Abstract
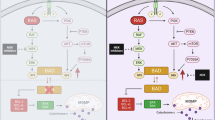
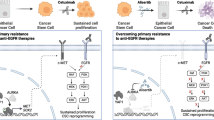
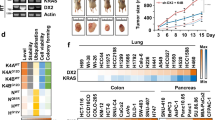
The role of KRAS, when activated through canonical mutations, has been well established in cancer1. Here we explore a secondary means of KRAS activation in cancer: focal high-level amplification of the KRAS gene in the absence of coding mutations. These amplifications occur most commonly in esophageal, gastric and ovarian adenocarcinomas2,3,4. KRAS-amplified gastric cancer models show marked overexpression of the KRAS protein and are insensitive to MAPK blockade owing to their capacity to adaptively respond by rapidly increasing KRAS–GTP levels. Here we demonstrate that inhibition of the guanine-exchange factors SOS1 and SOS2 or the protein tyrosine phosphatase SHP2 can attenuate this adaptive process and that targeting these factors, both genetically and pharmacologically, can enhance the sensitivity of KRAS-amplified models to MEK inhibition in both in vitro and in vivo settings. These data demonstrate the relevance of copy-number amplification as a mechanism of KRAS activation, and uncover the therapeutic potential for targeting of these tumors through combined SHP2 and MEK inhibition.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
09 August 2018
In the Supplementary Information originally published with this article, a lane was missing in the β-actin blot in Supplementary Fig. 2. The lane has been added. The error has been corrected in the Supplementary Information associated with this article.
References
Cox, A. D., Fesik, S. W., Kimmelman, A. C., Luo, J. & Der, C. J. Drugging the undruggable RAS: mission possible? Nat. Rev. Drug Discov. 13, 828–851 (2014).
Ross, J. S. et al. Comprehensive genomic profiling of epithelial ovarian cancer by next generation sequencing-based diagnostic assay reveals new routes to targeted therapies. Gynecol. Oncol. 130, 554–559 (2013).
Dulak, A. M. et al. Exome and whole-genome sequencing of esophageal adenocarcinoma identifies recurrent driver events and mutational complexity. Nat. Genet. 45, 478–486 (2013).
Dulak, A. M. et al. Gastrointestinal adenocarcinomas of the esophagus, stomach, and colon exhibit distinct patterns of genome instability and oncogenesis. Cancer Res. 72, 4383–4393 (2012).
Chen, Y. et al. Identification of druggable cancer driver genes amplified across TCGA datasets. PloS ONE 9, e98293 (2014).
Das, K. et al. Mutually exclusive FGFR2, HER2, and KRAS gene amplifications in gastric cancer revealed by multicolour FISH. Cancer Lett. 353, 167–175 (2014).
Birkeland, E. et al. KRAS gene amplification and overexpression but not mutation associates with aggressive and metastatic endometrial cancer. Br. J. Cancer 107, 1997–2004 (2012).
Pulciani, S., Santos, E., Long, L. K., Sorrentino, V. & Barbacid, M. ras gene amplification and malignant transformation. Mol. Cell. Biol. 5, 2836–2841 (1985).
Ahronian, L. G. et al. Clinical acquired resistance to RAF inhibitor combinations in BRAF-mutant colorectal cancer through MAPK pathway alterations. Cancer Discov. 5, 358–367 (2015).
Cargnelutti, M. et al. Activation of RAS family members confers resistance to ROS1 targeting drugs. Oncotarget 6, 5182–5194 (2015).
Oddo, D. et al. Molecular landscape of acquired resistance to targeted therapy combinations in BRAF-mutant colorectal cancer. Cancer Res. 76, 4504–4515 (2016).
Valtorta, E. et al. KRAS gene amplification in colorectal cancer and impact on response to EGFR-targeted therapy. Int. J. Cancer 133, 1259–1265 (2013).
Cox, A. D. & Der, C. J. Ras history: The saga continues. Small GTPases 1, 2–27 (2010).
Barretina, J. et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 483, 603–607 (2012).
Jokinen, E. & Koivunen, J. P. MEK and PI3K inhibition in solid tumors: rationale and evidence to date. Ther. Adv. Med. Oncol. 7, 170–180 (2015).
Sun, C. et al. Intrinsic resistance to MEK inhibition in KRAS mutant lung and colon cancer through transcriptional induction of ERBB3. Cell Rep. 7, 86–93 (2014).
Vigil, D., Cherfils, J., Rossman, K. L. & Der, C. J. Ras superfamily GEFs and GAPs: validated and tractable targets for cancer therapy? Nat. Rev. Cancer 10, 842–857 (2010).
Jeng, H. H., Taylor, L. J. & Bar-Sagi, D. Sos-mediated cross-activation of wild-type Ras by oncogenic Ras is essential for tumorigenesis. Nat. Commun. 3, 1168 (2012).
Prahallad, A. et al. PTPN11 is a central node in intrinsic and acquired resistance to targeted cancer drugs. Cell Rep. 12, 1978–1985 (2015).
Chen, Y. N. et al. Allosteric inhibition of SHP2 phosphatase inhibits cancers driven by receptor tyrosine kinases. Nature 535, 148–152 (2016).
Garcia Fortanet, J. et al. Allosteric inhibition of SHP2: identification of a potent, selective, and orally efficacious phosphatase inhibitor. J. Med. Chem. 59, 7773–7782 (2016).
Dance, M., Montagner, A., Salles, J. P., Yart, A. & Raynal, P. The molecular functions of Shp2 in the Ras/Mitogen-activated protein kinase (ERK1/2) pathway. Cell. Signal. 20, 453–459 (2008).
Bunda, S. et al. Inhibition of SHP2-mediated dephosphorylation of Ras suppresses oncogenesis. Nature Commun. 6, 8859 (2015).
Zhang, S. Q. et al. Shp2 regulates SRC family kinase activity and Ras/Erk activation by controlling Csk recruitment. Mol. Cell 13, 341–355 (2004).
Agazie, Y. M. & Hayman, M. J. Molecular mechanism for a role of SHP2 in epidermal growth factor receptor signaling. Mol. Cell. Biol. 23, 7875–7886 (2003).
Boykevisch, S. et al. Regulation of ras signaling dynamics by Sos-mediated positive feedback. Curr. Biol. 16, 2173–2179 (2006).
Araki, T., Nawa, H. & Neel, B. G. Tyrosyl phosphorylation of Shp2 is required for normal ERK activation in response to some, but not all, growth factors. J. Biol. Chem. 278, 41677–41684 (2003).
Buday, L., Warne, P. H. & Downward, J. Downregulation of the Ras activation pathway by MAP kinase phosphorylation of Sos. Oncogene 11, 1327–1331 (1995).
Kamioka, Y., Yasuda, S., Fujita, Y., Aoki, K. & Matsuda, M. Multiple decisive phosphorylation sites for the negative feedback regulation of SOS1 via ERK. J. Biol. Chem. 285, 33540–33548 (2010).
Porfiri, E. & McCormick, F. Regulation of epidermal growth factor receptor signaling by phosphorylation of the ras exchange factor hSOS1. J. Biol. Chem. 271, 5871–5877 (1996).
Manchado, E. et al. A combinatorial strategy for treating KRAS-mutant lung cancer. Nature 534, 647–651 (2016).
Lito, P. et al. Disruption of CRAF-mediated MEK activation is required for effective MEK inhibition in KRAS mutant tumors. Cancer Cell 25, 697–710 (2014).
Burgess, M. R. et al. KRAS allelic imbalance enhances fitness and modulates MAP kinase dependence in cancer. Cell 168, 817–829 (2017).
McNeill, R. S. et al. Combination therapy with potent PI3K and MAPK inhibitors overcomes adaptive kinome resistance to single agents in preclinical models of glioblastoma. Neuro-oncol. 19, 1469–1480 (2017).
Zawistowski, J. S. et al. Enhancer remodeling during adaptive bypass to MEK inhibition is attenuated by pharmacologic targeting of the P-TEFb complex. Cancer Discov. 7, 302–321 (2017).
Hunter, J. C. et al. Biochemical and structural analysis of common cancer-associated KRAS mutations. Mol. Cancer Res. 13, 1325–1335 (2015).
Lito, P., Solomon, M., Li, L. S., Hansen, R. & Rosen, N. Allele-specific inhibitors inactivate mutant KRAS G12C by a trapping mechanism. Science 351, 604–608 (2016).
Winter, J. J. et al. Small molecule binding sites on the Ras:SOS complex can be exploited for inhibition of Ras activation. J. Med. Chem. 58, 2265–2274 (2015).
Evelyn, C. R. et al. Rational design of small molecule inhibitors targeting the Ras GEF, SOS1. Chem. Biol. 21, 1618–1628 (2014).
Wang, W., Fang, G. & Rudolph, J. Ras inhibition via direct Ras binding—is there a path forward? Bioorg. Med. Chem. Lett. 22, 5766–5776 (2012).
Tokunaga, R. et al. Fibroblast growth factor receptor 2 expression, but not its genetic amplification, is associated with tumor growth and worse survival in esophagogastric junction adenocarcinoma. Oncotarget 7, 19748–19761 (2016).
Maron, S. B. et al. Targeted therapies for targeted populations: anti-EGFR treatment for EGFR amplified gastroesophageal adenocarcinoma. Cancer Discov. https://doi.org/10.1158/2159-8290.CD-17-1260 (2018).
The Cancer Genome Atlas Research Network. Comprehensive molecular characterization of gastric adenocarcinoma. Nature 513, 202–209 (2014).
Catenacci, D. V. et al. Durable complete response of metastatic gastric cancer with anti-Met therapy followed by resistance at recurrence. Cancer Discov. 1, 573–579 (2011).
Miyoshi, H. & Stappenbeck, T. S. In vitro expansion and genetic modification of gastrointestinal stem cells in spheroid culture. Nat. Protoc. 8, 2471–2482 (2013).
Catenacci, D. V. et al. Absolute quantitation of Met using mass spectrometry for clinical application: assay precision, stability, and correlation with MET gene amplification in FFPE tumor tissue. PloS ONE 9, e100586 (2014).
Hembrough, T. et al. Application of selected reaction monitoring for multiplex quantification of clinically validated biomarkers in formalin-fixed, paraffin-embedded tumor tissue. J. Mol. Diagn. 15, 454–465 (2013).
Acknowledgements
This research was supported by funding from Target Cancer Foundation, Sanofi Oncology (A.J.B., G.S.W. and K.J.), Twomey Family Fellowship in Esophageal Cancer Research (G.S.W. and J.Z.), a Research Scholar Grant from the American Cancer Society to A.J.B. and NIH grants P50 CA127003 (A.J.B.). A.J.B., K.-K.W., J.A.D. and A.K.R. were supported by NIH grant P01 CA098101. JSPS Kakenhi grant JP16H06259 and Kobayashi Foundation for Cancer Research supported Y.I. D.C. was supported by the Live Like Katie (LLK) Fund, Sal Ferrara II Fund for PANGEA, NIH K23 CA178203-01A1, University of Chicago Comprehensive Cancer Center (UCCCC) Precision Oncology-Cancer Center Support Grant P30 CA014599.
Author information
Authors and Affiliations
Contributions
G.S.W., A.J.B. and D.C. conceived the study and wrote and edited the manuscript. G.S.W., J.Z., J.B.L., Z.W., T.L., X.X., J.P., C.Z., A.D. and K.J. participated in the planning, data generation and analysis of in vitro and biochemical experiments. G.S.W., J.Z., J.B.L. and Z.W. performed tumor xenograft experiments. S.E.S., J.M., S.F., P.M., S.A.C. and R.B. performed genomic analysis. D.X., L.H., P.X., E.O’D., R.R., W.-l.L., F.C., T.H., S.S. and C.S. developed and maintained patient-derived cell lines, performed histochemical and mass spectrometric analysis. F.G., A.R., K.N., E.O., M.W., H.B. and Y.I. performed immunohistochemical and retrospective clinical outcomes analysis. A.K.R., K.-K.W. and J.A.D. provided critical input. All authors read and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
G.S.W. is now an employee of Novartis Institutes for Biomedical Research, Inc.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Wong, G.S., Zhou, J., Liu, J.B. et al. Targeting wild-type KRAS-amplified gastroesophageal cancer through combined MEK and SHP2 inhibition. Nat Med 24, 968–977 (2018). https://doi.org/10.1038/s41591-018-0022-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-018-0022-x
This article is cited by
-
DNMT1/miR-152-3p/SOS1 signaling axis promotes self-renewal and tumor growth of cancer stem-like cells derived from non-small cell lung cancer
Clinical Epigenetics (2024)
-
From bench to bedside: current development and emerging trend of KRAS-targeted therapy
Acta Pharmacologica Sinica (2024)
-
New clinical trial design in precision medicine: discovery, development and direction
Signal Transduction and Targeted Therapy (2024)
-
Biological and targeting differences between the rare KRAS A146T and canonical KRAS mutants in gastric cancer models
Gastric Cancer (2024)
-
Discovery of a novel SHP2 allosteric inhibitor using virtual screening, FMO calculation, and molecular dynamic simulation
Journal of Molecular Modeling (2024)