Abstract
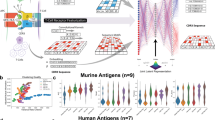
Identifying immunodominant T cell epitopes remains a significant challenge in the context of infectious disease, autoimmunity, and immuno-oncology. To address the challenge of antigen discovery, we developed a quantitative proteomic approach that enabled unbiased identification of major histocompatibility complex class II (MHCII)–associated peptide epitopes and biochemical features of antigenicity. On the basis of these data, we trained a deep neural network model for genome-scale predictions of immunodominant MHCII-restricted epitopes. We named this model bacteria originated T cell antigen (BOTA) predictor. In validation studies, BOTA accurately predicted novel CD4 T cell epitopes derived from the model pathogen Listeria monocytogenes and the commensal microorganism Muribaculum intestinale. To conclusively define immunodominant T cell epitopes predicted by BOTA, we developed a high-throughput approach to screen DNA-encoded peptide–MHCII libraries for functional recognition by T cell receptors identified from single-cell RNA sequencing. Collectively, these studies provide a framework for defining the immunodominance landscape across a broad range of immune pathologies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Source data are available for Figs. 1, 2, 5, and 7 and can be found in the Supplementary Information. There are no restrictions on source data availability. Data for Fig. 7 can be accessed through GEO accession GSE117166.
References
Babbitt, B. P., Allen, P. M., Matsueda, G., Haber, E. & Unanue, E. R. Binding of immunogenic peptides to Ia histocompatibility molecules. Nature 317, 359–361 (1985).
Stern, L. J. et al. Crystal structure of the human class II MHC protein HLA-DR1 complexed with an influenza virus peptide. Nature 368, 215–221 (1994).
Kim, A. & Sadegh-Nasseri, S. Determinants of immunodominance for CD4 T cells. Curr. Opin. Immunol. 34, 9–15 (2015).
Arunachalam, B., Phan, U. T., Geuze, H. J. & Cresswell, P. Enzymatic reduction of disulfide bonds in lysosomes: characterization of a gamma-interferon-inducible lysosomal thiol reductase (GILT). Proc. Natl Acad. Sci. USA 97, 745–750 (2000).
Hsieh, C. S., deRoos, P., Honey, K., Beers, C. & Rudensky, A. Y. A role for cathepsin L and cathepsin S in peptide generation for MHC class II presentation. J. Immunol. 168, 2618–2625 (2002).
Hsing, L. C. & Rudensky, A. Y. The lysosomal cysteine proteases in MHC class II antigen presentation. Immunol. Rev. 207, 229–241 (2005).
Miyazaki, T. et al. Mice lacking H2-M complexes, enigmatic elements of the MHC class II peptide-loading pathway. Cell 84, 531–541 (1996).
Schulze, M. S. & Wucherpfennig, K. W. The mechanism of HLA-DM induced peptide exchange in the MHC class II antigen presentation pathway. Curr. Opin. Immunol. 24, 105–111 (2012).
Rudensky, A. Y., Preston-Hurlburt, P., Hong, S. C., Barlow, A. & Janeway, C. A. Jr. Sequence analysis of peptides bound to MHC class II molecules. Nature 353, 622–627 (1991).
Hunt, D. F. et al. Peptides presented to the immune system by the murine class II major histocompatibility complex molecule I-Ad. Science 256, 1817–1820 (1992).
Chicz, R. M. et al. Predominant naturally processed peptides bound to HLA-DR1 are derived from MHC-related molecules and are heterogeneous in size. Nature 358, 764–768 (1992).
Chicz, R. M. et al. Specificity and promiscuity among naturally processed peptides bound to HLA-DR alleles. J. Exp. Med. 178, 27–47 (1993).
Sette, A. et al. Invariant chain peptides in most HLA-DR molecules of an antigen-processing mutant. Science 258, 1801–1804 (1992).
Lippolis, J. D. et al. Analysis of MHC class II antigen processing by quantitation of peptides that constitute nested sets. J. Immunol. 169, 5089–5097 (2002).
Sofron, A., Ritz, D., Neri, D. & Fugmann, T. High-resolution analysis of the murine MHC class II immunopeptidome. Eur. J. Immunol. 46, 319–328 (2016).
Mommen, G. P. et al. Sampling from the proteome to the human leukocyte antigen-DR (HLA-DR) ligandome proceeds via high specificity. Mol. Cell. Proteomics. 15, 1412–1423 (2016).
Dongre, A. R. et al. In vivo MHC class II presentation of cytosolic proteins revealed by rapid automated tandem mass spectrometry and functional analyses. Eur. J. Immunol. 31, 1485–1494 (2001).
Depontieu, F. R. et al. Identification of tumor-associated, MHC class II-restricted phosphopeptides as targets for immunotherapy. Proc. Natl Acad. Sci. USA 106, 12073–12078 (2009).
Suri, A., Walters, J. J., Rohrs, H. W., Gross, M. L. & Unanue, E. R. First signature of islet beta-cell-derived naturally processed peptides selected by diabetogenic class II MHC molecules. J. Immunol. 180, 3849–3856 (2008).
Seamons, A. et al. Competition between two MHC binding registers in a single peptide processed from myelin basic protein influences tolerance and susceptibility to autoimmunity. J. Exp. Med. 197, 1391–1397 (2003).
Nelson, C. A., Roof, R. W., McCourt, D. W. & Unanue, E. R. Identification of the naturally processed form of hen egg white lysozyme bound to the murine major histocompatibility complex class II molecule I-Ak. Proc. Natl Acad. Sci. USA 89, 7380–7383 (1992).
Brandwein, S. L. et al. Spontaneously colitic C3H/HeJBir mice demonstrate selective antibody reactivity to antigens of the enteric bacterial flora. J. Immunol. 159, 44–52 (1997).
Lodes, M. J. et al. Bacterial flagellin is a dominant antigen in Crohn disease. J. Clin. Invest. 113, 1296–1306 (2004).
Cong, Y., Feng, T., Fujihashi, K., Schoeb, T. R. & Elson, C. O. A dominant, coordinated T regulatory cell-IgA response to the intestinal microbiota. Proc. Natl Acad. Sci. USA 106, 19256–19261 (2009).
Janeway, C. A. Jr et al. Monoclonal antibodies specific for Ia glycoproteins raised by immunization with activated T cells: possible role of T cellbound Ia antigens as targets of immunoregulatory T cells. J. Immunol. 132, 662–667 (1984).
Andreatta, M., Schafer-Nielsen, C., Lund, O., Buus, S. & Nielsen, M. NNAlign: a web-based prediction method allowing non-expert end-user discovery of sequence motifs in quantitative peptide data. PLoS One 6, e26781 (2011).
Zhu, Y., Rudensky, A. Y., Corper, A. L., Teyton, L. & Wilson, I. A. Crystal structure of MHC class II I-Ab in complex with a human CLIP peptide: prediction of an I-Ab peptide-binding motif. J. Mol. Biol. 326, 1157–1174 (2003).
Liu, X. et al. Alternate interactions define the binding of peptides to the MHC molecule IA(b). Proc. Natl Acad. Sci. USA 99, 8820–8825 (2002).
Yu, N. Y. et al. PSORTb 3.0: improved protein subcellular localization prediction with refined localization subcategories and predictive capabilities for all prokaryotes. Bioinformatics 26, 1608–1615 (2010).
Tusnády, G. E. & Simon, I. Principles governing amino acid composition of integral membrane proteins: application to topology prediction. J. Mol. Biol. 283, 489–506 (1998).
Marchler-Bauer, A. et al. CDD: NCBI’s conserved domain database. Nucleic Acids Res. 43, D222–D226 (2015).
Scallan, E. et al. Foodborne illness acquired in the United States: major pathogens. Emerg. Infect. Dis. 17, 7–15 (2011).
Vita, R. et al. The immune epitope database (IEDB) 3.0. Nucleic Acids Res. 43, D405–D412 (2015).
Goyette, P. et al. High-density mapping of the MHC identifies a shared role for HLA-DRB1*01:03 in inflammatory bowel diseases and heterozygous advantage in ulcerative colitis. Nat. Genet. 47, 172–179 (2015).
Wang, P. et al. A systematic assessment of MHC class II peptide binding predictions and evaluation of a consensus approach. PLoS Comput. Biol. 4, e1000048 (2008).
Chatterjee, S. S. et al. Intracellular gene expression profile of Listeria monocytogenes. Infect. Immun. 74, 1323–1338 (2006).
Heng, T. S. et al. The Immunological Genome Project: networks of gene expression in immune cells. Nat. Immunol. 9, 1091–1094 (2008).
Weber, K. S. et al. Distinct CD4+ helper T cells involved in primary and secondary responses to infection. Proc. Natl Acad. Sci. USA 109, 9511–9516 (2012).
Palm, N. W., de Zoete, M. R. & Flavell, R. A. Immune-microbiota interactions in health and disease. Clin. Immunol. 159, 122–127 (2015).
Hall, A. B., Tolonen, A. C. & Xavier, R. J. Human genetic variation and the gut microbiome in disease. Nat. Rev. Genet. 18, 690–699 (2017).
Ormerod, K. L. et al. Genomic characterization of the uncultured Bacteroidales family S24-7 inhabiting the guts of homeothermic animals. Microbiome 4, 36 (2016).
Zeng, M. Y. et al. Gut microbiota-induced immunoglobulin G controls systemic infection by symbiotic bacteria and pathogens. Immunity 44, 647–658 (2016).
Christmann, B. S. et al. Human seroreactivity to gut microbiota antigens. J. Allergy Clin. Immunol. 136, 1378–1386.e1–5 (2015).
Stoll, M. L. et al. Altered microbiota associated with abnormal humoral immune responses to commensal organisms in enthesitis-related arthritis. Arthritis Res. Ther. 16, 486 (2014).
Conway, K. L. et al. Atg16l1 is required for autophagy in intestinal epithelial cells and protection of mice from Salmonella infection. Gastroenterology 145, 1347–1357 (2013).
Rappsilber, J., Mann, M. & Ishihama, Y. Protocol for micro-purification, enrichment, pre-fractionation and storage of peptides for proteomics using StageTips. Nat. Protoc. 2, 1896–1906 (2007).
Mertins, P. et al. Ischemia in tumors induces early and sustained phosphorylation changes in stress kinase pathways but does not affect global protein levels. Mol. Cell. Proteomics. 13, 1690–1704 (2014).
Eddy, S. R. Accelerated profile HMM searches. PLoS Comput. Biol. 7, e1002195 (2011).
Finn, R. D. et al. The Pfam protein families database: towards a more sustainable future. Nucleic Acids Res. 44, D279–D285 (2016).
Alipanahi, B., Delong, A., Weirauch, M. T. & Frey, B. J. Predicting the sequence specificities of DNA- and RNA-binding proteins by deep learning. Nat. Biotechnol. 33, 831–838 (2015).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
McDavid, A. et al. Modeling bi-modality improves characterization of cell cycle on gene expression in single cells. PLoS Comput. Biol. 10, e1003696 (2014).
Xu, C. & Su, Z. Identification of cell types from single-cell transcriptomes using a novel clustering method. Bioinformatics 31, 1974–1980 (2015).
Caporaso, J. G. et al. Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc. Natl Acad. Sci. USA 108 (Suppl 1), 4516–4522 (2011).
Caporaso, J. G. et al. Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J. 6, 1621–1624 (2012).
Krönke, J. et al. Lenalidomide induces ubiquitination and degradation of CK1α in del(5q) MDS. Nature 523, 183–188 (2015).
Bland, J. M. & Altman, D. G. Statistical methods for assessing agreement between two methods of clinical measurement. Lancet 1, 307–310 (1986).
Smyth, G. K. Linear models and empirical Bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol. 3, Article3 (2004).
Corpet, F. Multiple sequence alignment with hierarchical clustering. Nucleic Acids Res. 16, 10881–10890 (1988).
Binder, J. X. et al. COMPARTMENTS: unification and visualization of protein subcellular localization evidence. Database (Oxford) 2014, bau012 (2014).
Acknowledgements
We thank H. Vlamakis, T. Reimels, and I. Latorre for scientific input, J. Gracias for technical assistance, and P. Rogers for the FACS work. This work was supported by funding from The Leona M. and Harry B. Helmsley Charitable Trust, National Institutes of Health grants DK043351, AI109725, AT009708, and DK092405, and the Juvenile Diabetes Research Fund to R.J.X.
Author information
Authors and Affiliations
Contributions
D.B.G., C.L., and R.J.X. conceptualized the study. D.B.G., C.L., J.G.A., K.L.C., and S.A.C. constructed the study methodology. C.L. and M.Y. managed the software used in the study. C.L., M.V., and J.G.A. undertook the formal analysis of the data. D.B.G., J.G.A., C.G.M., A.L., G.J.J., E.M.B., D.J.O., and K.L.C. undertook the investigation. S.A.C. managed the resources. D.B.G. wrote the original manuscript draft. D.B.G. and R.J.X. supervised the study. R.J.X. acquired the funding for the study.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 and Supplementary Table 2
Supplementary Table 1
MHCII peptidomics
Supplementary Table 3
TCR pairing from Listeria-infected mice
Supplementary Table 4
Single-cell RNA-seq in T cells from Listeria-infected mice
Supplementary Table 5
16S rRNA sequencing from SICC-seq
Supplementary Table 6
TCR-seq and 5ʹ-DGE oligonucleotides
Rights and permissions
About this article
Cite this article
Graham, D.B., Luo, C., O’Connell, D.J. et al. Antigen discovery and specification of immunodominance hierarchies for MHCII-restricted epitopes. Nat Med 24, 1762–1772 (2018). https://doi.org/10.1038/s41591-018-0203-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-018-0203-7
This article is cited by
-
Cancer vaccines as promising immuno-therapeutics: platforms and current progress
Journal of Hematology & Oncology (2022)
-
pH sensing controls tissue inflammation by modulating cellular metabolism and endo-lysosomal function of immune cells
Nature Immunology (2022)
-
Genome binning of viral entities from bulk metagenomics data
Nature Communications (2022)
-
Immunopeptidomics-based design of mRNA vaccine formulations against Listeria monocytogenes
Nature Communications (2022)
-
Effects of Ethanol-Precipitated Argan Press Cake on the Caecal Microbiome of Mice Fed a High-Sucrose Diet
Waste and Biomass Valorization (2021)