Abstract
Non-neuronal cells are key to the complex cellular interplay that follows central nervous system insult. To understand this interplay, we generated a single-cell atlas of immune, glial and retinal pigment epithelial cells from adult mouse retina before and at multiple time points after axonal transection. We identified rare subsets in naive retina, including interferon (IFN)-response glia and border-associated macrophages, and delineated injury-induced changes in cell composition, expression programs and interactions. Computational analysis charted a three-phase multicellular inflammatory cascade after injury. In the early phase, retinal macroglia and microglia were reactivated, providing chemotactic signals concurrent with infiltration of CCR2+ monocytes from the circulation. These cells differentiated into macrophages in the intermediate phase, while an IFN-response program, likely driven by microglia-derived type I IFN, was activated across resident glia. The late phase indicated inflammatory resolution. Our findings provide a framework to decipher cellular circuitry, spatial relationships and molecular interactions following tissue injury.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Data generated during this study have been deposited in Gene Expression Omnibus under accession number GSE199317. The data can be visualized in the Broad Institute Single Cell Portal (https://singlecell.broadinstitute.org/single_cell/study/SCP1785).
Code availability
Scripts have been deposited to https://bitbucket.org/jerry00/onc-retina-script/src/master/.
References
Gadani, S. P., Walsh, J. T., Lukens, J. R. & Kipnis, J. Dealing with danger in the CNS: the response of the immune system to injury. Neuron 87, 47–62 (2015).
Shechter, R. & Schwartz, M. CNS sterile injury: just another wound healing? Trends Mol. Med. 19, 135–143 (2013).
Burda, J. E. & Sofroniew, M. V. Reactive gliosis and the multicellular response to CNS damage and disease. Neuron 81, 229–248 (2014).
Andries, L., De Groef, L. & Moons, L. Neuroinflammation and optic nerve regeneration: where do we stand in elucidating underlying cellular and molecular players? Curr. Eye Res. 45, 397–409 (2020).
Greenhalgh, A. D., David, S. & Bennett, F. C. Immune cell regulation of glia during CNS injury and disease. Nat. Rev. Neurosci. 21, 139–152 (2020).
Williams, P. R., Benowitz, L. I., Goldberg, J. L. & He, Z. Axon regeneration in the mammalian optic nerve. Annu. Rev. Vis. Sci. 6, 195–213 (2020).
Tran, N. M. et al. Single-cell profiles of retinal ganglion cells differing in resilience to injury reveal neuroprotective genes. Neuron 104, 1039–1055 (2019).
Bray, E. R. et al. Thrombospondin-1 mediates axon regeneration in retinal ganglion cells. Neuron 103, 642–657 (2019).
Jacobi, A. et al. Overlapping transcriptional programs promote survival and axonal regeneration of injured retinal ganglion cells. Neuron 110, 2625–2645 (2022).
Moalem, G. et al. Autoimmune T cells protect neurons from secondary degeneration after central nervous system axotomy. Nat. Med. 5, 49–55 (1999).
Sas, A. R. et al. A new neutrophil subset promotes CNS neuron survival and axon regeneration. Nat. Immunol. 21, 1496–1505 (2020).
Kurimoto, T. et al. Neutrophils express oncomodulin and promote optic nerve regeneration. J. Neurosci. 33, 14816–14824 (2013).
Guttenplan, K. A. et al. Neurotoxic reactive astrocytes drive neuronal death after retinal injury. Cell Rep. 31, 107776 (2020).
London, A. et al. Neuroprotection and progenitor cell renewal in the injured adult murine retina requires healing monocyte-derived macrophages. J. Exp. Med 208, 23–39 (2011).
Benhar, I., Reemst, K., Kalchenko, V. & Schwartz, M. The retinal pigment epithelium as a gateway for monocyte trafficking into the eye. EMBO J. 35, 1219–1235 (2016).
O'Koren, E. G. et al. Microglial function is distinct in different anatomical locations during retinal homeostasis and degeneration. Immunity 50, 723–737 (2019).
McMenamin, P. G., Saban, D. R. & Dando, S. J. Immune cells in the retina and choroid: two different tissue environments that require different defenses and surveillance. Prog. Retin. Eye Res. 70, 85–98 (2019).
Geisert, E. E. et al. Gene expression in the mouse eye: an online resource for genetics using 103 strains of mice. Mol. Vis. 15, 1730–1763 (2009).
Youkilis, J. C. & Bassnett, S. Single-cell RNA-sequencing analysis of the ciliary epithelium and contiguous tissues in the mouse eye. Exp. Eye Res. 213, 108811 (2021).
Lehmann, G. L. et al. Single-cell profiling reveals an endothelium-mediated immunomodulatory pathway in the eye choroid. J. Exp. Med. 217, e20190730 (2020).
Korin, B. et al. High-dimensional, single-cell characterization of the brain’s immune compartment. Nat. Neurosci. 20, 1300–1309 (2017).
Mrdjen, D. et al. High-dimensional single-cell mapping of central nervous system immune cells reveals distinct myeloid subsets in health, aging, and disease. Immunity 48, 380–395 (2018).
Van Hove, H. et al. A single-cell atlas of mouse brain macrophages reveals unique transcriptional identities shaped by ontogeny and tissue environment. Nat. Neurosci. 22, 1021–1035 (2019).
Jordão, M. J. C. et al. Single-cell profiling identifies myeloid cell subsets with distinct fates during neuroinflammation. Science 363, eaat7554 (2019).
Vecino, E., David Rodriguez, F., Ruzafa, N., Pereiro, X. & Sharma, S. C. Glia-neuron interactions in the mammalian retina. Prog. Retin. Eye Res. 51, 1–40 (2016).
Hammond, T. R. et al. Single-cell RNA sequencing of microglia throughout the mouse lifespan and in the injured brain reveals complex cell-state changes. Immunity 50, 253–271 (2019).
Ronning, K. E., Karlen, S. J., Miller, E. B. & Burns, M. E. Molecular profiling of resident and infiltrating mononuclear phagocytes during rapid adult retinal degeneration using single-cell RNA sequencing. Sci. Rep. 9, 4858 (2019).
Krasemann, S. et al. The TREM2–APOE pathway drives the transcriptional phenotype of dysfunctional microglia in neurodegenerative diseases. Immunity 47, 566–581 (2017).
Keren-Shaul, H. et al. A unique microglia type associated with restricting development of Alzheimer’s disease. Cell 169, 1276–1290 (2017).
Wieghofer, P. et al. Mapping the origin and fate of myeloid cells in distinct compartments of the eye by single-cell profiling. EMBO J. 40, e105123 (2021).
Hoang, T. et al. Gene regulatory networks controlling vertebrate retinal regeneration. Science 370, eabb8598 (2020).
Habib, N. et al. Disease-associated astrocytes in Alzheimer’s disease and aging. Nat. Neurosci. 23, 701–706 (2020).
Hasel, P., Rose, I. V. L., Sadick, J. S., Kim, R. D. & Liddelow, S. A. Neuroinflammatory astrocyte subtypes in the mouse brain. Nat. Neurosci. 24, 1475–1487 (2021).
Zamanian, J. L. et al. Genomic analysis of reactive astrogliosis. J. Neurosci. 32, 6391–6410 (2012).
Liddelow, S. A. et al. Neurotoxic reactive astrocytes are induced by activated microglia. Nature 541, 481–487 (2017).
Qu, J. & Jakobs, T. C. The time course of gene expression during reactive gliosis in the optic nerve. PLoS ONE 8, e67094 (2013).
Wohl, S. G., Schmeer, C. W., Kretz, A., Witte, O. W. & Isenmann, S. Optic nerve lesion increases cell proliferation and nestin expression in the adult mouse eye in vivo. Exp. Neurol. 219, 175–186 (2009).
Babcock, A. A., Kuziel, W. A., Rivest, S. & Owens, T. Chemokine expression by glial cells directs leukocytes to sites of axonal injury in the CNS. J. Neurosci. 23, 7922–7930 (2003).
Bosch, M. et al. Mammalian lipid droplets are innate immune hubs integrating cell metabolism and host defense. Science 370, eaay8085 (2020).
Zahabi, A. et al. A new efficient protocol for directed differentiation of retinal pigmented epithelial cells from normal and retinal disease induced pluripotent stem cells. Stem Cells Dev. 21, 2262–2272 (2012).
Zhao, C. et al. mTOR-mediated dedifferentiation of the retinal pigment epithelium initiates photoreceptor degeneration in mice. J. Clin. Invest. 121, 369–383 (2011).
Yang, J.-Y. et al. Retinal protection by sustained nanoparticle delivery of oncostatin M and ciliary neurotrophic factor into rodent models of retinal degeneration. Transl. Vis. Sci. Technol. 10, 6 (2021).
Reinhard, J., Roll, L. & Faissner, A. Tenascins in retinal and optic nerve neurodegeneration. Front. Integr. Neurosci. 11, 30 (2017).
Shemer, A. et al. Engrafted parenchymal brain macrophages differ from microglia in transcriptome, chromatin landscape and response to challenge. Nat. Commun. 9, 5206 (2018).
Eraslan, G. et al. Single-nucleus cross-tissue molecular reference maps toward understanding disease gene function. Science 376, eabl4290 (2022).
Jaitin, D. A. et al. Lipid-associated macrophages control metabolic homeostasis in a Trem2-dependent manner. Cell 178, 686–698 (2019).
Absinta, M. et al. A lymphocyte–microglia–astrocyte axis in chronic active multiple sclerosis. Nature 597, 709–714 (2021).
Margeta, M. A. et al. Apolipoprotein E4 impairs the response of neurodegenerative retinal microglia and prevents neuronal loss in glaucoma. Immunity 55, 1627–1644 (2022).
O’Koren, E. G., Mathew, R. & Saban, D. R. Fate mapping reveals that microglia and recruited monocyte-derived macrophages are definitively distinguishable by phenotype in the retina. Sci. Rep. 6, 20636 (2016).
Xu, H., Dawson, R., Forrester, J. V. & Liversidge, J. Identification of novel dendritic cell populations in normal mouse retina. Invest. Ophthalmol. Vis. Sci. 48, 1701–1710 (2007).
Yona, S. et al. Fate mapping reveals origins and dynamics of monocytes and tissue macrophages under homeostasis. Immunity 38, 79–91 (2013).
Mathys, H. et al. Temporal tracking of microglia activation in neurodegeneration at single-cell resolution. Cell Rep. 21, 366–380 (2017).
Sala Frigerio, C. et al. The major risk factors for Alzheimer’s disease: age, sex, and genes modulate the microglia response to Aβ plaques. Cell Rep. 27, 1293–1306 (2019).
Zhan, L. et al. Proximal recolonization by self-renewing microglia re-establishes microglial homeostasis in the adult mouse brain. PLoS Biol. 17, e3000134 (2019).
Huang, Y. et al. Repopulated microglia are solely derived from the proliferation of residual microglia after acute depletion. Nat. Neurosci. 21, 530–540 (2018).
Rothhammer, V. et al. Type I interferons and microbial metabolites of tryptophan modulate astrocyte activity and central nervous system inflammation via the aryl hydrocarbon receptor. Nat. Med. 22, 586–597 (2016).
Mostafavi, S. et al. Parsing the interferon transcriptional network and its disease associations. Cell 164, 564–578 (2016).
Kuse, Y., Tsuruma, K., Mizoguchi, T., Shimazawa, M. & Hara, H. Progranulin deficiency causes the retinal ganglion cell loss during development. Sci. Rep. 7, 1679 (2017).
Vigneswara, V., Berry, M., Logan, A. & Ahmed, Z. Pigment epithelium-derived factor is retinal ganglion cell neuroprotective and axogenic after optic nerve crush injury. Invest. Ophthalmol. Vis. Sci. 54, 2624–2633 (2013).
Jolly, S. et al. G protein-coupled receptor 37-like 1 modulates astrocyte glutamate transporters and neuronal NMDA receptors and is neuroprotective in ischemia. Glia 66, 47–61 (2018).
Ma, W. et al. Absence of TGFβ signaling in retinal microglia induces retinal degeneration and exacerbates choroidal neovascularization. eLife 8, e42049 (2019).
Fourgeaud, L. et al. TAM receptors regulate multiple features of microglial physiology. Nature 532, 240–244 (2016).
Sanmarco, L. M. et al. Gut-licensed IFNγ+ NK cells drive LAMP1+TRAIL+ anti-inflammatory astrocytes. Nature 590, 473–479 (2021).
Lückoff, A. et al. Interferon‐β signaling in retinal mononuclear phagocytes attenuates pathological neovascularization. EMBO Mol. Med. 8, 670–678 (2016).
Wang, W. et al. Type I interferon therapy limits CNS autoimmunity by inhibiting CXCR3-mediated trafficking of pathogenic effector T cells. Cell Rep. 28, 486–497 (2019).
Brennan, F. H. et al. Microglia coordinate cellular interactions during spinal cord repair in mice. Nat. Commun. 13, 4096 (2022).
Roy, E. R. et al. Type I interferon response drives neuroinflammation and synapse loss in Alzheimer disease. J. Clin. Invest. 130, 1912–1930 (2020).
Chakarov, S. et al. Two distinct interstitial macrophage populations coexist across tissues in specific subtissular niches. Science 363, eaau0964 (2019).
Kierdorf, K., Masuda, T., Jordão, M. J. C. & Prinz, M. Macrophages at CNS interfaces: ontogeny and function in health and disease. Nat. Rev. Neurosci. 20, 547–562 (2019).
Anderson, S. R. et al. Developmental apoptosis promotes a disease-related gene signature and independence from CSF1R signaling in retinal microglia. Cell Rep. 27, 2002–2013 (2019).
Vong, L. et al. Leptin action on GABAergic neurons prevents obesity and reduces inhibitory tone to POMC neurons. Neuron 71, 142–154 (2011).
Buffelli, M. et al. Genetic evidence that relative synaptic efficacy biases the outcome of synaptic competition. Nature 424, 430–434 (2003).
Saederup, N. et al. Selective chemokine receptor usage by central nervous system myeloid cells in CCR2–red fluorescent protein knock-in mice. PLoS ONE 5, e13693 (2010).
Fernandez-Godino, R., Garland, D. L. & Pierce, E. A. Isolation, culture and characterization of primary mouse RPE cells. Nat. Protoc. 11, 1206–1218 (2016).
Takahama, S. et al. Retinal astrocytes and GABAergic wide-field amacrine cells express PDGFRα: connection to retinal ganglion cell neuroprotection by PDGF-AA. Invest. Ophthalmol. Vis. Sci. 58, 4703–4711 (2017).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Mack, M. et al. Expression and characterization of the chemokine receptors CCR2 and CCR5 in mice. J. Immunol. 166, 4697–4704 (2001).
Okunuki, Y. et al. Retinal microglia initiate neuroinflammation in ocular autoimmunity. Proc. Natl Acad. Sci. USA 116, 9989–9998 (2019).
Ding, J. & Regev, A. Deep generative model embedding of single-cell RNA-Seq profiles on hyperspheres and hyperbolic spaces. Nat. Commun. 12, 2554 (2021).
Blondel, V. D., Guillaume, J.-L., Lambiotte, R. & Lefebvre, E. Fast unfolding of communities in large networks. J. Stat. Mech. 2008, P10008 (2008).
Levine, J. H. et al. Data-driven phenotypic dissection of AML reveals progenitor-like cells that correlate with prognosis. Cell 162, 184–197 (2015).
Ding, J. et al. Systematic comparison of single-cell and single-nucleus RNA-sequencing methods. Nat. Biotechnol. 38, 737–746 (2020).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 (2021).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Liberzon, A. et al. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
Schiebinger, G. et al. Optimal-transport analysis of single-cell gene expression identifies developmental trajectories in reprogramming. Cell 176, 928–943 (2019).
La Manno, G. et al. RNA velocity of single cells. Nature 560, 494–498 (2018).
Bergen, V., Lange, M., Peidli, S., Wolf, F. A. & Theis, F. J. Generalizing RNA velocity to transient cell states through dynamical modeling. Nat. Biotechnol. 38, 1408–1414 (2020).
Jerby-Arnon, L. & Regev, A. DIALOGUE maps multicellular programs in tissue from single-cell or spatial transcriptomics data. Nat. Biotechnol. 40, 1467–1477 (2022).
Baccin, C. et al. Combined single-cell and spatial transcriptomics reveal the molecular, cellular and spatial bone marrow niche organization. Nat. Cell Biol. 22, 38–48 (2020).
Hou, R., Denisenko, E., Ong, H. T., Ramilowski, J. A. & Forrest, A. R. R. Predicting cell-to-cell communication networks using NATMI. Nat. Commun. 11, 5011 (2020).
Wheeler, M. A. et al. MAFG-driven astrocytes promote CNS inflammation. Nature 578, 593–599 (2020).
Goldberger, J., Hinton, G. E., Roweis, S. & Salakhutdinov, R. R. Neighbourhood components analysis. In Advances in Neural Information Processing Systems 17 (NIPS 2004) (eds Saul, L. et al.) (MIT Press, 2004).
van den Brink, S. C. et al. Single-cell sequencing reveals dissociation-induced gene expression in tissue subpopulations. Nat. Methods 14, 935–936 (2017).
Gharahkhani, P. et al. Genome-wide meta-analysis identifies 127 open-angle glaucoma loci with consistent effect across ancestries. Nat. Commun. 12, 1258 (2021).
Han, X. et al. Automated AI labeling of optic nerve head enables insights into cross-ancestry glaucoma risk and genetic discovery in >280,000 images from UKB and CLSA. Am. J. Hum. Genet. 108, 1204–1216 (2021).
van Zyl, T. et al. Cell atlas of the human ocular anterior segment: tissue-specific and shared cell types. Proc. Natl Acad. Sci. USA 119, e2200914119 (2022).
Acknowledgements
We acknowledge the authors whose work could not be cited due to space limitations. We thank I. Avraham-Davidi, T. van Zyl, R. Kedmi, I. Shachar and M. Schwartz for helpful discussions, M. Schwartz and M. Mack for kindly providing the MC-21 antibody, Y. Okunuki and K. Connor for their help with the PLX experiment, L. Jerby-Arnon for assistance with the DIALOGUE analysis, K. Dey and K. Jagadeesh for help with genome-wide association study analysis, R. Harpaz for help with RPE image analysis and Z. Niziolek, S. Turney, Y. Li, R. Schaffer, M. Laboulaye, E. Martersteck, T. Delorey, D. Phillips and staff members of the Harvard University Bauer Core Facility and the Koch Institute Histology Core for technical assistance. We thank L. Gaffney and A. Hupalowska for help with figure preparation and the Richter family for support. This study was supported by the Human Frontier Science Program (I.B.), the Center for Integration in Science of the Israeli Ministry of Absorption (I.B.), HHMI (A.R.), the Klarman Family Foundation (A.R.), Wings for Life Spinal Cord Research Foundation (A.J.) and NIH grants EY028633 and MH105960 (J.R.S.). The funders had no role in the study design, experiments performed, data collection, data analysis and interpretation or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
I.B. and A.R. conceived the study. I.B., I.E.W., K.S., J.R.S. and A.R. designed experiments and analyzed data. I.B., I.E.W., A.J., M.S., G.B., N.M.T. and C.W. performed experiments. J.D., W.Y. and K.S. developed computational approaches and analyzed scRNA-seq data. Z.H., J.R.S. and A.R. provided supervision and acquired funding. I.B., J.D. and A.R. wrote the manuscript, with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
A.R. is a founder and equity holder of Celsius Therapeutics, an equity holder in Immunitas Therapeutics and, until 31 August 2020, was a SAB member of Syros Pharmaceuticals, Neogene Therapeutics, Asimov and Thermo Fisher Scientific. From 1 August 2020, A.R. is an employee of Genentech and has equity in Roche. J.R.S. is a consultant for Biogen. Z.H. is an advisor of Myro Therapeutics, Axonis and Rugen Therapeutics. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Immunology thanks Daniel Saban and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Ioana Visan, in collaboration with the Nature Immunology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Experimental strategy and reproducibility.
a, Experimental overview. b, Flow cytometry gating strategy used to enrich for retinal CD45+ immune cells, CD140a+ (Pdgfra+) astrocytes and GLAST+ Müller glia (MG). c, Distribution of expression levels of RNAs for the markers used to sort immune (Ptprc = CD45), Müller glia (Slc1a3 = GLAST), and astrocyte (Pdgfra = CD140a) subsets. d-f, Variation of cell composition across experiments. Fraction (y axis) of immune (d), MG (e) and astrocyte (f) subsets across 10x channels (x axis; legend).
Extended Data Fig. 2 Changes in cell composition and stability in cell marker expression throughout the time course.
a, 2D Uniform Manifold Approximation and Projection (UMAP) for 121,309 single cells profiled from the retina across 0, 0.5, 1, 2, 4, 7 and 14dpc, colored by time point (legend). b, ScPhere embedding of 121,309 single cell profiles (dots) from the retina, projected to 2D by the Equal Earth map projection method, colored by time point (legend). c, Fraction of expressing cells (dot size) and mean expression levels in expressing cells (dot color) of selected marker genes (columns) across 14 non-neuronal cell types (rows), plotted at each time point. d, UMAP for 21,275 cells profiled from mouse posterior eyecup at 0, 0.5, 1, 2dpc, colored by time point. e, Fraction of expressing cells (dot size) and normalized expression in expressing cells (dot color) of selected marker genes (columns) across 12 identified cell types in the eyecup (rows). f, Endothelial cells collected from mouse eyecup, colored by expression of genes associated with choroidal Kdrhi (left) and Kdrlo (right) cells20.
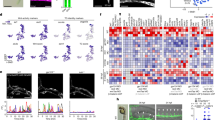
Extended Data Fig. 3 Visualizing immune cells in the retina and eyecup.
a, Fraction of expressing cells (dot size) and mean expression levels in expressing cells (dot color) of selected marker genes (columns) across 18 identified immune cell subsets (rows). b,c, Representative IHC on 0 and 0.5dpc retinal sections showing LY6G+ neutrophils that express MMP9 (red) and localize proximal to the optic nerve head (ONH) (n = 3 mice per time point) (b), and on 2dpc retinal whole-mounts from CCR2RFP/+ mice showing CCR2-RFP+ cells (red) in the GCL and IPL (c). Higher magnification inset of the outlined region in c is shown in Fig. 2d (n = 2). Scale bar, 50 μm. d, Representative IHC on 0dpc eyecup sections for IA-IE (MHC-II, magenta) and CD206 (green). Scale bar, 50 μm. Inset shows double positive cells, scale bar, 10 μm. BF, brightfield (n = 2-3 mice, representative of two independent experiments). e, Representative image of smFISH on 2dpc sections showing Ms4a7+Ccr2- cells (green) in the eyecup. This is a subset of the image presented in Fig. 5h. Scale bar, 50 μm. f,g, Representative images of IHC on eyecup whole-mounts (f) from CCR2RFP/+ mice showing that the majority of CCR2-RFP+ cells (red) in naïve eyecup are located posterior to the RPE, at the level of the choroid. ZO-1 (green) depicts tight junctions between individual RPE cells and nuclei are stained with Hoechst (blue) (n = 2 mice, representative of three independent experiments), and sections from 2dpc (g) showing that infiltrating CCR2-RFP+ cells (red) express the macrophage marker, IBA1 (green) (n = 2 mice). Arrowhead points to the RPE. Scale bars, 50 μm.
Extended Data Fig. 4 Glial reactivation and microglial proliferation after ONC.
a, Distribution of expression of genes differentially expressed between Müller glia (MG; n = 54,565 cells) and astrocytes (n = 18,959 cells). b, Fraction of expressing microglia cells (dot size) and mean expression levels in expressing cells (dot color) of microglia signature genes across time (0dpc (n = 4,479 cells); 0.5dpc (n = 2,136); 1dpc (n = 1,712); 2dpc (n = 3,616); 4dpc (n = 2,950), 7dpc (n = 4,433); 14dpc (n = 5,233)). c,d, Representative smFISH on 4dpc retinal sections (c) showing P2ry12+ microglia (green) expressing Mki67 (red) (n = 3 mice), and IHC on 4dpc retinal whole-mounts (d) of KI67+ (red) IBA1+ microglia (green) (n = 3 mice). Scale bars, 50 μm. e, Fraction of cells (dot size) in each MG subset, and mean expression level in expressing cells (dot color) of light and excitotoxin injury-induced genes31. f, Distribution of expression scores for signature genes of pan-reactive astrocytes35 across the astrocyte and astroIFN subsets throughout the time course (Astrocyte/Reactive astro: 0dpc (n = 1,009/17 cells); 0.5dpc (n = 586/299); 1dpc (n = 1,780/767); 2dpc (n = 2,268/439); 4dpc (n = 2,671/652), 7dpc (n = 2,051/396); 14dpc (n = 368/55)). g,h, Distribution of expression scores for Cluster 4 (‘inflammatory‘) signature from LPS-induced astrocytes33 (g) and for a tissue dissociation signature (h) on astrocytes from LPS-induced neuroinflammation33 classified to ONC astrocyte subsets (left) and across the ONC astrocyte subsets (right) (Student’s t-test, one sided). Boxplots in a and f denote medians and IQRs; whiskers are the lowest datum still within 1.5 IQR of the lower quartile and the highest datum still within 1.5 IQR of the upper quartile.
Extended Data Fig. 5 ONC-induced expression changes in the RPE.
a, Representative image of IHC on ocular sections, depicting the RPE with specific staining (RPE65, green) in relation to the retina and optic nerve. Scale bar, 50 μm. b, Fraction of expressing cells (dot size) and normalized expression level in expressing cells (dot color) of differentially expressed genes in the RPE between 0, 0.5, 1 and 2dpc. c,d, Distribution of expression of Plin2 (c) and Nog (d) on RPE cells by time (left; 0dpc (n = 2,605 cells); 0.5dpc (n = 1,746); 1dpc (n = 631); 2dpc (n = 3,031)) and across subsets (right). Boxplots denote medians and IQRs; whiskers are the lowest datum still within 1.5 IQR of the lower quartile and the highest datum still within 1.5 IQR of the upper quartile. e, Representative image of smFISH on 0 and 2dpc RPE for Nog (green) (n = 3 mice per time point). Scale bar, 50 μm.
Extended Data Fig. 6 Dynamics of resident and infiltrating mononuclear phagocytes in the retina, and conserved signature of Gpnmb+ macrophages.
a, Changes in frequency of Ms4a7+MHC-IIhi, Ms4a7+MHC-IIlo and Gpnmb+ macrophages across 0, 0.5, 1, 2, 4, 7, and 14dpc. b, Distribution of expression of genes associated with MHC-IIhi and MHC-IIlo BAMs23 on retinal Ms4a7+MHC-IIhi (n = 741) and Ms4a7+MHC-IIlo macrophages (n = 1,406). Boxplots denote medians and IQRs; whiskers are the lowest datum still within 1.5 IQR of the lower quartile and the highest datum still within 1.5 IQR of the upper quartile. c, Representative image of smFISH on 7dpc retinal for Gpnmb (blue), Ms4a7 (red) and Ptprc (green) (n = 3 mice). Scale bar, 25 μm. d, Distribution of expression of Gpnmb+ macrophage marker gene orthologs in immune cell scRNA-seq data from chronic lesions in human patients with multiple sclerosis47. e, Fraction of expressing cells (dot size) and normalized expression level in expressing cells (dot color) of the top 30 differentially expressed genes of Gpnmb+ macrophages across the RPE cell clusters. f, Representative IHC on uninjured (0 dpc) retinal whole-mounts showing perivascular and CB-adjacent IA-IE+ (green) cells. Blood vessels are labeled with IB4 (red) (representative of five independent experiments with n = 1-3 mice each). The middle panel shows a zoomed-in image of the perivascular macrophage in the inset of the left image. Scale bars, 50 μm g, Force-directed layout view of monocyte and macrophage subsets with optimal transport analysis. Each dot is a cell, color-coded by ancestor probabilities for Ms4a7+MHC-IIhi (top row) and Ms4a7+MHC-IIlo macrophages (bottom row), as estimated by Waddington-OT88.
Extended Data Fig. 7 A co-regulated IFN-response program in the injured retina.
a, Distribution of expression of genes differentially expressed in microgliaIFN by microglia repopulating the brain after depletion (Repop) compared to microglia from control brain55 (Mann-Whitney U test, two sided). b, Distribution of Ifit3 expression across time in MG (top) and astrocytes (bottom) across subsets. c, Distribution of a signature score of Cluster 8 (‘ISG‘)astrocytes from LPS-induced neuroinflammation33 in the LPS-induced astrocytes as classified to the ONC astrocyte subsets (left) and across the ONC astrocyte subsets (right) (Student’s t-test, one sided). d, Off-diagonal panels: Comparison of overall expression scores (y and x axes) for each cell component of MCP1 (rows and columns, labels on diagonal) across the samples; lines correspond to linear fit. Pearson correlation (r) and significance (***p < 0.001; Pearson correlation t-test, one-sided) are shown in the panels above the diagonal. Diagonal panels: Distribution of overall expression scores for each cell type component, with kernel density estimates91. e, Distribution of MCP1 expression scores by astrocytes (n = 18,956 cells), microglia (n = 20,000 cells) and MG (n = 19,998 cells) across 0, 0.5, 1, 2, 4, 7 and 14dpc. Numbers above violins indicate median expression scores. Boxplots in b, e denote medians and IQRs; whiskers are the lowest datum still within 1.5 IQR of the lower quartile and the highest datum still within 1.5 IQR of the upper quartile.
Extended Data Fig. 8 Schematic model summarizing the tissue dynamics along the response to ONC.
An inflammatory cascade is initiated early after injury (phase I), prior to RGC death, with activation of resident glia involving chemokine signals for leukocyte infiltration, which is observed in the oBRB and inner retina. At intermediate time points (phase II), concurrent with the peak rate of RGC death, infiltrating monocytes differentiate into distinct macrophage subsets, including Ms4a7+MHC-IIhi, Ms4a7+MHC-IIlo and Gpnmb+ macrophages. In parallel, during this phase, a synchronous interferon response program is induced in astrocytes, Müller glia and microglia. The latter also express a disease-associated microglia signature, which overlaps with that of Gpnmb+ macrophages. Finally, at 1-2wpc (phase III), glial cell proportions begin to return to their baseline levels, with enriched interactions among them including TGFβ signaling, collectively indicating restoration of homeostasis.
Supplementary information
Supplementary Information
Supplementary Notes 1–4, Figs. 1–5 and Table titles.
Supplementary Tables 1–4
Supplementary Tables 1–4. Titles are in the Supplementary Information file.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Benhar, I., Ding, J., Yan, W. et al. Temporal single-cell atlas of non-neuronal retinal cells reveals dynamic, coordinated multicellular responses to central nervous system injury. Nat Immunol 24, 700–713 (2023). https://doi.org/10.1038/s41590-023-01437-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-023-01437-w
This article is cited by
-
Intercellular communication atlas reveals Oprm1 as a neuroprotective factor for retinal ganglion cells
Nature Communications (2024)
-
Immune stimulation recruits a subset of pro-regenerative macrophages to the retina that promotes axonal regrowth of injured neurons
Acta Neuropathologica Communications (2023)