Abstract
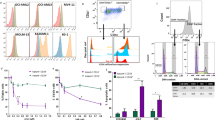
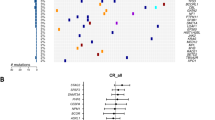
Mutations in the gene encoding isocitrate dehydrogenase 2 (IDH2) occur in several types of cancer, including acute myeloid leukemia (AML). In model systems, mutant IDH2 causes hematopoietic differentiation arrest. Enasidenib, a selective small-molecule inhibitor of mutant IDH2, produces a clinical response in 40% of treated patients with relapsed/refractory AML by promoting leukemic cell differentiation. Here, we studied the clonal basis of response and acquired resistance to enasidenib treatment. Using sequential patient samples, we determined the clonal structure of hematopoietic cell populations at different stages of differentiation. Before therapy, IDH2-mutant clones showed variable differentiation arrest. Enasidenib treatment promoted hematopoietic differentiation from either terminal or ancestral mutant clones; less frequently, treatment promoted differentiation of nonmutant cells. Analysis of paired diagnosis/relapse samples did not identify second-site mutations in IDH2 at relapse. Instead, relapse arose by clonal evolution or selection of terminal or ancestral clones, thus highlighting multiple bypass pathways that could potentially be targeted to restore differentiation arrest. These results show how mapping of clonal structure in cell populations at different stages of differentiation can reveal the response and evolution of clones during treatment response and relapse.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Abbosh, C. et al. Phylogenetic ctDNA analysis depicts early-stage lung cancer evolution. Nature 545, 446–451 (2017).
Jamal-Hanjani, M. et al. Tracking the evolution of nonsmall-cell lung cancer. N. Engl. J. Med. 376, 2109–2121 (2017).
Cancer Genome Atlas Research et al. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N. Engl. J. Med. 368, 2059–2074 (2013).
Paschka, P. et al. IDH1 and IDH2 mutations are frequent genetic alterations in acute myeloid leukemia and confer adverse prognosis in cytogenetically normal acute myeloid leukemia with NPM1 mutation without FLT3 internal tandem duplication. J. Clin. Oncol. 28, 3636–3643 (2010).
Green, C. L. et al. The prognostic significance of IDH2 mutations in AML depends on the location of the mutation. Blood 118, 409–412 (2011).
Craddock, C. F. et al. Outcome of azacitidine therapy in acute myeloid leukemia is not improved by concurrent vorinostat therapy but is predicted by a diagnostic molecular signature. Clin. Cancer Res. 23, 6430–6440 (2017).
Xu, W. et al. Oncometabolite 2-hydroxyglutarate is a competitive inhibitor of α-ketoglutarate-dependent dioxygenases. Cancer Cell 19, 17–30 (2011).
Lu, C. et al. IDH mutation impairs histone demethylation and results in a block to cell differentiation. Nature 483, 474–478 (2012).
Figueroa, M. E. et al. Leukemic IDH1 and IDH2 mutations result in a hypermethylation phenotype, disrupt TET2 function, and impair hematopoietic differentiation. Cancer Cell 18, 553–567 (2010).
Losman, J. A. et al. (R)-2-hydroxyglutarate is sufficient to promote leukemogenesis and its effects are reversible. Science 339, 1621–1625 (2013).
Wang, F. et al. Targeted inhibition of mutant IDH2 in leukemia cells induces cellular differentiation. Science 340, 622–626 (2013).
Kats, L. M. et al. Proto-oncogenic role of mutant IDH2 in leukemia initiation and maintenance. Cell Stem Cell 14, 329–341 (2014).
Yen, K. et al. AG-221, a first-in-class therapy targeting acute myeloid leukemia harboring oncogenic IDH2 mutations. Cancer Discov. 7, 478–493 (2017).
Shih, A. H. et al. Combination targeted therapy to disrupt aberrant oncogenic signaling and reverse epigenetic dysfunction in IDH2- and TET2-mutant acute myeloid leukemia. Cancer Discov. 7, 494–505 (2017).
Stein, E. M. et al. Enasidenib in mutant IDH2 relapsed or refractory acute myeloid leukemia. Blood 130, 722–731 (2017).
Amatangelo, M. D. et al. Enasidenib induces acute myeloid leukemia cell differentiation to promote clinical response. Blood 130, 732–741 (2017).
Jan, M. et al. Clonal evolution of preleukemic hematopoietic stem cells precedes human acute myeloid leukemia. Sci. Transl. Med. 4, 149ra118 (2012).
Corces-Zimmerman, M. R., Hong, W. J., Weissman, I. L., Medeiros, B. C. & Majeti, R. Preleukemic mutations in human acute myeloid leukemia affect epigenetic regulators and persist in remission. Proc. Natl Acad. Sci. USA 111, 2548–2553 (2014).
Shlush, L. I. et al. Identification of preleukaemic haematopoietic stem cells in acute leukaemia. Nature 506, 328–333 (2014).
Goardon, N. et al. Coexistence of LMPP-like and GMP-like leukemia stem cells in acute myeloid leukemia. Cancer Cell 19, 138–152 (2011).
Quek, L. et al. Genetically distinct leukemic stem cells in human CD34– acute myeloid leukemia are arrested at a hemopoietic precursor-like stage. J. Exp. Med. 213, 1513–1535 (2016).
Karamitros, D. et al. Single-cell analysis reveals the continuum of human lympho-myeloid progenitor cells. Nat. Immunol. 19, 85–97 (2018).
Maxson, J. E. et al. Oncogenic CSF3R mutations in chronic neutrophilic leukemia and atypical CML. N. Engl. J. Med. 368, 1781–1790 (2013).
Gaidzik, V. I. et al. RUNX1 mutations in acute myeloid leukemia: results from a comprehensive genetic and clinical analysis from the AML study group. J. Clin. Oncol. 29, 1364–1372 (2011).
Li, M. et al. Somatic mutations in the transcriptional corepressor gene BCORL1 in adult acute myelogenous leukemia. Blood 118, 5914–5917 (2011).
Papaemmanuil, E. et al. Genomic classification and prognosis in acute myeloid leukemia. N. Engl. J. Med. 374, 2209–2221 (2016).
Fasan, A. et al. GATA2 mutations are frequent in intermediate-risk karyotype AML with biallelic CEBPA mutations and are associated with favorable prognosis. Leukemia 27, 482–485 (2013).
Mason, C. C. et al. Age-related mutations and chronic myelomonocytic leukemia. Leukemia 30, 906–913 (2016).
Grimwade, D. et al. Refinement of cytogenetic classification in acute myeloid leukemia: determination of prognostic significance of rare recurring chromosomal abnormalities among 5876 younger adult patients treated in the United Kingdom Medical Research Council trials. Blood 116, 354–365 (2010).
Tauchert, M. J., Fourmann, J. B., Lührmann, R. & Ficner, R. Structural insights into the mechanism of the DEAH-box RNA helicase Prp43. eLife 6, e21510 (2017).
Murakami, K., Nakano, K., Shimizu, T. & Ohto, U. The crystal structure of human DEAH-box RNA helicase 15 reveals a domain organization of the mammalian DEAH/RHA family. Acta Crystallogr. F. Struct. Biol. Commun. 73, 347–355 (2017).
Arenas, J. E. & Abelson, J. N. Prp43: an RNA helicase-like factor involved in spliceosome disassembly. Proc. Natl Acad. Sci. USA 94, 11798–11802 (1997).
Fourmann, J. B. et al. Dissection of the factor requirements for spliceosome disassembly and the elucidation of its dissociation products using a purified splicing system. Genes Dev. 27, 413–428 (2013).
Faber, Z. J. et al. The genomic landscape of core-binding factor acute myeloid leukemias. Nat. Genet. 48, 1551–1556 (2016).
Farrar, J. E. et al. Genomic profiling of pediatric acute myeloid leukemia reveals a changing mutational landscape from disease diagnosis to relapse. Cancer Res. 76, 2197–2205 (2016).
Cordin, O., Banroques, J., Tanner, N. K. & Linder, P. The DEAD-box protein family of RNA helicases. Gene 367, 17–37 (2006).
Cools, J. et al. Prediction of resistance to small molecule FLT3 inhibitors: implications for molecularly targeted therapy of acute leukemia. Cancer Res. 64, 6385–6389 (2004).
Heidel, F. et al. Clinical resistance to the kinase inhibitor PKC412 in acute myeloid leukemia by mutation of Asn-676 in the FLT3 tyrosine kinase domain. Blood 107, 293–300 (2006).
Shah, N. P. et al. Multiple BCR-ABL kinase domain mutations confer polyclonal resistance to the tyrosine kinase inhibitor imatinib (STI571) in chronic phase and blast crisis chronic myeloid leukemia. Cancer Cell 2, 117–125 (2002).
Woyach, J. A. et al. Resistance mechanisms for the Bruton’s tyrosine kinase inhibitor ibrutinib. N. Engl. J. Med. 370, 2286–2294 (2014).
Kobayashi, S. et al. EGFR mutation and resistance of nonsmall-cell lung cancer to gefitinib. N. Engl. J. Med. 352, 786–792 (2005).
Choi, Y. L. et al. EML4-ALK mutations in lung cancer that confer resistance to ALK inhibitors. N. Engl. J. Med. 363, 1734–1739 (2010).
Figueroa, M. E. et al. DNA methylation signatures identify biologically distinct subtypes in acute myeloid leukemia. Cancer Cell 17, 13–27 (2010).
Kulis, M. et al. Epigenomic analysis detects widespread gene-body DNA hypomethylation in chronic lymphocytic leukemia. Nat. Genet. 44, 1236–1242 (2012).
Li, S. et al. Distinct evolution and dynamics of epigenetic and genetic heterogeneity in acute myeloid leukemia. Nat. Med. 22, 792–799 (2016).
Shaffer, S. M. et al. Rare cell variability and drug-induced reprogramming as a mode of cancer drug resistance. Nature 546, 431–435 (2017).
Sharma, S. V. et al. A chromatin-mediated reversible drug-tolerant state in cancer cell subpopulations. Cell 141, 69–80 (2010).
Landau, D. A. et al. Evolution and impact of subclonal mutations in chronic lymphocytic leukemia. Cell 152, 714–726 (2013).
Morrissy, A. S. et al. Divergent clonal selection dominates medulloblastoma at recurrence. Nature 529, 351–357 (2016).
Raffel, S. et al. BCAT1 restricts αKG levels in AML stem cells leading to IDHmut-like DNA hypermethylation. Nature 551, 384–388 (2017).
Rathert, P. et al. Transcriptional plasticity promotes primary and acquired resistance to BET inhibition. Nature 525, 543–547 (2015).
He, J. et al. Integrated genomic DNA/RNA profiling of hematologic malignancies in the clinical setting. Blood 127, 3004–3014 (2016).
Lunter, G. & Goodson, M. Stampy: a statistical algorithm for sensitive and fast mapping of Illumina sequence reads. Genome Res. 21, 936–939 (2011).
Potter, N. E. et al. Single-cell mutational profiling and clonal phylogeny in cancer. Genome Res. 23, 2115–2125 (2013).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, (15–21 (2013).
Katz, Y., Wang, E. T., Airoldi, E. M. & Burge, C. B. Analysis and design of RNA sequencing experiments for identifying isoform regulation. Nat. Methods 7, 1009–1015 (2010).
Acknowledgements
We thank the patients and clinical staff for the samples studied. L.Q. was supported by an Oxford-Celgene Fellowship; P.V. acknowledges funding from the MRC Disease Team Awards (G1000729/94931 and MR/L008963/1), MRC Molecular Haematology Unit and the Oxford Partnership Comprehensive Biomedical Research Centre (NIHR BRC Funding scheme. oxfbrc-2012-1). V.P.-L. and S.D.B. acknowledge funding from the French National Institute of Health (INSERM-AVIESAN), the National Cancer Institute (INCa-DGOS-Inserm_6043 and INCa 2012-1-RT-09), SIRIC-SOCRATE 2.0 and the Fondation Association pour la Recherche sur le Cancer (ARC, Programme). M.D.D. is funded by a fellowship from the Institut National du Cancer (INCa-DGOS_5733). M.H. is supported as a fellow of the Fondation Philanthropia, Ecole des Sciences du Cancer, Gustave Roussy, Villejuif, France. We acknowledge the Core Flow Cytometry and Next Generation Sequencing Facilities at the WIMM; the Imaging, Cytometry and Integrated Biology platforms at Gustave Roussy (P. Rameau, Y. Lecluse, N. Droin, M. K. Diop and UMS AMMICa); the clinical departments at Gustave Roussy (J.-B. Micol for clinical specimens; N. Auger for cytogenetic analyses; and C. Marzac and E. Leclercq for FLT3 genotyping) and Memorial Sloan Kettering. R.L.L. is supported by grants from the NIH, including R35 CA197594-01A1 and a Memorial Sloan Kettering Cancer Center Support Grant (NIH P30 CA008748, including a supplement to R.L.L.). The views expressed are those of the authors and not necessarily those of the NHS, the NIHR or the Department of Health.
Author information
Authors and Affiliations
Contributions
L.Q. and M.D.D. designed and performed experiments, and analyzed data; A.K., M.M., M.A., B.S., C.Q., M.H., C.W., V.S. and S. Alsafadi performed experiments and analyzed data; M.S.V. and G.S.V. analyzed data; M.A., A.S., A.P., K.Y., S. Agresta, S.D.B., R.L., E.S., K.M. and A.T. provided reagents/samples/clinical data; O.A.B., S.D.B., A.T., R.L., V.P.-L. and P.V. designed the experiments and analyzed the data. L.Q. and P.V. wrote the manuscript. All authors edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
P.V. has received research grant support from Celgene and is on its speaker bureau. L.Q. has received research grant support from Celgene.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Supplementary Text and Figures
Supplementary Tables 1–8 and Supplementary Figures 1–12
Rights and permissions
About this article
Cite this article
Quek, L., David, M.D., Kennedy, A. et al. Clonal heterogeneity of acute myeloid leukemia treated with the IDH2 inhibitor enasidenib. Nat Med 24, 1167–1177 (2018). https://doi.org/10.1038/s41591-018-0115-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-018-0115-6
This article is cited by
-
Isocitrate dehydrogenase 2 regulates the proliferation of triple-negative breast cancer through the ferroptosis pathway
Scientific Reports (2024)
-
Mutation order in acute myeloid leukemia identifies uncommon patterns of evolution and illuminates phenotypic heterogeneity
Leukemia (2024)
-
Perturbed epigenetic transcriptional regulation in AML with IDH mutations causes increased susceptibility to NK cells
Leukemia (2023)
-
Update on Small Molecule Targeted Therapies for Acute Myeloid Leukemia
Current Treatment Options in Oncology (2023)
-
FLT3 targeting in the modern era: from clonal selection to combination therapies
International Journal of Hematology (2023)