Abstract
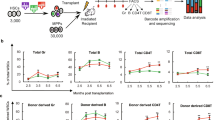
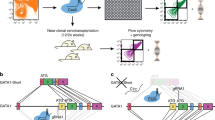
Hematopoietic stem and progenitor cells (HSPC) are endowed with the role of generating and maintaining lifelong the extremely diverse pool of blood cells1. Clinically, transplantation of human HSPC from an allogeneic healthy donor or infusion of autologous gene-corrected HSPC can effectively replenish defective blood cell production caused by congenital or acquired disorders2,3,4,5,6,7,8,9. However, due to methodological and ethical constraints that have limited the study of human HSPC primarily to in vitro assays10 or xenotransplantation models11,12, the in vivo activity of HSPC has to date remained relatively unexplored in humans13,14,15,16. Here we report a comprehensive study of the frequencies, dynamics and output of seven HSPC subtypes in humans that was performed by tracking 148,093 individual clones in six patients treated with lentiviral gene therapy using autologous HSPC transplantation and followed for up to 5 years. We discovered that primitive multipotent progenitor and hematopoietic stem cell (HSC) populations have distinct roles during the initial reconstitution after transplant, compared with subsequent steady-state phases. Furthermore, we showed that a fraction of in vitro–activated HSC are resilient and undergo a defined delayed activation period upon transplant. Finally, our data support the concept that early lymphoid-biased progenitors might be capable of long-term survival, such that they can be maintained independently of their continuous production from HSC. Overall, this study provides comprehensive data on HSPC dynamics after autologous transplantation and gene therapy in humans.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this published article (and its supplementary information files). Additional raw data files will be made available by the corresponding authors upon reasonable request.
References
Doulatov, S., Notta, F., Laurenti, E. & Dick, J. E. Hematopoiesis: a human perspective. Cell Stem Cell 10, 120–136 (2012).
De Kouchkovsky, I. & Abdul-Hay, M. Acute myeloid leukemia: a comprehensive review and 2016 update. Blood Cancer J. 6, e441 (2016).
Jabbour, E., O’Brien, S., Konopleva, M. & Kantarjian, H. New insights into the pathophysiology and therapy of adult acute lymphoblastic leukemia. Cancer 121, 2517–2528 (2015).
Pai, S.-Y. et al. Transplantation outcomes for severe combined immunodeficiency, 2000–2009. N. Engl. J. Med. 371, 434–446 (2014).
Hacein-Bey Abina, S. et al. Outcomes following gene therapy in patients with severe Wiskott–Aldrich syndrome. JAMA 313, 1550–1563 (2015).
Cicalese, M. P. & Aiuti, A. Clinical applications of gene therapy for primary immunodeficiencies. Hum. Gene Ther. 26, 210–219 (2015).
Sessa, M. et al. Lentiviral haemopoietic stem-cell gene therapy in early-onset metachromatic leukodystrophy: an ad-hoc analysis of a non-randomised, open-label, phase 1/2 trial. Lancet 388, 476–487 (2016).
Aiuti, A. et al. Lentiviral hematopoietic stem cell gene therapy in patients with Wiskott–Aldrich syndrome. Science 341, 1233151 (2013).
Hacein-Bey-Abina, S. et al. Efficacy of gene therapy for X-linked severe combined immunodeficiency. N. Engl. J. Med. 363, 355–364 (2010).
Notta, F. et al. Distinct routes of lineage development reshape the human blood hierarcy across ontogeny. Science 351, aab2116.1-9 (2016).
Doulatov, S. et al. Revised map of the human progenitor hierarchy shows the origin of macrophages and dendritic cells in early lymphoid development. Nat. Immunol. 11, 585–593 (2010).
Zonari, E. et al. Efficient ex vivo engineering and expansion of highly purified human hematopoietic stem and progenitor cell populations for gene therapy. Stem Cell Rep. 8, 977–990 (2017).
Busch, K. et al. Fundamental properties of unperturbed haematopoiesis from stem cells in vivo. Nature 518, 542–546 (2015).
Sun, J. et al. Clonal dynamics of native hematopoiesis. Nature 514, 322–327 (2015).
Sawai, C. M. et al. Hematopoietic stem cells are the major source of multilineage hematopoiesis in adult animals. Immunity 45, 597–609 (2016).
Yu, V. W. C. et al. Epigenetic memory underlies cell-autonomous heterogeneous behavior of hematopoietic stem cells. Cell 167, 1310–1322 (2016).
Biasco, L. et al. In vivo tracking of human hematopoiesis reveals patterns of clonal dynamics during early and steady-state reconstitution phases. Cell Stem Cell 19, 107–119 (2016).
Cavazzana-Calvo, M. et al. Is normal hematopoiesis maintained solely by long-term multipotent stem cells? Blood 117, 4420–4424 (2011).
Wang, G. P. et al. Dynamics of gene-modified progenitor cells analyzed by tracking retroviral integration sites in a human SCID-X1 gene therapy trial. Blood 115, 4356–4366 (2010).
Basso-Ricci, L. et al. Multiparametric whole blood dissection: a one-shot comprehensive picture of the human hematopoietic system. Cytom. A 91, 952–965 (2017).
Biasco, L. et al. In vivo tracking of T cells in humans unveils decade-long survival and activity of genetically modified T memory stem cells. Sci. Transl. Med. 7, 30–33 (2015).
Arredondo-Vega, F. X. et al. Adenosine deaminase deficiency with mosaicism for a ‘second-site suppressor’ of a splicing mutation: decline in revertant T lymphocytes during enzyme replacement therapy. Blood 99, 1005–1013 (2002).
Konno, A. et al. Differential contribution of Wiskott–Aldrich syndrome protein to selective advantage in T and B cell lineages. Blood 141, 1491–1494 (2004).
Aiuti, A. & Naldini, L. Safer conditioning for blood stem cell transplants. Nat. Biotechnol. 34, 721–723 (2016).
Simons, L. et al. Generation of adult human T-cell progenitors for immunotherapeutic applications. J. Allergy Clin. Immunol. 141, 1491–1494 (2018).
Zakrzewski, J. L. et al. Tumor immunotherapy across MHC barriers using allogeneic T-cell precursors. Nat. Biotechnol. 26, 453–461 (2008).
Kim, S. et al. Dynamics of HSPC repopulation in nonhuman primates revealed by a decade-long clonal-tracking study. Cell Stem Cell 14, 473–485 (2014).
Wu, C. et al. Clonal tracking of rhesus macaque hematopoiesis highlights a distinct lineage origin for natural killer cells. Cell Stem Cell 14, 486–499 (2014).
Jansen, R. et al. A Bayesian networks approach for predicting protein–protein interactions from genomic data. Science 302, 449–453 (2003).
Pittavino, M. et al. Comparison between generalized linear modelling and additive Bayesian network; identification of factors associated with the incidence of antibodies against Leptospira interrogans sv pomona in meat workers in New Zealand. Acta Tropica 173, 191–199 (2017).
Heckerman, D. et al. Learning Bayesian networks: the combination of knowledge and statistical data. Mach. Learn. 20, 197–243 (1995).
Koivisto, M. & Sood, K. Exact Bayesian structure discovery in Bayesian networks. J. Mach. Learn. Res. 5, 549–573 (2004).
Acknowledgements
This work was supported by Fondazione Telethon (TIGET Core Grant B2, A.A.), the Italian Ministero della Salute (Programma di Rete, NET-2011-02350069, A.A.) and the European Commission (ERARE-3-JTC 2015 EUROCID, A.A.). The work of D.P. and L.B. on bioinformatics and statistical analyses of the results and on manuscript preparation phases was performed utilizing the resources of the Gene Therapy Program of the Dana Farber/Boston Children’s Cancer and Blood Disorders Center. We thank F. Ciceri, M. G. Roncarolo and all medical and nursing staff of the Pediatric Immunohematology and Bone Marrow Transplantation Unit of the San Raffaele Scientific Institute and the San Raffaele Stem Cell Programme; S. Zancan, M. Casiraghi, S. Darin, M. Facchini, G. Tomaselli and all TIGET Clinical Trial Office personnel for clinical trial management and support; the GlaxoSmithKline research and development team for revising the manuscript; M. Gabaldo for support with project management; and P. Massariello, G. Vallanti, M. Manfredini and other MolMed staff for patient cell manipulation. We thank A. Ditadi for critically reviewing the manuscript. We thank C. Villa, E. Canonico and M. Romanò of the Flow Cytometry Resource, Advanced Cytometry Technical Applications Laboratory (FRACTAL) at Ospedale San Raffaele for cell sorting and technical help with instrumentation. We are indebted to the patients and their families for their commitment.
Author information
Authors and Affiliations
Contributions
S.S., L.B.-R., F.D. and D.P. are coauthors of this work. S.S. performed phenotypic characterization, IS retrieval of HSPC progenitors and additional molecular testing for VCN estimation, interpreted the data and wrote the manuscript; L.B.-R. performed phenotypic characterization and isolation of HSPC subpopulations, interpreted the data and wrote the manuscript; F.D. performed LAM–PCR and VCN estimation for all BM and PB patient samples; D.P. mapped IS, designed and applied mathematical models of hematopoietic hierarchy and analyzed lymphoid/myeloid contribution of subsets isolated from GT patients; F.A.S. and S.G. performed isolation of patient cell lineages; L.L. generated the bioinformatics pipeline for IS mapping; M.P.C. and F.F. provided WAS patients’ BM and PB samples and clinical data; A.A. contributed as PI by interpreting data, supervising the project and revising the manuscript; L.B. designed IS analyses, supervised the project and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The WAS gene therapy trial (NCT01515462) was originally sponsored by Fondazione Telethon and promoted by San Raffaele Telethon Institute for Gene Therapy (SR-TIGET). GlaxoSmithKline subsequently in-licensed the investigational product (GSK2696275) and became the financial and regulatory sponsor of the study. In April 2018 the license was transferred to Orchard Therapeutics. A.A. is the principal investigator (PI) of the clinical trial. L.B. is a consultant to GlaxoSmithKline for the assessment of safety of the WAS GT. All the other authors declare no competing interests. Readers are welcome to comment on the online version of the paper.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10, Supplementary Tables 1–6 and Supplementary Methods
Supplementary Dataset 1
Recapture probabilities of IS
Rights and permissions
About this article
Cite this article
Scala, S., Basso-Ricci, L., Dionisio, F. et al. Dynamics of genetically engineered hematopoietic stem and progenitor cells after autologous transplantation in humans. Nat Med 24, 1683–1690 (2018). https://doi.org/10.1038/s41591-018-0195-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-018-0195-3
This article is cited by
-
Gene therapy for inborn errors of immunity: past, present and future
Nature Reviews Immunology (2023)
-
Genotoxic effects of base and prime editing in human hematopoietic stem cells
Nature Biotechnology (2023)
-
Hematopoietic reconstitution dynamics of mobilized- and bone marrow-derived human hematopoietic stem cells after gene therapy
Nature Communications (2023)
-
Clonal reconstruction from co-occurrence of vector integration sites accurately quantifies expanding clones in vivo
Nature Communications (2022)
-
CRISPR–Cas9-mediated gene editing of the BCL11A enhancer for pediatric β0/β0 transfusion-dependent β-thalassemia
Nature Medicine (2022)