Abstract
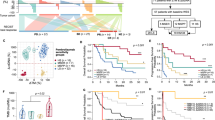
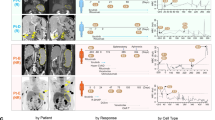
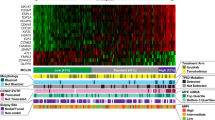
Ibrutinib plus venetoclax is a highly effective combination in mantle cell lymphoma. However, strategies to enable the evaluation of therapeutic response are required. Our prospective analyses of patients within the AIM study revealed genomic profiles that clearly dichotomized responders and nonresponders. Mutations in ATM were present in most patients who achieved a complete response, while chromosome 9p21.1–p24.3 loss and/or mutations in components of the SWI–SNF chromatin-remodeling complex were present in all patients with primary resistance and two-thirds of patients with relapsed disease. Circulating tumor DNA analysis revealed that these alterations could be dynamically monitored, providing concurrent information on treatment response and tumor evolution. Functional modeling demonstrated that compromise of the SWI–SNF complex facilitated transcriptional upregulation of BCL2L1 (Bcl-xL) providing a selective advantage against ibrutinib plus venetoclax. Together these data highlight important insights into the molecular basis of therapeutic response and provide a model for real-time assessment of innovative targeted therapies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author upon request. The sequencing data that supports the findings of this study has been deposited into the sequence read archive, which is hosted by the National Centre for Biotechnology Information. The BioProject accession number is PRJNA489753.
References
Cheah, C. Y., Seymour, J. F. & Wang, M. L. Mantle cell lymphoma. J. Clin. Oncol. 34, 1256–1269 (2016).
Le Gouill, S. et al. Rituximab after autologous stem-cell transplantation in mantle-cell lymphoma. N. Engl. J. Med. 377, 1250–1260 (2017).
Martin, P., Ruan, J. & Leonard, J. P. The potential for chemotherapy-free strategies in mantle cell lymphoma. Blood 130, 1881–1888 (2017).
Souers, A. J. et al. ABT-199, a potent and selective BCL-2 inhibitor, achieves antitumor activity while sparing platelets. Nat. Med. 19, 202–208 (2013).
Wang, M. L. et al. Targeting BTK with ibrutinib in relapsed or refractory mantle-cell lymphoma. N. Engl. J. Med. 369, 507–516 (2013).
Davids, M. S. et al. Phase I first-in-human study of venetoclax in patients with relapsed or refractory non-Hodgkin lymphoma. J. Clin. Oncol. 35, 826–833 (2017).
Dreyling, M. et al. Ibrutinib versus temsirolimus in patients with relapsed or refractory mantle-cell lymphoma: an international, randomised, open-label, phase 3 study. Lancet 387, 770–778 (2016).
Zhao, X. et al. Combination of ibrutinib with ABT-199: synergistic effects on proliferation inhibition and apoptosis in mantle cell lymphoma cells through perturbation of BTK, AKT and BCL2 pathways. Br. J. Haematol. 168, 765–768 (2015).
Axelrod, M. et al. Combinatorial drug screening identifies synergistic co-targeting of Bruton’s tyrosine kinase and the proteasome in mantle cell lymphoma. Leukemia 28, 407–410 (2014).
Li, Y. et al. FBXO10 deficiency and BTK activation upregulate BCL2 expression in mantle cell lymphoma. Oncogene 35, 6223–6234 (2016).
Beltran, E. et al. A cyclin-D1 interaction with BAX underlies its oncogenic role and potential as a therapeutic target in mantle cell lymphoma. Proc. Natl Acad. Sci. USA 108, 12461–12466 (2011).
Tam, C. S. et al. Ibrutinib plus venetoclax for the treatment of mantle-cell lymphoma. N. Engl. J. Med. 378, 1211–1223 (2018).
Bea, S. et al. Landscape of somatic mutations and clonal evolution in mantle cell lymphoma. Proc. Natl Acad. Sci. USA 110, 18250–18255 (2013).
Zhang, J. et al. The genomic landscape of mantle cell lymphoma is related to the epigenetically determined chromatin state of normal B cells. Blood 123, 2988–2996 (2014).
Kridel, R. et al. Whole transcriptome sequencing reveals recurrent NOTCH1 mutations in mantle cell lymphoma. Blood 119, 1963–1971 (2012).
Meissner, B. et al. The E3 ubiquitin ligase UBR5 is recurrently mutated in mantle cell lymphoma. Blood 121, 3161–3164 (2013).
Rahal, R. et al. Pharmacological and genomic profiling identifies NF-κB-targeted treatment strategies for mantle cell lymphoma. Nat. Med. 20, 87–92 (2014).
Greiner, T. C. et al. Mutation and genomic deletion status of ataxia telangiectasia mutated (ATM) and p53 confer specific gene expression profiles in mantle cell lymphoma. Proc. Natl Acad. Sci. USA 103, 2352–2357 (2006).
Saba, N. S. et al. Pathogenic role of B-cell receptor signaling and canonical NF-κB activation in mantle cell lymphoma. Blood 128, 82–92 (2016).
Wu, C. et al. Genetic heterogeneity in primary and relapsed mantle cell lymphomas: impact of recurrent CARD11 mutations. Oncotarget 7, 38180–38190 (2016).
Zhao, X. et al. Unification of de novo and acquired ibrutinib resistance in mantle cell lymphoma. Nat. Commun. 8, 14920 (2017).
Tahir, S. K. et al. Potential mechanisms of resistance to venetoclax and strategies to circumvent it. BMC Cancer 17, 399 (2017).
van der Velden, V. H. et al. Analysis of minimal residual disease by Ig/TCR gene rearrangements: guidelines for interpretation of real-time quantitative PCR data. Leukemia 21, 604–611 (2007).
Cheminant, M. et al. Minimal residual disease monitoring by 8-color flow cytometry in mantle cell lymphoma: an EU-MCL and LYSA study. Haematologica 101, 336–345 (2016).
Ferrero, S., Dreyling, M. & European Mantle Cell Lymphoma Network. Minimal residual disease in mantle cell lymphoma: are we ready for a personalized treatment approach? Haematologica 102, 1133–1136 (2017).
Eskelund, C. W. et al. TP53 mutations identify younger mantle cell lymphoma patients who do not benefit from intensive chemoimmunotherapy. Blood 130, 1903–1910 (2017).
Halldorsdottir, A. M. et al. Impact of TP53 mutation and 17p deletion in mantle cell lymphoma. Leukemia 25, 1904–1908 (2011).
Nordstrom, L. et al. SOX11 and TP53 add prognostic information to MIPI in a homogenously treated cohort of mantle cell lymphoma--a Nordic Lymphoma Group study. Br. J. Haematol. 166, 98–108 (2014).
Delfau-Larue, M. H. et al. High-dose cytarabine does not overcome the adverse prognostic value of CDKN2A and TP53 deletions in mantle cell lymphoma. Blood 126, 604–611 (2015).
Aukema, S. M. et al. Expression of TP53 is associated with outcome of MCL independent of MIPI and Ki-67 in trials of the European-MCL Network. Blood 131, 417–420 (2017).
Woyach, J. A. et al. Resistance mechanisms for the Bruton’s tyrosine kinase inhibitor ibrutinib. N. Engl. J. Med. 370, 2286–2294 (2014).
Mermel, C. H. et al. GISTIC2.0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome Biol. 12, R41 (2011).
Hartmann, E. M. et al. Pathway discovery in mantle cell lymphoma by integrated analysis of high-resolution gene expression and copy number profiling. Blood 116, 953–961 (2010).
Salaverria, I. et al. Specific secondary genetic alterations in mantle cell lymphoma provide prognostic information independent of the gene expression-based proliferation signature. J. Clin. Oncol. 25, 1216–1222 (2007).
Dreyling, M. H. et al. Alterations of the cyclin D1/p16-pRB pathway in mantle cell lymphoma. Cancer Res. 57, 4608–4614 (1997).
Kadoch, C. & Crabtree, G. R. Mammalian SWI/SNF chromatin remodeling complexes and cancer: mechanistic insights gained from human genomics. Sci. Adv. 1, e1500447 (2015).
Chueh, A. C. et al. ATF3 repression of BCL-XL determines apoptotic sensitivity to HDAC inhibitors across tumor types. Clin. Cancer Res. 23, 5573–5584 (2017).
Yeh, P. et al. Circulating tumour DNA reflects treatment response and clonal evolution in chronic lymphocytic leukaemia. Nat. Commun. 8, 14756 (2017).
Herling, C. D. et al. Clonal dynamics towards the development of venetoclax resistance in chronic lymphocytic leukemia. Nat. Commun. 9, 727 (2018).
Dawson, M. A. The cancer epigenome: concepts, challenges, and therapeutic opportunities. Science 355, 1147–1152 (2017).
Dawson, S. J. et al. Analysis of circulating tumor DNA to monitor metastatic breast cancer. N. Engl. J. Med. 368, 1199–1209 (2013).
Needleman, S. B. & Wunsch, C. D. A general method applicable to the search for similarities in the amino acid sequence of two proteins. J. Mol. Biol. 48, 443–453 (1970).
Koboldt, D. C. et al. VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res. 22, 568–576 (2012).
Doig, K. D. et al. PathOS: a decision support system for reporting high throughput sequencing of cancers in clinical diagnostic laboratories. Genome Med. 9, 38 (2017).
Wong, S. Q. et al. Assessing the clinical value of targeted massively parallel sequencing in a longitudinal, prospective population-based study of cancer patients. Br. J. Cancer 112, 1411–1420 (2015).
Wong, S. Q. et al. UV-associated mutations underlie the etiology of MCV-negative Merkel cell carcinomas. Cancer Res. 75, 5228–5234 (2015).
Scheinin, I. et al. DNA copy number analysis of fresh and formalin-fixed specimens by shallow whole-genome sequencing with identification and exclusion of problematic regions in the genome assembly. Genome Res. 24, 2022–2032 (2014).
Fong, C. Y. et al. BET inhibitor resistance emerges from leukaemia stem cells. Nature 525, 538–542 (2015).
Anderson, M. A. et al. The BCL2 selective inhibitor venetoclax induces rapid onset apoptosis of CLL cells in patients via a TP53-independent mechanism. Blood 127, 3215–3224 (2016).
Leverson, J. D. et al. Exploiting selective BCL-2 family inhibitors to dissect cell survival dependencies and define improved strategies for cancer therapy. Sci. Transl. Med. 7, 279ra240 (2015).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Heinze, G. & Schemper, M. A solution to the problem of monotone likelihood in Cox regression. Biometrics 57, 114–119 (2001).
Acknowledgements
The clinical study was funded by Abbvie and Janssen. The work included in this manuscript was funded by the Leukemia and Lymphoma Society (grant no. 0862-15 (S.-J.D., M.A.D., C.S.T., J.F.S.) and SCOR grant no. 11283-17 (A.W.R., D.C.S.H.)). We thank K. Doig, G. Arnau, T. Semple, T. Holloway and J. Carmody for advice and assistance with genomic sequencing, G. Lessene for providing A-1331852 and the following funders for fellowship, scholarship and grant support: CSL Centenary fellowship (S.-J.D.), Leukaemia Foundation Australia senior fellowship and Howard Hughes Medical Institute international research scholarship 55008729 (M.A.D.), NHMRC postgraduate scholarship 1114242 (R.A.) and HSANZ new investigator fellowship (R.A.), NHMRC fellowships (1089072 (C.E.T.), 1090236 (D.H.D.G.) and 1079560 (A.W.R.), NHMRC grant no. 1104549 (S.-J.D., M.A.D., C.S.T., J.F.S.) and programs 1113577, 1016701 (A.W.R., D.C.S.H.) and Leukemia and Lymphoma Society Independent Research Institutes Infrastructure Support Scheme grant no. 9000220 (R.T.), Snowdome Foundation (M.A.A.), Maddie Riewoldt’s Vision 064728 (Y.-C.C.), Victorian State Government Operational Infrastructure Support grant and Cancer Council Victoria grant (nos. 1146518, 1102104) and the Peter MacCallum Cancer Foundation. Mass cytometry was performed in part at the Materials Characterization and Fabrication Platform at the University of Melbourne and the Victorian Node of the Australian National Fabrication Facility, with support from the Victorian Comprehensive Cancer Centre.
Author information
Authors and Affiliations
Contributions
R.A. and Y.-C.C. performed the majority of the experiments, helped develop the overall concept behind the analysis, and helped write the manuscript. T.H., D.V., C.E.T., R.T., P.Y., S.Q.W., S.F., E.Y.N.L., M.A.A., C.P., O.G., C.C.B., K.K., P.B., K.R., A.Z., J.L., M.W. and D.H.D.G. contributed to data analysis. D.C.S.H. provided critical reagents. C.S.T., J.F.S. and A.W.R initiated and conducted the AIM clinical trial, provided patient data, facilitated biospecimen collection and contributed to the interpretation of the research findings. S.-J.D. and M.A.D. developed the overall concept behind the study, supervised the experiments, analyzed the data and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
C.E.T., R.T., M.A.A., D.C.S.H., D.H.D.G. and A.W.R. are employees of Walter and Eliza Hall Institute of Medical Research which receives milestone and royalty payments related to venetoclax. C.S.T., J.F.S. and A.W.R. received funding from Janssen and AbbVie to conduct the AIM clinical trial. The remaining authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10 and Supplementary Tables 4, 6 and 8–11
Supplementary Table 1
Clinicopathological and molecular characteristics of MCL cases on AIM study
Supplementary Table 2
Clinical samples used for tumor and plasma whole-exome sequencing (WES), targeted sequencing (TS) and low-coverage whole-genome sequencing (LC-WGS)
Supplementary Table 3
All variants identified from whole-exome sequencing
Supplementary Table 5
Differential expressed genes of RNA-seq data comparing SMARCA4 knockdown in Z-138 cells to control
Supplementary Table 7
List of primers used for targeted amplicon sequencing
Rights and permissions
About this article
Cite this article
Agarwal, R., Chan, YC., Tam, C.S. et al. Dynamic molecular monitoring reveals that SWI–SNF mutations mediate resistance to ibrutinib plus venetoclax in mantle cell lymphoma. Nat Med 25, 119–129 (2019). https://doi.org/10.1038/s41591-018-0243-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-018-0243-z
This article is cited by
-
Genetic and prognostic analysis of blastoid and pleomorphic mantle cell lymphoma: a multicenter analysis in China
Annals of Hematology (2024)
-
SWI/SNF complexes in hematological malignancies: biological implications and therapeutic opportunities
Molecular Cancer (2023)
-
Beyond Bruton’s tyrosine kinase inhibitors in mantle cell lymphoma: bispecific antibodies, antibody–drug conjugates, CAR T-cells, and novel agents
Journal of Hematology & Oncology (2023)
-
Utility of Measurable Residual Disease (MRD) Assessment in Mantle Cell Lymphoma
Current Treatment Options in Oncology (2023)
-
Clinical experiences with venetoclax and other pro-apoptotic agents in lymphoid malignancies: lessons from monotherapy and chemotherapy combination
Journal of Hematology & Oncology (2022)