Abstract
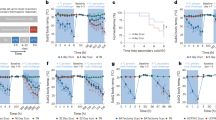
Recent research has focused on environmental effects that control tissue functionality and systemic metabolism. However, whether such stimuli affect human thermogenesis and body mass index (BMI) has not been explored. Here we show retrospectively that the presence of brown adipose tissue (BAT) and the season of conception are linked to BMI in humans. In mice, we demonstrate that cold exposure (CE) of males, but not females, before mating results in improved systemic metabolism and protection from diet-induced obesity of the male offspring. Integrated analyses of the DNA methylome and RNA sequencing of the sperm from male mice revealed several clusters of co-regulated differentially methylated regions (DMRs) and differentially expressed genes (DEGs), suggesting that the improved metabolic health of the offspring was due to enhanced BAT formation and increased neurogenesis. The conclusions are supported by cell-autonomous studies in the offspring that demonstrate an enhanced capacity to form mature active brown adipocytes, improved neuronal density and more norepinephrine release in BAT in response to cold stimulation. Taken together, our results indicate that in humans and in mice, seasonal or experimental CE induces an epigenetic programming of the sperm such that the offspring harbor hyperactive BAT and an improved adaptation to overnutrition and hypothermia.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Change history
07 August 2018
In the version of this article originally published, the bars in the mean temperature graph in Fig. 1a were incorrectly aligned. The left-most bar should have been aligned with the Apr label on the projected month of conception axis. The error has been corrected in the print, PDF and HTML versions of this article.
07 August 2018
In the version of this article originally published, the months on the axis labeled projected month of conception in Fig. 1a were out of order. April and March should have been the first and last months listed, respectively. The error has been corrected in the print, PDF and HTML versions of this article.
References
World Health Organization. Obesity and overweight. Retrieved 22 June, 2018 http://www.who.int/news-room/fact-sheets/detail/obesity-and-overweight (2017).
Rosen, E. D. & Spiegelman, B. M. What we talk about when we talk about fat. Cell 156, 20–44 (2014).
Tseng, Y. H., Cypess, A. M. & Kahn, C. R. Cellular bioenergetics as a target for obesity therapy. Nat. Rev. Drug Discov. 9, 465–482 (2010).
Frontini, A. & Cinti, S. Distribution and development of brown adipocytes in the murine and human adipose organ. Cell Metab. 11, 253–256 (2010).
Bartelt, A. & Heeren, J. Adipose tissue browning and metabolic health. Nat. Rev. Endocrinol. 10, 24–36 (2014).
Cannon, B. & Nedergaard, J. Brown adipose tissue: function and physiological significance. Physiol. Rev. 84, 277–359 (2004).
Rosenwald, M. & Wolfrum, C. The origin and definition of brite versus white and classical brown adipocytes. Adipocyte 3, 4–9 (2014).
Hany, T. F. et al. Brown adipose tissue: a factor to consider in symmetrical tracer uptake in the neck and upper chest region. Eur. J. Nucl. Med. Mol. Imaging 29, 1393–1398 (2002).
Cypess, A. M. et al. Identification and importance of brown adipose tissue in adult humans. N. Engl. J. Med. 360, 1509–1517 (2009).
Saito, M. et al. High incidence of metabolically active brown adipose tissue in healthy adult humans: effects of cold exposure and adiposity. Diabetes 58, 1526–1531 (2009).
van Marken Lichtenbelt, W. D. et al. Cold-activated brown adipose tissue in healthy men. N. Engl. J. Med. 360, 1500–1508 (2009).
Virtanen, K. A. et al. Functional brown adipose tissue in healthy adults. N. Engl. J. Med. 360, 1518–1525 (2009).
Zingaretti, M. C. et al. The presence of UCP1 demonstrates that metabolically active adipose tissue in the neck of adult humans truly represents brown adipose tissue. FASEB J. 23, 3113–3120 (2009).
Nedergaard, J., Bengtsson, T. & Cannon, B. Unexpected evidence for active brown adipose tissue in adult humans. Am. J. Physiol. Endocrinol. Metab. 293, E444–E452 (2007).
Cypess, A. M. et al. Activation of human brown adipose tissue by a β3-adrenergic receptor agonist. Cell Metab. 21, 33–38 (2015).
Yoneshiro, T. et al. Recruited brown adipose tissue as an anti-obesity agent in humans. J. Clin. Invest. 123, 3404–3408 (2013).
Carone, B. R. et al. Paternally induced transgenerational environmental reprogramming of metabolic gene expression in mammals. Cell 143, 1084–1096 (2010).
Ng, S. F. et al. Chronic high-fat diet in fathers programs beta cell dysfunction in female rat offspring. Nature 467, 963–966 (2010).
Jaenisch, R. & Bird, A. Epigenetic regulation of gene expression: how the genome integrates intrinsic and environmental signals. Nat. Genet. 33, 245–254 (2003).
Seong, K. H., Li, D., Shimizu, H., Nakamura, R. & Ishii, S. Inheritance of stress-induced, ATF-2-dependent epigenetic change. Cell 145, 1049–1061 (2011).
Anderson, L. M. et al. Preconceptional fasting of fathers alters serum glucose in offspring of mice. Nutrition 22, 327–331 (2006).
Ng, S. F. et al. Paternal high-fat diet consumption induces common changes in the transcriptomes of retroperitoneal adipose and pancreatic islet tissues in female rat offspring. FASEB J. 28, 1830–1841 (2014).
Phillips, D. I. & Young, J. B. Birth weight, climate at birth and the risk of obesity in adult life. Int. J. Obes. Relat. Metab. Disord. 24, 281–287 (2000).
Kaufman, M. H. The Atlas of Mouse Development (Academic Press, London, 1994).
Rosenwald, M., Perdikari, A., Rülicke, T. & Wolfrum, C. Bidirectional interconversion of brite and white adipocytes. Nat. Cell Biol. 15, 659–667 (2013).
Hondares, E. et al. Thermogenic activation induces FGF21 expression and release in brown adipose tissue. J. Biol. Chem. 286, 12983–12990 (2011).
Jones, P. A. Functions of DNA methylation: islands, start sites, gene bodies and beyond. Nat. Rev. Genet. 13, 484–492 (2012).
Yang, X. et al. Gene body methylation can alter gene expression and is a therapeutic target in cancer. Cancer Cell 26, 577–590 (2014).
Öst, A. et al. Paternal diet defines offspring chromatin state and intergenerational obesity. Cell 159, 1352–1364 (2014).
Lv, J. et al. The associations of month of birth with body mass index, waist circumference and leg length: findings from the China Kadoorie Biobank of 0.5 million adults. J. Epidemiol. 25, 221–230 (2015).
Speakman, J. R. & Heidari-Bakavoli, S. Type 2 diabetes, but not obesity, prevalence is positively associated with ambient temperature. Sci. Rep. 6, 30409 (2016).
Valdés, S. et al. Ambient temperature and prevalence of obesity in the Spanish population: The Di@bet.es study. Obes. (Silver Spring) 22, 2328–2332 (2014).
Yang, H. K. et al. Ambient temperature and prevalence of obesity: a nationwide population-based study in Korea. PLoS One 10, e0141724 (2015).
Afonso, A. Immigration and its impacts in Switzerland. Mediterranean Quarterly (Duke University Press) 15(4), 147–166 (2004).
Au-Yong, I. T. H., Thorn, N., Ganatra, R., Perkins, A. C. & Symonds, M. E. Brown adipose tissue and seasonal variation in humans. Diabetes 58, 2583–2587 (2009).
Adefuye, A. O., Sales, K. J. & Katz, A. A. Seminal plasma induces the expression of IL-1α in normal and neoplastic cervical cells via EP2–EGFR–PI3K–AKT pathway. J. Mol. Signal. 9, 8 (2014).
Zhang, Z. et al. Functional analysis of the cooled rat testis. J. Androl. 25, 57–68 (2004).
Fischer, A. W., Cannon, B. & Nedergaard, J. Optimal housing temperatures for mice to mimic the thermal environment of humans: an experimental study. Mol. Metab. 7, 161–170 (2018).
Bartelt, A. et al. Brown adipose tissue activity controls triglyceride clearance. Nat. Med. 17, 200–205 (2011).
Shabalina, I. G. et al. UCP1 in brite (beige) adipose tissue mitochondria is functionally thermogenic. Cell Rep. 5, 1196–1203 (2013).
Bronnikov, G., Houstĕk, J. & Nedergaard, J. β-adrenergic, cAMP-mediated stimulation of proliferation of brown fat cells in primary culture. Mediation via β1- but not via β3-adrenoceptors. J. Biol. Chem. 267, 2006–2013 (1992).
Shea, J. M. et al. Genetic and epigenetic variation, but not diet, shape the sperm methylome. Dev. Cell 35, 750–758 (2015).
Rando, O. J. Intergenerational transfer of epigenetic information in sperm. Cold Spring Harb. Perspect. Med. 6, a022988 (2016).
Greer, E. L. & Shi, Y. Histone methylation: a dynamic mark in health, disease and inheritance. Nat. Rev. Genet. 13, 343–357 (2012).
Daxinger, L. & Whitelaw, E. Understanding transgenerational epigenetic inheritance via the gametes in mammals. Nat. Rev. Genet. 13, 153–162 (2012).
Becker, A. S., Nagel, H. W., Wolfrum, C. & Burger, I. A. Anatomical grading for metabolic activity of brown adipose tissue. PLoS One 11, e0149458 (2016).
Ho, D. E., Imai, K., King, G. & Stuart, E. A. Matching as nonparametric preprocessing for reducing model dependence in parametric causal inference. Polit. Anal. 15, 199–236 (2007).
Kazak, L. et al. A creatine-driven substrate cycle enhances energy expenditure and thermogenesis in beige fat. Cell 163, 643–655 (2015).
Haueter, S. et al. Genetic vasectomy–overexpression of PRM1-EGFP fusion protein in elongating spermatids causes dominant male sterility in mice. Genesis 48, 151–160 (2010).
Whittle, A. J. et al. BMP8B increases brown adipose tissue thermogenesis through both central and peripheral actions. Cell 149, 871–885 (2012).
Abreu-Vieira, G. et al. Cidea improves the metabolic profile through expansion of adipose tissue. Nat. Commun. 6, 7433 (2015).
Pryce, C. R., Bettschen, D., Nanz-Bahr, N. I. & Feldon, J. Comparison of the effects of early handling and early deprivation on conditioned stimulus, context and spatial learning and memory in adult rats. Behav. Neurosci. 117, 883–893 (2003).
Spandl, J., White, D. J., Peychl, J. & Thiele, C. Live-cell multicolor imaging of lipid droplets with a new dye, LD540. Traffic 10, 1579–1584 (2009).
Meissburger, B. et al. Adipogenesis and insulin sensitivity in obesity are regulated by retinoid-related orphan receptor gamma. EMBO Mol. Med. 3, 637–651 (2011).
Sanchez-Gurmaches, J. & Guertin, D. A. Adipocytes arise from multiple lineages that are heterogeneously and dynamically distributed. Nat. Commun. 5, 4099 (2014).
Mehlem, A., Hagberg, C. E., Muhl, L., Eriksson, U. & Falkevall, A. Imaging of neutral lipids by oil red O for analyzing the metabolic status in health and disease. Nat. Protoc. 8, 1149–1154 (2013).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
von Meyenn, F. et al. Comparative principles of dna methylation reprogramming during human and mouse in vitro primordial germ-cell specification. Dev. Cell 39, 104–115 (2016).
Krueger, F. & Andrews, S. R. Bismark: a flexible aligner and methylation caller for bisulfite-seq applications. Bioinformatics 27, 1571–1572 (2011).
Acknowledgements
We are grateful to M. Stoffel, J. Krützfeldt and members of the Wolfrum lab for helpful discussions, K. Tabbada for assistance with WGBS high-throughput sequencing, and F. Krueger and S. Andrews for help with bioinformatics analysis. We thank K. De Bock and F. Zheng for the IB4 antibody and K. A. Rollins for editing the manuscript. Data produced and analyzed in this paper were generated in collaboration with the Genetic Diversity Center (GDC) and Functional Genomics Center Zurich (FGCZ). The work was supported by the Swiss National Science Foundation (SNSF; C.W. and F.v.M.).
Author information
Authors and Affiliations
Contributions
W.S. and C.W. designed the study; W.S. and H.D. performed all of the experimental work, except that described below; P.P. performed the IVF; S.M. helped with the Seahorse experiments; D.H.D. characterized the Ucp1-DTR-GFP mice; C.W., V.E., M.B. and D.H.D. contributed to the tracing of radiolabeled glucose; E.K. did paraffin sectioning; G.G. quantified lipid droplet sizes; A.P. helped with FACS; V.E. performed automated image analysis; L.G.S. helped with indirect calorimetry analysis; G.S. helped in the analysis of maternal behavior; D.P.-R. and W.S. did the microdialysis studies; A.S.B., I.A.B., S.B. and C.Z. performed the retrospective analysis of BAT in humans; L.O. contributed to RNA-seq data analysis; F.v.M. and W.R. did DNA methylation sequencing and bioinformatic analysis; W.S. and C.W. wrote the manuscript; and F.v.M., A.S.B., I.A.B., D.H.D., S.M., M.B. and L.B. helped with the editing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–11 and Supplementary Tables 3 and 4
Supplementary Table 1
Differentially expressed gene lists of iBAT RNA sequencing
Supplementary Table 2
Differentially methylated gene lists of sperm whole genome bisulfite sequencing
Rights and permissions
About this article
Cite this article
Sun, W., Dong, H., Becker, A.S. et al. Cold-induced epigenetic programming of the sperm enhances brown adipose tissue activity in the offspring. Nat Med 24, 1372–1383 (2018). https://doi.org/10.1038/s41591-018-0102-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-018-0102-y
This article is cited by
-
Neural circuits of long-term thermoregulatory adaptations to cold temperatures and metabolic demands
Nature Reviews Neuroscience (2024)
-
A latest progress in the study of fish behavior: cross-generational effects of behavior under pollution pressure and new technologies for behavior monitoring
Environmental Science and Pollution Research (2024)
-
Hypoxia induces alterations in tRNA modifications involved in translational control
BMC Biology (2023)
-
Epitranscriptomics in metabolic disease
Nature Metabolism (2023)
-
Cold shock induces a terminal investment reproductive response in C. elegans
Scientific Reports (2022)