Abstract
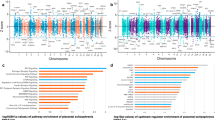
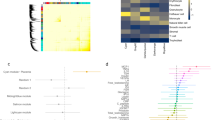
Defining the environmental context in which genes enhance disease susceptibility can provide insight into the pathogenesis of complex disorders. We report that the intra-uterine environment modulates the association of schizophrenia with genomic risk (in this study, genome-wide association study–derived polygenic risk scores (PRSs)). In independent samples from the United States, Italy, and Germany, the liability of schizophrenia explained by PRS is more than five times greater in the presence of early-life complications (ELCs) compared with their absence. Patients with ELC histories have significantly higher PRS than patients without ELC histories, which is confirmed in additional samples from Germany and Japan. The gene set composed of schizophrenia loci that interact with ELCs is highly expressed in placenta, is differentially expressed in placentae from complicated in comparison with normal pregnancies, and is differentially upregulated in placentae from male compared with female offspring. Pathway analyses reveal that genes driving the PRS-ELC interaction are involved in cellular stress response; genes that do not drive such interaction implicate orthogonal biological processes (for example, synaptic function). We conclude that a subset of the most significant genetic variants associated with schizophrenia converge on a developmental trajectory sensitive to events that affect the placental response to stress, which may offer insights into sex biases and primary prevention.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Saha, S., Chant, D., Welham, J. & McGrath, J. A systematic review of the prevalence of schizophrenia. PLoS Med. 2, e141 (2005).
McGrath, J., Saha, S., Chant, D. & Welham, J. Schizophrenia: a concise overview of incidence, prevalence, and mortality. Epidemiol. Rev. 30, 67–76 (2008).
Sullivan, P. F. The genetics of schizophrenia. PLoS Med. 2, e212 (2005).
Lichtenstein, P. et al. Common genetic determinants of schizophrenia and bipolar disorder in Swedish families: a population-based study. Lancet 373, 234–239 (2009).
Patterson, P. H. Neuroscience. Maternal effects on schizophrenia risk. Science 318, 576–577 (2007).
Gottesman, I. I. & Wolfgram, D. L. Schizophrenia Genesis: The Origins of Madness (Freeman, 1991).
Mikaelsson, M. A., Constancia, M., Dent, C. L., Wilkinson, L. S. & Humby, T. Placental programming of anxiety in adulthood revealed by Igf2-null models. Nat. Commun. 4, 2311 (2013).
Bronson, S. L. & Bale, T. L. Prenatal stress-induced increases in placental inflammation and offspring hyperactivity are male-specific and ameliorated by maternal antiinflammatory treatment. Endocrinology 155, 2635–2646 (2014).
Bronson, S. L. & Bale, T. L. The placenta as a mediator of stress effects on neurodevelopmental reprogramming. Neuropsychopharmacology 41, 207–218 (2016).
Bonnin, A. & Levitt, P. Placental source for 5-HT that tunes fetal brain development. Neuropsychopharmacology 37, 299–300 (2012).
Weinberger, D. R. From neuropathology to neurodevelopment. Lancet 346, 552–557 (1995).
Brown, A. S. & Derkits, E. J. Prenatal infection and schizophrenia: a review of epidemiologic and translational studies. Am. J. Psychiatry 167, 261–280 (2010).
Cannon, M., Jones, P. B. & Murray, R. M. Obstetric complications and schizophrenia: historical and meta-analytic review. Am. J. Psychiatry 159, 1080–1092 (2002).
Nicodemus, K. K. et al. Serious obstetric complications interact with hypoxia-regulated/vascular-expression genes to influence schizophrenia risk. Mol. Psychiatry 13, 873–877 (2008).
Schmidt-Kastner, R., van Os, J., Esquivel, G., Steinbusch, H. W. & Rutten, B. P. An environmental analysis of genes associated with schizophrenia: hypoxia and vascular factors as interacting elements in the neurodevelopmental model. Mol. Psychiatry 17, 1194–1205 (2012).
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014).
Stefansson, H. et al. Common variants conferring risk of schizophrenia. Nature 460, 744–747 (2009).
Ingason, A. et al. Copy number variations of chromosome 16p13.1 region associated with schizophrenia. Mol. Psychiatry 16, 17–25 (2011).
Purcell, S. M. et al. A polygenic burden of rare disruptive mutations in schizophrenia. Nature 506, 185–190 (2014).
International Schizophrenia Consortium et al. Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature 460, 748–752 (2009).
Wray, N. R., Goddard, M. E. & Visscher, P. M. Prediction of individual genetic risk to disease from genome-wide association studies. Genome Res. 17, 1520–1528 (2007).
McNeil, T. F. et al. Obstetric complications in histories of monozygotic twins discordant and concordant for schizophrenia. Acta Psychiatr. Scand. 89, 196–204 (1994).
McNeil, T. F., Cantor-Graae, E. & Sjostrom, K. Obstetric complications as antecedents of schizophrenia: empirical effects of using different obstetric complication scales. J. Psychiatr. Res. 28, 519–530 (1994).
Ursini, G. et al. Stress-related methylation of the catechol-O-methyltransferase Val 158 allele predicts human prefrontal cognition and activity. J. Neurosci. 31, 6692–6698 (2011).
Verdoux, H., Sutter, A. L., Glatigny-Dallay, E. & Minisini, A. Obstetrical complications and the development of postpartum depressive symptoms: a prospective survey of the MATQUID cohort. Acta Psychiatr. Scand. 106, 212–219 (2002).
Wall, J. D. & Pritchard, J. K. Haplotype blocks and linkage disequilibrium in the human genome. Nat. Rev. Genet. 4, 587–597 (2003).
Won, H. et al. Chromosome conformation elucidates regulatory relationships in developing human brain. Nature 538, 523–527 (2016).
Jaffe, A. E. et al. Developmental regulation of human cortex transcription and its clinical relevance at single base resolution. Nat. Neurosci. 18, 154–161 (2015).
Jaffe, A. E. et al. Mapping DNA methylation across development, genotype and schizophrenia in the human frontal cortex. Nat. Neurosci. 19, 40–47 (2016).
Burton, G. J. & Jauniaux, E. The cytotrophoblastic shell and complications of pregnancy. Placenta 60, 134–139 (2017).
Cotechini, T. & Graham, C. H. Aberrant maternal inflammation as a cause of pregnancy complications: a potential therapeutic target?. Placenta 36, 960–966 (2015).
Maltepe, E. & Fisher, S. J. Placenta: the forgotten organ. Annu. Rev. Cell Dev. Biol. 31, 523–552 (2015).
Maltepe, E., Bakardjiev, A. I. & Fisher, S. J. The placenta: transcriptional, epigenetic, and physiological integration during development. J. Clin. Invest. 120, 1016–1025 (2010).
Leavey, K. et al. Unsupervised placental gene expression profiling identifies clinically relevant subclasses of human preeclampsia. Hypertension 68, 137–147 (2016).
Redman, C. Pre-eclampsia: a complex and variable disease. Pregnancy Hypertens. 4, 241–242 (2014).
Ferguson, K. K., Meeker, J. D., McElrath, T. F., Mukherjee, B. & Cantonwine, D. E. Repeated measures of inflammation and oxidative stress biomarkers in preeclamptic and normotensive pregnancies. Am. J. Obstet. Gynecol. 216, 527.e521–527.e529 (2017).
Redman, C. W., Sacks, G. P. & Sargent, I. L. Preeclampsia: an excessive maternal inflammatory response to pregnancy. Am. J. Obstet. Gynecol. 180, 499–506 (1999).
Hwang, S. S. et al. Maternal substance use disorders and infant outcomes in the first year of life among Massachusetts singletons, 2003-2010. J. Pediatr. 191, 69–75 (2017).
Mol, B. W. J. et al. Pre-eclampsia. Lancet 387, 999–1011 (2016).
Lisonkova, S. & Joseph, K. S. Incidence of preeclampsia: risk factors and outcomes associated with early- versus late-onset disease. Am. J. Obstet. Gynecol. 209, 544.e541–544.e512 (2013).
Wollmann, H. A. Intrauterine growth restriction: definition and etiology. Horm. Res. 49 (Suppl. 2), 1–6 (1998).
Dalman, C., Allebeck, P., Cullberg, J., Grunewald, C. & Koster, M. Obstetric complications and the risk of schizophrenia: a longitudinal study of a national birth cohort. Arch. Gen. Psychiatry 56, 234–240 (1999).
Byrne, M., Agerbo, E., Bennedsen, B., Eaton, W. W. & Mortensen, P. B. Obstetric conditions and risk of first admission with schizophrenia: a Danish national register based study. Schizophr. Res. 97, 51–59 (2007).
Xiao, X. et al. HSF1 is required for extra-embryonic development, postnatal growth and protection during inflammatory responses in mice. EMBO J. 18, 5943–5952 (1999).
Hashimoto-Torii, K. et al. Roles of heat shock factor 1 in neuronal response to fetal environmental risks and its relevance to brain disorders. Neuron 82, 560–572 (2014).
McGrath, J. et al. A systematic review of the incidence of schizophrenia: the distribution of rates and the influence of sex, urbanicity, migrant status and methodology. BMC Med. 2, 13 (2004).
Aleman, A., Kahn, R. S. & Selten, J. P. Sex differences in the risk of schizophrenia: evidence from meta-analysis. Arch. Gen. Psychiatry 60, 565–571 (2003).
Hjorthoj, C., Sturup, A. E., McGrath, J. J. & Nordentoft, M. Years of potential life lost and life expectancy in schizophrenia: a systematic review and meta-analysis. Lancet Psychiatry 4, 295–301 (2017).
Ochoa, S., Usall, J., Cobo, J., Labad, X. & Kulkarni, J. Gender differences in schizophrenia and first-episode psychosis: a comprehensive literature review. Schizophr. Res. Treatment 2012, 916198 (2012).
Abel, K. M., Drake, R. & Goldstein, J. M. Sex differences in schizophrenia. Int. Rev. Psychiatry 22, 417–428 (2010).
Angermeyer, M. C., Kuhn, L. & Goldstein, J. M. Gender and the course of schizophrenia: differences in treated outcomes. Schizophr. Bull. 16, 293–307 (1990).
Bloom, B., Cohen, R. A. & Freeman, G. Summary health statistics for U.S. children: National Health Interview Survey, 2010. Vital Health Stat. 10, 1–80 (2011).
Burton, G. J., Moffett, A. & Keverne, B. Human evolution: brain, birthweight and the immune system. Philos. Trans. R. Soc. Lond. B Biol. Sci. 370, 20140061 (2015).
Teffer, K. & Semendeferi, K. Human prefrontal cortex: evolution, development, and pathology. Prog. Brain Res. 195, 191–218 (2012).
Rosenberg, K. R. & Trevathan, W. R. An anthropological perspective on the evolutionary context of preeclampsia in humans. J. Reprod. Immunol. 76, 91–97 (2007).
Sekar, A. et al. Schizophrenia risk from complex variation of complement component 4. Nature 530, 177–183 (2016).
Niemela, S., et al. Prenatal nicotine exposure and risk of schizophrenia among offspring in a national birth cohort. Am. J. Psychiatry 73, 799–806 (2016).
Glasson, E. J. et al. Perinatal factors and the development of autism: a population study. Arch. Gen. Psychiatry 61, 618–627 (2004).
Stoner, R. et al. Patches of disorganization in the neocortex of children with autism. N. Engl. J. Med. 370, 1209–1219 (2014).
Ross, R. G. et al. Perinatal phosphatidylcholine supplementation and early childhood behavior problems: evidence for CHRNA7 moderation. Am. J. Psychiatry 173, 509–516 (2016).
Rasetti, R. et al. Altered cortical network dynamics: a potential intermediate phenotype for schizophrenia and association with ZNF804A. Arch. Gen. Psychiatry 68, 1207–1217 (2011).
Ursini, G. et al. BDNF rs6265 methylation and genotype interact on risk for schizophrenia. Epigenetics 11, 1–13 (2016).
Begemann, M. et al. Modification of cognitive performance in schizophrenia by complexin 2 gene polymorphisms. Arch. Gen. Psychiatry 67, 879–888 (2010).
Ribbe, K. et al. The cross-sectional GRAS sample: a comprehensive phenotypical data collection of schizophrenic patients. BMC Psychiatry 10, 91 (2010).
Ohi, K. et al. Glutamate networks implicate cognitive impairments in schizophrenia: genome-wide association studies of 52 cognitive phenotypes. Schizophr. Bull. 41, 909–918 (2015).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Human Genet. 81, 559–575 (2007).
Howie, B., Marchini, J. & Stephens, M. Genotype imputation with thousands of genomes. G3 (Bethesda) 1, 457–470 (2011).
Delaneau, O., Marchini, J. & Zagury, J. F. A linear complexity phasing method for thousands of genomes. Nat. Methods 9, 179–181 (2012).
Stepniak, B. et al. Accumulated environmental risk determining age at schizophrenia onset: a deep phenotyping-based study. Lancet Psychiatry 1, 444–453 (2014).
Danilack, V. A., Nunes, A. P. & Phipps, M. G. Unexpected complications of low-risk pregnancies in the United States. Am. J. Obstet. Gynecol. 212, 809.e1–809.e6 (2015).
Hewson, D. & Bennett, A. Childbirth research data: medical records or women’s reports? Am. J. Epidemiol. 125, 484–491 (1987).
O’Callaghan, E., Larkin, C. & Waddington, J. L. Obstetric complications in schizophrenia and the validity of maternal recall. Psychol. Med. 20, 89–94 (1990).
R Development Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2008).
Keller, M. C. Gene x environment interaction studies have not properly controlled for potential confounders: the problem and the (simple) solution. Biol. Psychiatry 75, 18–24 (2014).
Zhu, Z. et al. Integration of summary data from GWAS and eQTL studies predicts complex trait gene targets. Nat. Genet. 48, 481–487 (2016).
Roadmap Epigenomics Consortium et al. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Lawrence, M. et al. Software for computing and annotating genomic ranges. PLoS Comput. Biol. 9, e1003118 (2013).
Mi, H., Muruganujan, A., Casagrande, J. T. & Thomas, P. D. Large-scale gene function analysis with the PANTHER classification system. Nat. Protoc. 8, 1551–1566 (2013).
Acknowledgements
We are grateful to the Lieber and Maltz families for their visionary support that funded the analytic work of this project. We thank all of the participants in the study and their families. We thank Joo Heon Shin for help with the gene expression analyses, and Barbara Gelao, Marina Mancini, Raffaella Romano, Rita Masellis, and Grazia Caforio for help with data acquisition. We also thank Sally Cheung and John Meyer for data management, and Susan Fisher for guidance with placental datasets and review of the manuscript. We thank the Psychiatry Genomics Consortium for providing the statistics for PRS calculation, and Thomas F. McNeil for providing the scale for ELC scoring. We also thank all of the authors of the publicly available placental datasets that have been used in this study. The collection of the ELC and genetic data for the American samples was supported by direct funding from the Intramural Research Program of the NIMH to the Clinical Brain Disorders Branch (D.R.W., PI, protocol 95-M-0150, NCT00001486, annual report number: ZIA MH002942-05), with supplemental analytic support from the Clinical and Translational Neuroscience Branch (K.F.B., PI). G.U. received partial support from P50MH094268. The collection of ELCs and genetic data for the Italian sample has been supported by the NARSAD grant no. 20337 and the “Ricerca Finalizzata” grant no. PE-2011-02347951 (both awarded to A.B.). The GRAS data collection has been supported by funding from the Max Planck Society, the Max Planck Förderstiftung, the DFG (CNMPB), EXTRABRAIN EU-FP7, and the Niedersachsen-Research Network on Neuroinfectiology (N-RENNT) (to H.E.).
Author information
Authors and Affiliations
Contributions
G.U., G.P., and D.R.W. designed the study and interpreted the results. G.U., G.P., J.F.R., E.G.H., A.E.J., and C.C. carried out statistical analyses. G.U., Q.C., M.M., R.E.S., H.E., and D.R.W. organized and performed genotyping, imputation, and risk profile scoring. G.U., S.M., M.B., J.S., K.F.B., M.F.E., R.E.S., G.B., R.H., D.R., H.E., A.B., and D.R.W. organized and carried out subject recruitment and biological material collection in the discovery sample and in the replication samples, whereas G.U., G.P., S.M., A.P., G.M., M.B., H.Y., R.H., D.R., and H.E. carried out ELC assessment. J.F.R. and E.G.H. contributed to the collection of the placental tissue used in the RNA-sequencing analysis and, together with G.U., G.P., C.C., and D.R.W., interpreted the results of the gene set enrichment analyses in placental samples from complicated pregnancies compared with controls. G.U., G.P., and D.R.W. drafted the manuscript, and all authors contributed to the final version of the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–21, Supplementary Tables 2–8, 10–14, and 19–21 and Supplementary Notes
Supplementary Table 1
Polygenic risk profile score SNPs
Supplementary Table 9
PRS1 and PRS2 genes and their enrichment in placental datasets
Supplementary Table 15
Pathway enrichment results
Supplementary Table 16
Function enrichment results
Supplementary Table 17
Gene Ontology enrichment results
Supplementary Table 18
Upstream regulators enrichment results
Rights and permissions
About this article
Cite this article
Ursini, G., Punzi, G., Chen, Q. et al. Convergence of placenta biology and genetic risk for schizophrenia. Nat Med 24, 792–801 (2018). https://doi.org/10.1038/s41591-018-0021-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-018-0021-y
This article is cited by
-
Genetic effects of sequence-conserved enhancer-like elements on human complex traits
Genome Biology (2024)
-
Abundant pleiotropy across neuroimaging modalities identified through a multivariate genome-wide association study
Nature Communications (2024)
-
Divergent epigenetic responses to perinatal asphyxia in severe mental disorders
Translational Psychiatry (2024)
-
Sex differences in DNA methylation across gestation: a large scale, cross-cohort, multi-tissue analysis
Cellular and Molecular Life Sciences (2024)
-
Association between daily breakfast habit during pregnancy and neurodevelopment in 3-year-old offspring: The Japan Environment and Children’s Study
Scientific Reports (2024)