Abstract
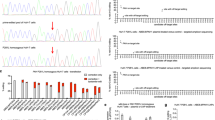
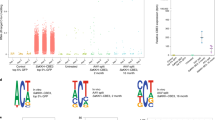
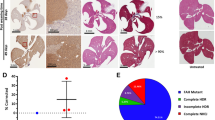
CRISPR–Cas-based genome editing holds great promise for targeting genetic disorders, including inborn errors of hepatocyte metabolism. Precise correction of disease-causing mutations in adult tissues in vivo, however, is challenging. It requires repair of Cas9-induced double-stranded DNA (dsDNA) breaks by homology-directed mechanisms, which are highly inefficient in nondividing cells. Here we corrected the disease phenotype of adult phenylalanine hydroxylase (Pah)enu2 mice, a model for the human autosomal recessive liver disease phenylketonuria (PKU)1, using recently developed CRISPR–Cas-associated base editors2,3,4. These systems enable conversion of C∙G to T∙A base pairs and vice versa, independent of dsDNA break formation and homology-directed repair (HDR). We engineered and validated an intein-split base editor, which allows splitting of the fusion protein into two parts, thereby circumventing the limited cargo capacity of adeno-associated virus (AAV) vectors. Intravenous injection of AAV-base editor systems resulted in Pahenu2 gene correction rates that restored physiological blood phenylalanine (l-Phe) levels below 120 µmol/l [5]. We observed mRNA correction rates up to 63%, restoration of phenylalanine hydroxylase (PAH) enzyme activity, and reversion of the light fur phenotype in Pahenu2 mice. Our findings suggest that targeting genetic diseases in vivo using AAV-mediated delivery of base-editing agents is feasible, demonstrating potential for therapeutic application.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All requests for materials and data are promptly reviewed by the ETH Zurich to verify whether the request is subject to any intellectual property or confidentiality obligations. Any materials and data that can be shared will be released via a Material Transfer Agreement. High-throughput sequencing data can be found at EMBL-EBI Array Express (accession number: E-MTAB-7154).
References
Shedlovsky, A., McDonald, J. D., Symula, D. & Dove, W. F. Mouse models of human phenylketonuria. Genetics 134, 1205–1210 (1993).
Kim, Y. B. et al. Increasing the genome-targeting scope and precision of base editing with engineered Cas9-cytidine deaminase fusions. Nat. Biotechnol. 35, 371–376 (2017).
Gaudelli, N. M. et al. Programmable base editing of A∙T to G∙C in genomic DNA without DNA cleavage. Nature 551, 464–471 (2017).
Komor, A. C., Kim, Y. B., Packer, M. S., Zuris, J. A. & Liu, D. R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420–424 (2016).
Blau, N., van Spronsen, F. J. & Levy, H. L. Phenylketonuria. Lancet 376, 1417–1427 (2010).
Mitchell, J. J., Trakadis, Y. J. & Scriver, C. R. Phenylalanine hydroxylase deficiency. Genet. Med. 13, 697–707 (2011).
Scriver, C. R. & Clow, C. L. Phenylketonuria: epitome of human biochemical genetics. N. Engl. J. Med. 303, 1394–1400 (1980).
Martynyuk, A. E. et al. Epilepsy in phenylketonuria: a complex dependence on serum phenylalanine levels. Epilepsia 48, 1143–1150 (2007).
Martynyuk, A. E., van Spronsen, F. J. & Van der Zee, E. A. Animal models of brain dysfunction in phenylketonuria. Mol. Genet. Metab. 99, S100–S105 (2010).
Ding, Z., Georgiev, P. & Thöny, B. Administration-route and gender-independent long-term therapeutic correction of phenylketonuria (PKU) in a mouse model by recombinant adeno-associated virus 8 pseudotyped vector-mediated gene transfer. Gene Ther. 13, 587–593 (2006).
Viecelli, H. M. et al. Treatment of phenylketonuria using minicircle-based naked-DNA gene transfer to murine liver. Hepatology 60, 1035–1043 (2014).
Harding, C. O. et al. Complete correction of hyperphenylalaninemia following liver-directed, recombinant AAV2/8 vector-mediated gene therapy in murine phenylketonuria. Gene Ther. 13, 457–462 (2006).
Yin, H. et al. Therapeutic genome editing by combined viral and non-viral delivery of CRISPR system components in vivo. Nat. Biotechnol. 34, 328–333 (2016).
Yin, H. et al. Genome editing with Cas9 in adult mice corrects a disease mutation and phenotype. Nat. Biotechnol. 32, 551–553 (2014).
Yang, Y. et al. A dual AAV system enables the Cas9-mediated correction of a metabolic liver disease in newborn mice. Nat. Biotechnol. 34, 334–338 (2016).
Mao, Z., Bozzella, M., Seluanov, A. & Gorbunova, V. Comparison of nonhomologous end joining and homologous recombination in human cells. DNA Repair (Amst.) 7, 1765–1771 (2008).
Mao, Z., Bozzella, M., Seluanov, A. & Gorbunova, V. DNA repair by nonhomologous end joining and homologous recombination during cell cycle in human cells. Cell Cycle 7, 2902–2906 (2008).
Ma, H. et al. Correction of a pathogenic gene mutation in human embryos. Nature 548, 413–419 (2017).
Gao, X. et al. Treatment of autosomal dominant hearing loss by in vivo delivery of genome editing agents. Nature 553, 217–221 (2018).
Nishida, K. et al. Targeted nucleotide editing using hybrid prokaryotic and vertebrate adaptive immune systems. Science 353, 553–563 (2016).
Nishimasu, H. et al. Crystal structure of Cas9 in complex with guide RNA and target DNA. Cell 156, 935–949 (2014).
Chadwick, A. C., Wang, X. & Musunuru, K. In vivo base editing of PCSK9 (proprotein convertase subtilisin/kexin type 9) as a therapeutic alternative to genome editing. Arterioscler. Thromb. Vasc. Biol. 9, 1741–1747 (2017).
Ryu, S.-M. et al. Adenine base editing in mouse embryos and an adult mouse model of Duchenne muscular dystrophy. Nat. Biotechnol. 36, 536–539 (2018).
Kleinstiver, B. P. et al. Engineered CRISPR–Cas9 nucleases with altered PAM specificities. Nature 523, 481–485 (2015).
Zetsche, B. et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR–Cas system. Cell 163, 759–771 (2015).
Gao, L. et al. Engineered Cpf1 variants with altered PAM specificities. Nat. Biotechnol. 35, 789–792 (2017).
Li, X. et al. Base editing with a Cpf1–cytidine deaminase fusion. Nat. Biotechnol. 36, 324–327 (2018).
Mingozzi, F. & High, K. A. Therapeutic in vivo gene transfer for genetic disease using AAV: progress and challenges. Nat. Rev. Genet. 12, 341–355 (2011).
Kay, M. A. State-of-the-art gene-based therapies: the road ahead. Nat. Rev. Genet. 12, 316–328 (2011).
Ran, F. A. et al. In vivo genome editing using Staphylococcus aureus Cas9. Nature 520, 186–191 (2015).
Nishimasu, H. et al. Crystal structure of Staphylococcus aureus Cas9. Cell 162, 1113–1126 (2015).
Truong, D. J. J. et al. Development of an intein-mediated split-Cas9 system for gene therapy. Nucleic Acids Res. 43, 6450–6458 (2015).
Nair, N. et al. Computationally designed liver-specific transcriptional modules and hyperactive factor IX improve hepatic gene therapy. Blood 123, 3195–3199 (2014).
Davidoff, A. M., Ng, C. Y. C., Zhou, J., Spence, Y. & Nathwani, A. C. Sex significantly influences transduction of murine liver by recombinant adeno-associated viral vectors through an androgen-dependent pathway. Blood 102, 480–488 (2003).
Gao, B., Jeong, W. I. & Tian, Z. Liver: an organ with predominant innate immunity. Hepatology 47, 729–736 (2008).
Racanelli, V. & Rehermann, B. The liver as an immunological organ. Hepatology 43, S54–S62 (2006).
Thöny, B., Ding, Z., Rebuffat, A. & Viecelli, H. M. Phenotypic reversion of fair hair upon gene therapy of the phenylketonuria mice. Hum. Gene Ther. 25, 573–574 (2014).
Haeussler, M. et al. Evaluation of off-target and on-target scoring algorithms and integration into the guide RNA selection tool CRISPOR. Genome Biol. 17, 148 (2016).
Kleinstiver, B. P. et al. Broadening the targeting range of Staphylococcus aureus CRISPR–Cas9 by modifying PAM recognition. Nat. Biotechnol. 33, 1293–1298 (2015).
Farin, H. F., Van Es, J. H. & Clevers, H. Redundant sources of Wnt regulate intestinal stem cells and promote formation of Paneth cells. Gastroenterology 143, 1518–1529.e7 (2012).
Barker, N. et al. Lgr5(+ve) stem cells drive self-renewal in the stomach and build long-lived gastric units in vitro. Cell Stem Cell 6, 25–36 (2010).
Zhang, J., Kobert, K., Flouri, T. & Stamatakis, A. PEAR: a fast and accurate Illumina Paired-End reAd mergeR. Bioinformatics 30, 614–620 (2014).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA–MEM. Preprint at https://arxiv.org/abs/1303.3997 (2013).
Lindsay, H. et al. CrispRVariants charts the mutation spectrum of genome engineering experiments. Nat. Biotechnol. 34, 701–702 (2016).
Pagès, H., Aboyoun, P., Gentleman, R. & DebRoy, S. Biostrings: efficient manipulation of biological strings (R Package Version 2.0, 2017).
Heintz, C., Troxler, H., Martinez, A., Thöny, B. & Blau, N. Quantification of phenylalanine hydroxylase activity by isotope-dilution liquid chromatography–electrospray ionization tandem mass spectrometry. Mol. Genet. Metab. 105, 559–565 (2012).
Acknowledgements
We thank J.-C. Paterna from the Viral Vector Facility of the Neuroscience Center Zurich for supplying vectors, the Functional Genomics Centre Zurich for sequencing in vivo samples, M. Rodriguez, A. Apladas, N. Rimann for their support with animal work, A. Garcete and I. Kleiber from the immunohistology laboratory for surgical research of the University Hospital Zurich for immunhistochemistry of liver samples. pJL-SaKKH-BE3 and pBK-YE1-BE3 were a gift from D. Liu (Addgene plasmid no. 85170 and no. 85174). WN10151 was a gift from E. Welker (Addgene plasmid no. 80441). pX601-AAV-CMV::NLS-SaCas9-NLS-3xHA-bGHpA;U6::BsaI-sgRNA was a gift from F. Zhang (Addgene plasmid no. 61591), pLenti CMV GFP Puro (658-5) was a gift from E. Campeau and P. Kaufman (Addgene plasmid no. 17448). pCMV-VSV-G was a gift from B. Weinberg (Addgene plasmid no. 8454) and psPAX2 was a gift from D. Trono (Addgene plasmid no. 12260). This work was funded by the SNSF (31003A_160230). L.V. holds an MD/PhD scholarship from the Swiss National Science Foundation.
Author information
Authors and Affiliations
Contributions
L.V. designed the research, performed experiments, analyzed data, and wrote the manuscript. F.R. conducted cryosections, fluorescent imaging, and western blot experiments and analysis thereof. J.M. and C.B.P performed western blot experiments. H.M.G.-C. and G.A. assisted with in vitro PAH assays and data analysis thereof. H.L. wrote all R scripts and assisted with HTS data analysis. R.F. analyzed blood l-Phe levels. B.T., M.D.R., J.H., and H.M.G.-C. assisted with the design of experiments G.S. designed and supervised the research and wrote the manuscript. All authors approved the final version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9 and Supplementary Tables 1–3
Rights and permissions
About this article
Cite this article
Villiger, L., Grisch-Chan, H.M., Lindsay, H. et al. Treatment of a metabolic liver disease by in vivo genome base editing in adult mice. Nat Med 24, 1519–1525 (2018). https://doi.org/10.1038/s41591-018-0209-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-018-0209-1
This article is cited by
-
Eukaryotic-driven directed evolution of Cas9 nucleases
Genome Biology (2024)
-
Continuous directed evolution of a compact CjCas9 variant with broad PAM compatibility
Nature Chemical Biology (2024)
-
Efficient prime editing in mouse brain, liver and heart with dual AAVs
Nature Biotechnology (2024)
-
Precise genome-editing in human diseases: mechanisms, strategies and applications
Signal Transduction and Targeted Therapy (2024)
-
DNA and RNA base editors can correct the majority of pathogenic single nucleotide variants
npj Genomic Medicine (2024)