the enrichment in CD and blue color represents the enrichment in UC. Box-and-whisker plots in g,j display first and third quartiles and whiskers are from each quartile to the minimum or maximum and possible outliers. Consistent metabolic subsystems increased in CD belonged to B-vitamin and LPS biosynthesis, heparan sulfate and chondroitin sulfate degradation and fatty acid oxidation. The BH-FDR was applied to correct for multiple testing and q < 0.05 between groups was considered significant (f,i). Fisher’s exact test was performed to determine if the subsystem was overrepresented among the significantly different reactions; subsystems with P < 0.05 were considered enriched (g,j). 494 CD and 447 UC samples in Cohort 1 and 230 CD, 195 UC and 770 non-IBD samples in Cohort 2 were used for analysis (a–d).
Extended Data Fig. 5 Unique microbial taxa identified as IBD signatures across different species using unsupervised meta-analysis of the published IBD studies.
a, The Euclidean clustering of IBD patients and animal models of IBD including dogs/cats diagnosed with IBD and mice with genetically and/or chemically (DSS or TNBS) induced colitis was performed using information of taxa identified significantly changing between disease groups and is plotted using the categorical information of the study models such as according to race/ethnicity, gender, median age, species, subject number, sample type, sample size, sequencing method and experimental model of IBD induction. b, The Spearman correlation heatmap shows the correlation between 123 different human and animal IBD studies based on identified 96 differentially abundant microbial taxa that are characterized in a. The correlation values ranging from 0 to 1 show positive correlation (in red) and the values ranging from −1 to 0 show negative correlation (in blue) between compared IBD studies. c, Statistical information of data of 123 independent human and animal IBD studies in total (a,b) is shown on the same matrix. Non-parametric two-tailed Spearman correlation test was performed and P < 0.05 was considered significant. Green color shows significant correlation between taxa plotted in a.
Extended Data Fig. 6 Microbial stability over time in longitudinally studied IBD patients and correlation of intestinal inflammation with microbial abundance.
a, Biopsies were collected from 22 individuals in Cohort 1 and 12 individuals in Cohort 2 over several years (1–9 years and 0.25–2 years, respectively). Each row corresponds to the time course of an individual patient. The resulting data comprised 176 biopsy samples. b, PCoA on Bray–Curtis dissimilarity distance matrix for longitudinally collected (as shown in a) 77 CD and 44 UC samples in Cohort 1 and 49 CD and 14 UC samples in Cohort 2 are plotted. Each color in represents an individual IBD patient. Ellipsoids represent a 95% confidence interval surrounding each disease group. c, The relative abundance changes for Bacteroides, Firmicutes and Proteobacteria phyla in IBD patients are plotted based on their disease severity changes over time. The x axis shows the relative abundance difference of a given phylum compared to previous sampling time point. A value higher than zero indicates that the phylum increases and a value lower than zero indicates that the phylum decreases compared to the previous sampling time point. Disease severity worsening over time is labeled as ‘decreasing’, improving over time is labeled as ‘increasing’ and stable disease severity is identified as ‘steady’ on the y axis. d,e, Fecal calprotectin that is positively correlated with Enterobacteriaceae∙ and Klebsiella for 79 CD patients (d) and negatively correlated with Ruminococcus∙∙∙ and Prevotella in 42 UC patients are shown on continuous data plot generated in MaAsLin pipeline with q < 0.05 (e). Spearman’s rank correlation coefficient for taxa in CD: 0.284 and 0.147 and for taxa in UC: −0.322 and −0.2, respectively. Adonis test was used to determine significant differences between the distance matrix of each group (b). Data shown in b was not significant when longitudinal samples were compared for individuals (P > 0.05). However, significant microbial differences were only observed between patients (P < 0.05). Taxa significantly associated with disease severity and fecal calprotectin were identified in MaAsLin pipeline (c,d,e) with BH-FDR correction and significant taxa are plotted. The q < 0.05 was considered significant.
Extended Data Fig. 7 Microbial profile along the gut in IBD patients.
a,b, Species richness of samples collected along the gut including Ileum (I), right colon (RC), transverse colon (CT), left colon (CL) and rectum (R) were calculated with Shannon index for 494 CD and 447 UC samples shown in Cohort 1 (a) and 230 CD and 195 UC in Cohort 2 (b). c,d, Beta diversity of these samples are shown for Cohort 1 (c) and Cohort 2 (d). e,f, Samples collected from same patients cluster intra-individually, as depicted CD (e) and UC (f) patients. g–j, Species richness calculated with Shannon and Simpson indices for 494 CD and 447 UC samples shown in Cohort 1 and 230 CD and 195 UC in Cohort 2 with different inflammation status are shown in g for Cohort 1 and in h for Cohort 2. Beta diversity of these samples individually analyzed for CD and UC are shown for Cohort 1 (i) and Cohort 2 (j). Significant differences between groups were determined by one-way ANOVA corrected for multiple comparisons using BH-FDR and there is no significance between compared groups (q > 0.05) (a,b,g,h). Lines indicate mean values and error bars are standard deviations (a,b,g,h). Adonis test was used to determine significant differences between the dissimilarity distance matrix of each group and groups are not significantly different than each other (c,d,i,j). The edges strongly similar to each other are connected with a solid line (pure edge) and the edges partially similar to each other are connected with dashed lines (mixed edge) (e,f).
Extended Data Fig. 8 Co-occurrence patterns and degree centrality scores identify the important components of IBD.
a,b, Ecosystem-specific co-occurrence patterns are visualized using network diagrams where microbial phyla represent nodes and the presence of a positive co-occurrence relationship based on correlation is represented by an edge in Cohort 1 (a) and Cohort 2 (b). Co-occurrence relationships with less strong Spearman’s correlation coefficients (ρ value > 0.25 and P < 0.05) are depicted with network diagram for each disease. c,d, The value of eigenvector and betweenness centralities for CD, UC and non-IBD samples were calculated in Cohort 1 (c) and in Cohort 2 (d). Significant differences between groups were determined by non-parametric two-sided Mann–Whitney U-test (c) and ordinary one-way ANOVA corrected for multiple comparisons using BH-FDR correction (d) and P < 0.05 was considered significant and significant results are shown on the plot. Lines indicate mean values and error bars are standard deviations. 65 CD and 61 UC taxa from Cohort 1 and 48 CD, 44 UC and 41 non-IBD taxa from Cohort 2 were used for analysis in c and d. e,f, Important genera based on their between centrality score for Cohort 1 (e) and Cohort 2 (f) are depicted for each cohort (CD in red, UC in blue and Non-IBD in green). g–j, Prominent and influential taxa were identified using in- and out-degree scores are shown for CD (g,i) and UC (h,j) for corresponding cohorts: Cohort 1 (g,h) and Cohort 2 (i,j). Taxa with ρ value > 0.25 and P < 0.05 are plotted (e–j). (Cohort 1, CD phyla 9 nodes (N) and 13 edges (E); UC phyla 8 N and 12E; CD genera 64 N, 473E; UC genera, 60 N, 440E and Cohort 2: CD phyla 6 N, 7E; UC 9 N, 11E; CD genera 40 N, 273E; UC genera 38 N, 276E).
Extended Data Fig. 9 Gut microbiota differences in IBD patients with different lifestyles and different responsiveness to disease.
a–d, Major taxonomic changes were observed in 494 CD samples in Cohort 1 when samples were analyzed for sport activities (a), smoking status (b), alcohol abuse (c) and family history of disease (d). e–g, Species richness biopsy samples obtained from patients responsive (success) or unresponsive (failure) to anti-TNF-α therapies (e,f) and corticosteroid therapies (g,h) are shown in e,g for Cohort 1 and in f,h for Cohort 2. i,j, Microbial clustering of intestinal biopsy samples from IBD patients responding or non-responding to corticosteroids therapies for Cohort 1 (i) and Cohort 2 (j) is shown with PCoA on Bray–Curtis distance dissimilarity metrics. CD (solid line) and UC (dashed line) are used to identify the disease groups on PCoA plot. k–m, Unique microbial taxa identified as a signature of responding and non-responding groups in CD (182 with success and 47 with failure) (k) and in UC (l) are shown for Cohort 1 (131 with success and 36 with failure) and in UC (146 with success and 16 with failure) for Cohort 2 (m). n,o, Species richness (of biopsy samples obtained from patients with different disease activities, characterized by the frequency of exacerbations (active) and remissions (quiescent) are shown for Cohort 1 (n) and for Cohort 2 (o). Data is not significant in n and o. p,q, Microbial clustering of intestinal biopsy samples from IBD patients with different disease activities for Cohort 1 (p) and Cohort 2 (q) is shown with PCoA on Bray–Curtis distance dissimilarity metrics. CD (solid line) and UC (dashed line) are used to identify the disease groups on PCoA plot. 494 samples in CD and 447 samples in UC for Cohort 1 (p) and 226 samples in CD and 195 samples in UC for Cohort 2 (q) were analyzed. Microbial profiles were analyzed using MaAsLin pipeline with BH-FDR correction (q value) and q < 0.05 was considered significant (a–d,k–m). Significant taxa (a–d,k–m) are plotted using notched box whisker showing first and third quartiles and median value. (a) Sport: actively (several times per week), sometimes (once or twice a week) and rarely (less than once a week); (c) alcohol abuse; (d) family history: N (None), Y (Yes). Mann–Whitney U-test was used for statistical analysis of alpha diversity (e–h,n,o). No significant differences in species richness observed between groups (P > 0.05). Box-and-whisker plots display quartiles and range with standard deviations with possible outlier shown with dots in e–h,n–o. Adonis test assessed the significant difference differences between the dissimilarity distance matrix of each group (i,j,p,q). P < 0.05 for each compared group in each cohort (i,j,p,q).
Extended Data Fig. 10 The relative abundance changes in Cluster CDA taxa with disease activity and intestinal inflammation.
a, The relative abundance changes of taxa in CDA cluster in longitudinally studied 34 IBD patients (77 CD and 44 UC samples in Cohort 1 and 49 CD and 14 UC samples in Cohort 2) based on the clinically defined changes in disease activity over time as described in Fig. 6c. b, The correlation between fecal calprotectin of 78 CD patients and Cluster CDA taxa. Data was analyzed using MaAsLin with BH-FDR correction and data was not significant (NS; q > 0.05).
Supplementary information
Supplementary Information
Supplementary Tables 1–9
Rights and permissions
About this article
Cite this article
Yilmaz, B., Juillerat, P., Øyås, O. et al. Microbial network disturbances in relapsing refractory Crohn’s disease. Nat Med 25, 323–336 (2019). https://doi.org/10.1038/s41591-018-0308-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-018-0308-z
This article is cited by
-
Gut microbial network signatures of early colonizers in preterm neonates with extrauterine growth restriction
BMC Microbiology (2024)
-
Predictive biomarkers for anti-TNF alpha therapy in IBD patients
Journal of Translational Medicine (2024)
-
Short-chain fatty acids: linking diet, the microbiome and immunity
Nature Reviews Immunology (2024)
-
Mucosal host-microbe interactions associate with clinical phenotypes in inflammatory bowel disease
Nature Communications (2024)
-
The gut ileal mucosal virome is disturbed in patients with Crohn’s disease and exacerbates intestinal inflammation in mice
Nature Communications (2024)