Abstract
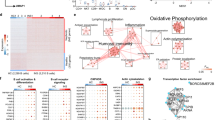
Understanding the mechanisms underlying autoantibody development will accelerate therapeutic target identification in autoimmune diseases such as systemic lupus erythematosus (SLE)1. Follicular helper T cells (TFH cells) have long been implicated in SLE pathogenesis. Yet a fraction of autoantibodies in individuals with SLE are unmutated, supporting that autoreactive B cells also differentiate outside germinal centers2. Here, we describe a CXCR5−CXCR3+ programmed death 1 (PD1)hiCD4+ helper T cell population distinct from TFH cells and expanded in both SLE blood and the tubulointerstitial areas of individuals with proliferative lupus nephritis. These cells produce interleukin-10 (IL-10) and accumulate mitochondrial reactive oxygen species as the result of reverse electron transport fueled by succinate. Furthermore, they provide B cell help, independently of IL-21, through IL-10 and succinate. Similar cells are generated in vitro upon priming naive CD4+ T cells with plasmacytoid dendritic cells activated with oxidized mitochondrial DNA, a distinct class of interferogenic toll-like receptor 9 ligand3. Targeting this pathway might blunt the initiation and/or perpetuation of extrafollicular humoral responses in SLE.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Expression array data from pDCs is available at the Gene Expression Omnibus (GEO) database under accession GSE93679. Expression array data from in vitro–generated CD4+ cells is available at the GEO database under accession GSE118951. Expression array data from ex vivo–isolated CD4+ cells is available at the GEO database under accession GSE109843. ATAC-seq data is available at the GEO database under accession GSE110017. Uncropped data for Supplementary Fig. 2g can be accessed in Supplementary Fig. 8a. All other relevant data are available from the corresponding author directly.
References
Tsokos, G. C. Systemic lupus erythematosus. N. Engl. J. Med. 365, 2110–2121 (2011).
Tipton, C. M. et al. Diversity, cellular origin and autoreactivity of antibody-secreting cell population expansions in acute systemic lupus erythematosus. Nat. Immunol. 16, 755–765 (2015).
Caielli, S. et al. Oxidized mitochondrial nucleoids released by neutrophils drive type I interferon production in human lupus. J. Exp. Med. 213, 697–713 (2016).
Guiducci, C. et al. TLR recognition of self nucleic acids hampers glucocorticoid activity in lupus. Nature 465, 937–941 (2010).
Means, T. K. et al. Human lupus autoantibody-DNA complexes activate DCs through cooperation of CD32 and TLR9. J. Clin. Investig. 115, 407–417 (2005).
Gilliet, M. & Liu, Y. J. Human plasmacytoid-derived dendritic cells and the induction of T-regulatory cells. Hum. Immunol. 63, 1149–1155 (2002).
Jego, G. et al. Plasmacytoid dendritic cells induce plasma cell differentiation through type I interferon and interleukin 6. Immunity 19, 225–234 (2003).
Ito, T. et al. Plasmacytoid dendritic cells prime IL-10-producing T regulatory cells by inducible costimulator ligand. J. Exp. Med. 204, 105–115 (2007).
Gilliet, M., Cao, W. & Liu, Y. J. Plasmacytoid dendritic cells: sensing nucleic acids in viral infection and autoimmune diseases. Nat. Rev. Immunol. 8, 594–606 (2008).
Grouard, G. et al. The enigmatic plasmacytoid T cells develop into dendritic cells with interleukin (IL)-3 and CD40-ligand. J. Exp. Med. 185, 1101–1111 (1997).
Penna, G., Sozzani, S. & Adorini, L. Cutting edge: selective usage of chemokine receptors by plasmacytoid dendritic cells. J. Immunol. 167, 1862–1866 (2001).
Glimcher, L. H. & Murphy, K. M. Lineage commitment in the immune system: the T helper lymphocyte grows up. Genes Dev. 14, 1693–1711 (2000).
Acosta-Rodriguez, E. V. et al. Surface phenotype and antigenic specificity of human interleukin 17-producing T helper memory cells. Nat. Immunol. 8, 639–646 (2007).
Jackson, S. K., DeLoose, A. & Gilbert, K. M. The ability of antigen, but not interleukin-2, to promote n-butyrate-induced T helper 1 cell anergy is associated with increased expression and altered association patterns of cyclin-dependent kinase inhibitors. Immunology 106, 486–495 (2002).
Sena, L. A. et al. Mitochondria are required for antigen-specific T cell activation through reactive oxygen species signaling. Immunity 38, 225–236 (2013).
Nag, S., Picard, P. & Stewart, D. J. Expression of nitric oxide synthases and nitrotyrosine during blood-brain barrier breakdown and repair after cold injury. Lab. Investig. 81, 41–49 (2001).
Guaras, A. et al. The CoQH2/CoQ ratio serves as a sensor of respiratory chain efficiency. Cell Rep. 15, 197–209 (2016).
Murphy, M. P. How mitochondria produce reactive oxygen species. Biochem. J. 417, 1–13 (2009).
Tannahill, G. M. et al. Succinate is an inflammatory signal that induces IL-1β through HIF-1ɑ. Nature 496, 238–242 (2013).
Oaks, Z., Winans, T., Huang, N., Banki, K. & Perl, A. Activation of the mechanistic target of rapamycin in SLE: explosion of evidence in the last five years. Curr. Rheumatol. Rep. 18, 73 (2016).
Rousset, F. et al. Interleukin 10 is a potent growth and differentiation factor for activated human B lymphocytes. Proc. Natl Acad. Sci. USA 89, 1890–1893 (1992).
Moore, K. W., de Waal Malefyt, R., Coffman, R. L. & O’Garra, A. Interleukin-10 and the interleukin-10 receptor. Annu. Rev. Immunol. 19, 683–765 (2001).
Ueno, H., Banchereau, J. & Vinuesa, C. G. Pathophysiology of T follicular helper cells in humans and mice. Nat. Immunol. 16, 142–152 (2015).
Mills, E. & O’Neill, L. A. Succinate: a metabolic signal in inflammation. Trends Cell Biol. 24, 313–320 (2014).
Buck, M. D., O’Sullivan, D. & Pearce, E. L. T cell metabolism drives immunity. J. Exp. Med. 212, 1345–1360 (2015).
Keir, M. E., Butte, M. J., Freeman, G. J. & Sharpe, A. H. PD-1 and its ligands in tolerance and immunity. Annu. Rev. Immunol. 26, 677–704 (2008).
Locci, M. et al. Human circulating PD-1+CXCR3−CXCR5+ memory TFH cells are highly functional and correlate with broadly neutralizing HIV antibody responses. Immunity 39, 758–769 (2013).
Morita, R. et al. Human blood CXCR5+CD4+ T cells are counterparts of T follicular cells and contain specific subsets that differentially support antibody secretion. Immunity 34, 108–121 (2011).
Patil, V. S. et al. Precursors of human CD4+ cytotoxic T lymphocytes identified by single-cell transcriptome analysis. Sci. Immunol, 3, eaan8664 (2018).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Arce, E. et al. Increased frequency of pre-germinal center B cells and plasma cell precursors in the blood of children with systemic lupus erythematosus. J. Immunol. 167, 2361–2369 (2001).
Dorner, T. & Lipsky, P. E. Correlation of circulating CD27high plasma cells and disease activity in systemic lupus erythematosus. Lupus 13, 283–289 (2004).
Rubtsov, A. V. et al. Toll-like receptor 7 (TLR7)-driven accumulation of a novel CD11c+ B-cell population is important for the development of autoimmunity. Blood 118, 1305–1315 (2011).
Wang, S. et al. IL-21 drives expansion and plasma cell differentiation of autoreactive CD11chiT-bet+ B cells in SLE. Nat. Commun. 9, 1758 (2018).
Blanco, P. et al. Increase in activated CD8+ T lymphocytes expressing perforin and granzyme B correlates with disease activity in patients with systemic lupus erythematosus. Arthritis Rheum. 52, 201–211 (2005).
Chang, A. et al. In situ B cell-mediated immune responses and tubulointerstitial inflammation in human lupus nephritis. J. Immunol. 186, 1849–1860 (2011).
Rao, D. A. et al. Pathologically expanded peripheral T helper cell subset drives B cells in rheumatoid arthritis. Nature 542, 110–114 (2017).
Peti-Peterdi, J. High glucose and renin release: the role of succinate and GPR91. Kidney Int. 78, 1214–1217 (2010).
Weening, J. J. et al. The classification of glomerulonephritis in systemic lupus erythematosus revisited. Kidney Int. 65, 521–530 (2004).
Hochberg, M. C. Updating the American College of Rheumatology revised criteria for the classification of systemic lupus erythematosus. Arthritis Rheum. 40, 1725 (1997).
Anoopkumar-Dukie, S. et al. Resazurin assay of radiation response in cultured cells. Br. J. Radiol. 78, 945–947 (2005).
Schmitt, N. et al. The cytokine TGF-β co-opts signaling via STAT3-STAT4 to promote the differentiation of human TFH cells. Nat. Immunol. 15, 856–865 (2014).
Garrone, P. et al. Fas ligation induces apoptosis of CD40-activated human B lymphocytes. J. Exp. Med. 182, 1265–1273 (1995).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Yu, G., Wang, L. G. & He, Q. Y. ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 31, 2382–2383 (2015).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Salabei, J. K., Gibb, A. A. & Hill, B. G. Comprehensive measurement of respiratory activity in permeabilized cells using extracellular flux analysis. Nat. Protoc. 9, 421–438 (2014).
Acknowledgements
The authors thank C. Kusminski and R. Gordillo for helping with the metabolic Seahorse experiments. We also thank N. Baldwin, R. Marches and especially our study subjects, healthy donors and their parents. This work was supported by NIH grants P50 AR054083-01 and U19 AIO82715 (V.P.), by the Baylor Scott & White Health Care Research Foundation and by the Drukier Institute for Children’s Health at Weill Cornell Medicine.
Author information
Authors and Affiliations
Contributions
S.C. performed and analyzed most of the experiments, participated in their design, provided critical discussions and co-wrote the manuscript. P.B., B.D., E.M., M.C., S.A., C.H.C. and L.W. performed and analyzed several experiments. R.B., Z.X. and D.T.V perfomed gene expression and ATAC-seq analyses. J. Baisch coordinated the sample drawing and institutional review board–related issues. T.W., M.P., L.N., K.S., J.F. and J.Z. provided subject samples and data. D.U. supervised the ATAC-seq analyses. H.U. provided help designing experiments with TFH cells. J. Banchereau provided critical suggestions and discussions throughout the study and contributed to writing the manuscript. V.P. conceived and supervised this study, was involved in the design and evaluation of all experiments and wrote the manuscript with comments from co-authors.
Corresponding author
Ethics declarations
Competing interests
V.P. has received a research grant and consulting honorarium from Sanofi-Pasteur.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8
Rights and permissions
About this article
Cite this article
Caielli, S., Veiga, D.T., Balasubramanian, P. et al. A CD4+ T cell population expanded in lupus blood provides B cell help through interleukin-10 and succinate. Nat Med 25, 75–81 (2019). https://doi.org/10.1038/s41591-018-0254-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-018-0254-9
This article is cited by
-
T cell expressions of aberrant gene signatures and Co-inhibitory receptors (Co-IRs) as predictors of renal damage and lupus disease activity
Journal of Biomedical Science (2024)
-
Cytotoxic Tph subset with low B-cell helper functions and its involvement in systemic lupus erythematosus
Communications Biology (2024)
-
Helios characterized circulating follicular helper T cells with enhanced functional phenotypes and was increased in patients with systemic lupus erythematosus
Clinical and Experimental Medicine (2024)
-
Distinct transcriptomes and autocrine cytokines underpin maturation and survival of antibody-secreting cells in systemic lupus erythematosus
Nature Communications (2024)
-
The effect of PD-1/PD-L1 signaling axis on the interaction between CD19+B cells and CD4+T cells in peripheral blood of patients with systemic lupus erythematosus
Advances in Rheumatology (2023)