Abstract
Patients with loss of function in the gene encoding the master regulator of central tolerance AIRE suffer from a devastating disorder called autoimmune polyendocrine syndrome type 1 (APS-1), characterized by a spectrum of autoimmune diseases and severe mucocutaneous candidiasis. Although the key mechanisms underlying the development of autoimmunity in patients with APS-1 are well established, the underlying cause of the increased susceptibility to Candida albicans infection remains less understood. Here, we show that Aire+MHCII+ type 3 innate lymphoid cells (ILC3s) could sense, internalize and present C. albicans and had a critical role in the induction of Candida-specific T helper 17 (TH17) cell clones. Extrathymic Rorc-Cre-mediated deletion of Aire resulted in impaired generation of Candida-specific TH17 cells and subsequent overgrowth of C. albicans in the mucosal tissues. Collectively, our observations identify a previously unrecognized regulatory mechanism for effective defense responses against fungal infections.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Anderson, M. S. et al. Projection of an immunological self shadow within the thymus by the aire protein. Science 298, 1395–1401 (2002).
Klein, L., Kyewski, B., Allen, P. M. & Hogquist, K. A. Positive and negative selection of the T cell repertoire: what thymocytes see (and don’t see). Nat. Rev. Immunol. 14, 377–391 (2014).
Liston, A., Lesage, S., Wilson, J., Peltonen, L. & Goodnow, C. C. Aire regulates negative selection of organ-specific T cells. Nat. Immunol. 4, 350–354 (2003).
Aschenbrenner, K. et al. Selection of Foxp3+ regulatory T cells specific for self antigen expressed and presented by Aire+ medullary thymic epithelial cells. Nat. Immunol. 8, 351–358 (2007).
Malchow, S. et al. Aire enforces immune tolerance by directing autoreactive T cells into the regulatory T cell lineage. Immunity 44, 1102–1113 (2016).
Abramson, J. & Husebye, E. S. Autoimmune regulator and self-tolerance: molecular and clinical aspects. Immunol. Rev. 271, 127–140 (2016).
Husebye, E. S., Anderson, M. S. & Kämpe, O. Autoimmune polyendocrine syndromes. N. Engl. J. Med. 378, 2543–2544 (2018).
Bruserud, Ø. et al. A longitudinal follow-up of autoimmune polyendocrine syndrome type 1. J. Clin. Endocrinol. Metab. 101, 2975–2983 (2016).
Perheentupa, J. Autoimmune polyendocrinopathy-candidiasis-ectodermal dystrophy. J. Clin. Endocrinol. Metab. 91, 2843–2850 (2006).
Okada, S. et al. IMMUNODEFICIENCIES. Impairment of immunity to Candida and Mycobacterium in humans with bi-allelic RORC mutations. Science 349, 606–613 (2015).
Puel, A. et al. Chronic mucocutaneous candidiasis in humans with inborn errors of interleukin-17 immunity. Science 332, 65–68 (2011).
Milner, J. D. et al. Impaired T(H)17 cell differentiation in subjects with autosomal dominant hyper-IgE syndrome. Nature 452, 773–776 (2008).
Ferwerda, B. et al. Human dectin-1 deficiency and mucocutaneous fungal infections. N. Engl. J. Med. 361, 1760–1767 (2009).
Glocker, E. O. et al. A homozygous CARD9 mutation in a family with susceptibility to fungal infections. N. Engl. J. Med. 361, 1727–1735 (2009).
Liu, L. et al. Gain-of-function human STAT1 mutations impair IL-17 immunity and underlie chronic mucocutaneous candidiasis. J. Exp. Med. 208, 1635–1648 (2011).
Conti, H. R. & Gaffen, S. L. IL-17-mediated immunity to the opportunistic fungal pathogen Candida albicans. J. Immunol. 195, 780–788 (2015).
Kisand, K. et al. Chronic mucocutaneous candidiasis in APECED or thymoma patients correlates with autoimmunity to Th17-associated cytokines. J. Exp. Med. 207, 299–308 (2010).
Puel, A. et al. Autoantibodies against IL-17A, IL-17F, and IL-22 in patients with chronic mucocutaneous candidiasis and autoimmune polyendocrine syndrome type I. J. Exp. Med. 207, 291–297 (2010).
Yamano, T. et al. Aire-expressing ILC3-like cells in the lymph node display potent APC features. J. Exp. Med. 216, 1027–1037 (2019).
Schlitzer, A. et al. IRF4 transcription factor-dependent CD11b+ dendritic cells in human and mouse control mucosal IL-17 cytokine responses. Immunity 38, 970–983 (2013).
Jouault, T. et al. Candida albicans phospholipomannan is sensed through toll-like receptors. J. Infect. Dis. 188, 165–172 (2003).
Blasi, E. et al. Biological importance of the two Toll-like receptors, TLR2 and TLR4, in macrophage response to infection with Candida albicans. FEMS Immunol. Med. Microbiol. 44, 69–79 (2005).
Brown, G. D. et al. Dectin-1 mediates the biological effects of beta-glucans. J. Exp. Med. 197, 1119–1124 (2003).
Gantner, B. N., Simmons, R. M. & Underhill, D. M. Dectin-1 mediates macrophage recognition of Candida albicans yeast but not filaments. EMBO J. 24, 1277–1286 (2005).
Kohatsu, L., Hsu, D. K., Jegalian, A. G., Liu, F. T. & Baum, L. G. Galectin-3 induces death of Candida species expressing specific beta-1,2-linked mannans. J. Immunol. 177, 4718–4726 (2006).
Jouault, T. et al. Specific recognition of Candida albicans by macrophages requires galectin-3 to discriminate Saccharomyces cerevisiae and needs association with TLR2 for signaling. J. Immunol. 177, 4679–4687 (2006).
Veldhoen, M., Hocking, R. J., Atkins, C. J., Locksley, R. M. & Stockinger, B. TGFbeta in the context of an inflammatory cytokine milieu supports de novo differentiation of IL-17-producing T cells. Immunity 24, 179–189 (2006).
Bettelli, E. et al. Reciprocal developmental pathways for the generation of pathogenic effector TH17 and regulatory T cells. Nature 441, 235–238 (2006).
Mangan, P. R. et al. Transforming growth factor-beta induces development of the TH17 lineage. Nature 441, 231–234 (2006).
Dobeš, J. et al. A novel conditional Aire allele enables cell-specific ablation of the immune tolerance regulator Aire. Eur. J. Immunol. 48, 546–548 (2018).
Eberl, G. & Littman, D. R. Thymic origin of intestinal alphabeta T cells revealed by fate mapping of RORgammat+ cells. Science 305, 248–251 (2004).
Jiang, T. T. et al. Commensal fungi recapitulate the protective benefits of intestinal bacteria. Cell Host Microbe 22, 809–816.e804 (2017).
Shao, T. Y. et al. Commensal Candida albicans positively calibrates systemic Th17 immunological responses. Cell Host Microbe 25, 404–417.e406 (2019).
Sonnenberg, G. F., Monticelli, L. A., Elloso, M. M., Fouser, L. A. & Artis, D. CD4+ lymphoid tissue-inducer cells promote innate immunity in the gut. Immunity 34, 122–134 (2011).
Solis, N. V. & Filler, S. G. Mouse model of oropharyngeal candidiasis. Nat. Protoc. 7, 637–642 (2012).
Altieri, D. C. Survivin: the inconvenient IAP. Semin. Cell Dev. Biol. 39, 91–96 (2015).
DiToro, D. et al. Insulin-like growth factors are key regulators of T helper 17 regulatory T cell balance in autoimmunity. Immunity 52, 650–667 (2020).
Sonnenberg, G. F. & Hepworth, M. R. Functional interactions between innate lymphoid cells and adaptive immunity. Nat. Rev. Immunol. 19, 599–613 (2019).
Hepworth, M. R. et al. Innate lymphoid cells regulate CD4+ T-cell responses to intestinal commensal bacteria. Nature 498, 113–117 (2013).
Hepworth, M. R. et al. Immune tolerance. Group 3 innate lymphoid cells mediate intestinal selection of commensal bacteria-specific CD4+ T cells. Science 348, 1031–1035 (2015).
Melo-Gonzalez, F. et al. Antigen-presenting ILC3 regulate T cell-dependent IgA responses to colonic mucosal bacteria. J. Exp. Med. 216, 728–742 (2019).
Oliphant, C. J. et al. MHCII-mediated dialog between group 2 innate lymphoid cells and CD4+ T cells potentiates type 2 immunity and promotes parasitic helminth expulsion. Immunity 41, 283–295 (2014).
von Burg, N. et al. Activated group 3 innate lymphoid cells promote T-cell-mediated immune responses. Proc. Natl Acad. Sci. U S A 111, 12835–12840 (2014).
Kärner, J. et al. Anti-cytokine autoantibodies suggest pathogenetic links with autoimmune regulator deficiency in humans and mice. Clin. Exp. Immunol. 171, 263–272 (2013).
Dobeš, J. et al. Gastrointestinal autoimmunity associated with loss of central tolerance to enteric α-defensins. Gastroenterology 149, 139–150 (2015).
Gavanescu, I., Kessler, B., Ploegh, H., Benoist, C. & Mathis, D. Loss of Aire-dependent thymic expression of a peripheral tissue antigen renders it a target of autoimmunity. Proc. Natl Acad. Sci. U S A 104, 4583–4587 (2007).
Ma, C. S. et al. Deficiency of Th17 cells in hyper IgE syndrome due to mutations in STAT3. J. Exp. Med. 205, 1551–1557 (2008).
Minegishi, Y. et al. Human tyrosine kinase 2 deficiency reveals its requisite roles in multiple cytokine signals involved in innate and acquired immunity. Immunity 25, 745–755 (2006).
Break, T. J. et al. Aberrant type 1 immunity drives susceptibility to mucosal fungal infections. Science 371, eaay5731 (2021).
Wang, J. et al. Single-cell multiomics defines tolerogenic extrathymic Aire-expressing populations with unique homology to thymic epithelium. Sci. Immunol. 6, eabl5053 (2021).
Lyu, M. et al. ILC3s select for RORγt+ Tregs and establish tolerance to intestinal microbiota. Preprint at bioRxiv, https://doi.org/10.1101/2022.04.25.489463 (2022).
Jiang, W., Anderson, M. S., Bronson, R., Mathis, D. & Benoist, C. Modifier loci condition autoimmunity provoked by Aire deficiency. J. Exp. Med. 202, 805–815 (2005).
Gardner, J. M. et al. Deletional tolerance mediated by extrathymic Aire-expressing cells. Science 321, 843–847 (2008).
Taylor, P. R. et al. Dectin-1 is required for beta-glucan recognition and control of fungal infection. Nat. Immunol. 8, 31–38 (2007).
Gordon, J. et al. Specific expression of lacZ and cre recombinase in fetal thymic epithelial cells by multiplex gene targeting at the Foxn1 locus. BMC Dev. Biol. 7, 69 (2007).
Colnot, C., Fowlis, D., Ripoche, M. A., Bouchaert, I. & Poirier, F. Embryonic implantation in galectin 1/galectin 3 double mutant mice. Dev. Dyn. 211, 306–313 (1998).
Barnden, M. J., Allison, J., Heath, W. R. & Carbone, F. R. Defective TCR expression in transgenic mice constructed using cDNA-based alpha- and beta-chain genes under the control of heterologous regulatory elements. Immunol. Cell Biol. 76, 34–40 (1998).
Mombaerts, P. et al. RAG-1-deficient mice have no mature B and T lymphocytes. Cell 68, 869–877 (1992).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Fonzi, W. A. & Irwin, M. Y. Isogenic strain construction and gene mapping in Candida albicans. Genetics 134, 717–728 (1993).
Igyártó, B. Z. et al. Skin-resident murine dendritic cell subsets promote distinct and opposing antigen-specific T helper cell responses. Immunity 35, 260–272 (2011).
Jaitin, D. A. et al. Massively parallel single-cell RNA-seq for marker-free decomposition of tissues into cell types. Science 343, 776–779 (2014).
Kohen, R. et al. UTAP: user-friendly transcriptome analysis pipeline. BMC Bioinf. 20, 154 (2019).
Moon, J. J. et al. Naive CD4+ T cell frequency varies for different epitopes and predicts repertoire diversity and response magnitude. Immunity 27, 203–213 (2007).
Pfaffl, M. W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 29, e45 (2001).
Acknowledgements
Research in the Abramson laboratory is kindly supported by the European Research Council (ERC-2016-CoG-724821); Israel Science Foundation (1796/16 and 1819/21); Chan Zuckerberg Initiative; Bill and Marika Glied and Family Fund, Binational Science Foundation; Pasteur-Weizmann Delegation; Enoch Foundation, Ruth and Samuel David Gameroff Family Foundation; and Erica Drake Fund and Lilly Fulop Fund for Multiple Sclerosis Research. J.D. was supported by the Dean of Faculty Fellowship by Weizmann Institute of Science and by the Weizmann Institute of Science – Czech Academy of Sciences Bilateral Fellowship by Czech Academy of Sciences. J.D., K.K., H.B. and E.V. are also supported by Charles University PRIMUS grant (Primus/21/MED/003) and Czech Science Foundation JUNIOR STAR grant (21-22435M). B.E.O. and E.S.H. are supported by the K.G. Jebsen Center for autoimmune disorders, the Norwegian Research Council, the Novo Nordisk Foundation and the Regional Health Authorities of Western Norway. D.F. is supported by the Grant Agency of the Czech Republic (17-25365S).
Author information
Authors and Affiliations
Contributions
J.A. and J.D. conceived the project, designed the experiments and wrote the manuscript; J.D. performed the experiments and analyzed the data; J.A. supervised the study; A.B., Y. Goldfarb, O.B.-N., N.K., Y. Gruper, T.G., I.Z., H.B., K.K., E.V., B.E.O. and E.S.H. assisted at different aspects of the study, including data or sample acquisition and analysis. D.F. contributed by the essential reagent. L.S.B. and Z.S. performed the two-photon microscopy and analyzed the data.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Immunology thanks the anonymous reviewers for their contribution to the peer review of this work. Ioana Visan was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1
a–c, GO enrichment analysis of upregulated differentially expressed genes from unstimulated vs. C. albicans- stimulated ILC3 subsets including cILC3s (related to Fig. 2a) (a); MHCII+ ILC3s (related to Fig. 2b) (b); Aire+ ILC3s (related to Fig. 2c) (c). d, Heatmap of Pearson correlation according to gene expression values between individual samples analyzed in Fig. 2a-c.
Extended Data Fig. 2
a, Imaging flow cytometry analyzing the physical interaction between HKCA with DCs or with Aire+ ILC3s. CD11c+ DCs or Lineage negative cells were isolated from mouse pLNs using MACS-beads depletion and then incubated with CPD-stained HKCA for 90 minutes. Samples were stained for Aire, MHCII, CD11c, Lamp1 (lysosomal marker) and DAPI and analyzed by Imaging flow cytometry. In both, Aire+ ILC3 cells (Aire+) and DCs (CD11c+) HKCA associate with lysosomes (Lamp1). Representative images out of five independent repetitions of the experiment are shown. b, Comparison of the capacity of different immune subsets (B cells, Ly6C+ MNPs, CD11b+ MNPs, CD11c+ MNPs and Aire+ ILC3s) to endocytose CPD-labeled HKCA in an in vitro endocytosis assay. Subsets were FACS-sorted and incubated with HKCA-CPD for three hours. The frequency of cells that have successfully internalized HKCA-CPD was then assessed by flow cytometry (mean ± SD; n = 3). c,d, Comparison of the capacity of either wild-type or Galectin-3/Dectin-1-double knockout (dKO) Aire+ ILC3s (c) or DCs (d) to endocytose CPD-labeled HKCA at 37 °C or at 4 °C. Corresponding populations were isolated from WT or dKO mice and kept either at 37 °C or at 4 °C. The internalization of HKCA-CPD by the cells was measured by FACS (mean ± SD; n = 3). e, Representative gating strategy of APC populations related to Fig. 3c–g. f, FACS analysis of in vitro assay related to Fig. 3c. g, Graphical summary of an experimental setting relevant for data shown in h. Wild-type mice were intravenously stimulated by HKCA in specified timepoints and APC population were isolated using gating strategy depicted in e. h, FACS analysis of in vitro assay related to Fig. 3d–f assessing the antigen presentation of the corresponding cell subsets.
Extended Data Fig. 3
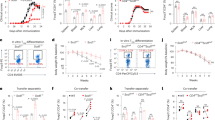
a, FACS analysis validating that Aire protein expression in the entire LN-resident cell compartment is exclusively restricted to a CD45 positive, lineage negative (Lin 1: TCR-β, CD19, Gr1, F4/80, CD11b, CD11c, NK1.1; Lin 2: CD3, B220), NKp46-negative, MHCII positive, Rorγt positive cell subset – previously identified as Aire+ ILC3s. b, Back-gating FACS analysis of Aire-expressing cells identified in a. c, Representative flow cytometry dot plots showing the frequencies of transferred CD45.1+ OT-II T cell (red gate) vs. CD45.1/CD45.2 double positive control T cell (blue gate) populations 2 days after the transfer. d, Statistical analysis of ratios related to b (n = 5 per group, mean ± SD, two-tailed Student’s t-test) of OT-II vs. control T two days after the transfer. e,f, Survival curves of WT (Aire+/+) and knockout (Aire–/–) mice (n ≥ 10 mice per group) on either on NOD (e) and C57Bl/6 (f) genetic background after systemic challenge with live C. albicans. Mice were i.v. injected every second day by heat-killed C. albicans (HKCA) for the duration of three weeks. Subsequently, mice were infected by alive C. albicans and monitored for survival. Long-rank (Mantel-Cox) test was used to calculate the indicated P-value. g,h, Quantitative PCR analysis assessing the presence of C. albicans-specific DNA in the liver (g) and small intestine (h) from Rorc-Cre– Airefl/fl (WT) and Rorc-Cre+Airefl/fl (ILC3ΔAire) mice (n = 6, mean ± SD, two-tailed Student’s t-test). i,j, ELISA assessing the amount of IL-17 (i) or (IL-22) (j) autoantibodies in the sera of 8-week old untreated Aire+/+ vs. Aire–/– mice on NOD and C57Bl/6 (B6) genetic background (n = 6 per group, two-tailed Student’s t-test). Data are shown as mean of optical density ± SD. Data are shown as mean of optical density ± SD. P-value indicators: *** = P-value < 0.0001, ** = P-value < 0.001, ns = not significant.
Extended Data Fig. 4
a, Experimental outline relevant to data shown in Fig. 6a, b: Wild-type mice were orally colonized by C. albicans and analyzed in indicated time points. b, Representative FACS gating strategy of Als1-Tet+ T cells. c, Experimental outline relevant to data shown in (d) and (e) WT (Aire+/+) and Aire−/− were orally colonized by C. albicans and analyzed after two weeks. d, Representative FACS plot of Als1-Tet+ T cells. Cells were isolated from pLNs and spleens of mice described in (c). Counts of Als1-Tet+ cells are highlighted in red rectangles (left panel). Statistical analysis of the same representative experiment showing the total counts (mean ± SD, two-tailed Student’s t-test, n = 6). Representative experiment is shown. e, Quantitative PCR analysis assessing the presence of C. albicans-specific DNA in the ileal part of small intestine from WT and Aire-deficient mice (n = 5, mean ± SD, two-tailed Student’s t-test). f, Experimental outline relevant to data shown in (g) and (h). Bone marrow (BM) chimeras restricting Aire expression either to hematopoietic (Aire+/+ BM→Aire–/–) or stromal compartment (Aire–/– BM→Aire+/+) were generated by reciprocal BM transfer to recipient mice after 900 rad whole-body irradiation. Six weeks after the BM transfer, the mice were orally colonized by C. albicans and analyzed after two weeks. g, Representative FACS plot of Als1-Tet+ T cells. Counts of Als1-Tet+ cells are highlighted in red rectangles (left panel). Statistical analysis of the same representative experiment showing the total counts (n = 6, mean ± SD, two-tailed Student’s t-test). Representative experiment is shown. h, Quantitative PCR analysis assessing the presence of C. albicans-specific DNA in the ileal part of small intestine from reciprocal bone marrow chimeras (n = 6, mean ± SD, two-tailed Student’s t-test). i, Experimental outline relevant to data shown in j and k. CD90-disperate chimeras were created by adoptive transfer of T cells and B-lymphocytes from CD90.1 mice to Rag1−/− (CD90.2) recipients and let to proliferate for 2 months. Mice were treated by anti-CD90.2 or isotype control antibody prior the C. albicans oral colonization and then each third day and analyzed after two weeks. j, Representative FACS plot of Als1-Tet+ T cells. Counts of tetramer positive cells are highlighted in red rectangles (left panel). Statistical analysis of the same representative experiment showing the total counts (n = 6, mean ± SD, two-tailed Student’s t-test). Representative experiment is shown. k, Quantitative PCR analysis assessing the presence of C. albicans-specific DNA in the ileal part of small intestine from CD90-disparate chimeras (n = 6, mean ± SD, two-tailed Student’s t-test). P-value indicators: *** = P-value < 0.0001, ** = P-value < 0.001, * = P-value < 0.05, ns = not significant.
Extended Data Fig. 5
a–e, Flow cytometry analysis of C albicans-specific T cells (using Als1-Tet) after oral colonization. Aire whole-body knockout mice (Aire−/−), their wild-type littermates (Aire+/+), WT (Rorc-Cre–Airefl/fl) and ILC3ΔAire (Rorc-Cre+Airefl/fl) were orally colonized by C. albicans and analyzed after 24 hours (left panel) or two weeks (right panel). a, Representative FACS plot of Als1-Tet+ T cells. Cells were isolated from pLNs and spleens (SLO), oral mucosa, esophagus and intestine of mice described above 24 hours (left panel) or 2 weeks (right panel) post C. albicans colonization. Counts of Als1-Tet+ cells are highlighted in red rectangles. b–e, Statistical analysis of the Als1-Tet+ cells counts (n = 6, mean ± SD, two-tailed Student’s t-test in SLO (b), oral mucosa (c), esophagus (d) and small intestine (e) 24 hours (left panel) or 2 weeks (right panel) post C. albicans colonization (n = 6, mean ± SD, two-tailed Student’s t-test). P-value indicators: *** = P-value < 0.0001, ** = P-value < 0.001, * = P-value < 0.05, ns = not significant.
Extended Data Fig. 6
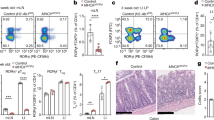
a, Representative FACS gating strategy of Roryt+ CD4+ TCR-β+ TH17 cells in pLNs two weeks after C. albicans colonization of Rorc-Cre– Airefl/fl (WT) and Rorc-Cre+Airefl/fl (ILC3ΔAire) mice. b, Statistical analysis of related to a). The plot is showing the frequency from parent gate (n = 6, mean ± SD, two-tailed Student’s t-test). c, Representative FACS gating strategy of Roryt+ CD4+ TCR-β+ TH17 cells in mesenteric lymph nodes (mLN) two weeks after C. albicans colonization of Rorc-Cre– Airefl/fl (WT) and Rorc-Cre+Airefl/fl (ILC3ΔAire) mice. d, Statistical analysis of related to c). The plot is showing the frequency from parent gate (n = 6, mean ± SD, two-tailed Student’s t-test). e, Representative FACS gating strategy of Roryt+ CD4+ TCR-β+ TH17 cells in lamina propria two weeks after C. albicans colonization of Rorc-Cre– Airefl/fl (WT) and Rorc-Cre+Airefl/fl (ILC3ΔAire) mice. f), Statistical analysis of related to e). The plot is showing the frequency from parent gate (n = 6, mean ± SD, two-tailed Student’s t-test). P-value indicators: ** =P-value < 0.001, * = P-value < 0.05, ns = not significant.
Extended Data Fig. 7
a–d, Colony forming units (CFU)-based assay determining the overgrowth of C albicans 1 day (left panel) and 14 days (right panel) after the oral colonization. The tissues were isolated from Aire whole-body knockout mice (Aire−/−), their wild-type littermates (Aire+/+), Rorc-Cre– Airefl/fl (WT) and Rorc-Cre+Airefl/fl (ILC3ΔAire) mice. CFU was determined by plating the lysates obtained from the kidney (a), oral cavity (b), esophagus (c), small intestine (d) (n = 6, mean ± SD, two-tailed Student’s t-test). P-value indicators: *** = P-value < 0.0001, ** = P-value < 0.001, * = P-value < 0.05, ns = not significant.
Extended Data Fig. 8
a, Experimental outline relevant to data shown in b–c. Rorc-Cre– Airefl/fl (WT) and Rorc-Cre+Airefl/fl (ILC3ΔAire) were first primed by repeated injections of heat-killed C. albicans (HKCA) for two weeks, then exposed to prolonged protocol of oropharyngeal candidiasis and analyzed five days later. b, Representative FACS plot of Als1-Tet+ T cells. Cells were isolated from pLNs and spleens of mice described in a. Counts of Als1-Tet+ cells are highlighted in red rectangles (left panel). Statistical analysis of the same representative experiment showing the total counts of Als1-Tet+ cells (n = 6, mean ± SD, two-tailed Student’s t-test). Representative experiment is shown. c, Representative FACS plot of Als1-Tet+ T cells. Cells were isolated from tongue mucosae of mice described in a. Counts of Als1-Tet+ cells are highlighted in red rectangles (left panel). Statistical analysis of the same representative experiment showing the total counts of Als1-Tet+ cells (n = 6, mean ± SD, two-tailed Student’s t-test). Representative experiment is shown.
Extended Data Fig. 9
a, Representative FACS-sorting strategy of Aire+ ILC3s from reporter AireGFP mice. The plots are showing the frequency from parent gates. Lineage: CD3, CD19, CD11c, CD11b, N.K.1, F4/80, Gr1, B220. b, Back-gating of AireGFP+ cells.
Extended Data Fig. 10
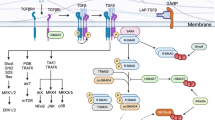
a, Experimental outline relevant to data shown in b and c. WT (Rorc-Cre–Airefl/fl) and ILC3ΔAire (Rorc-Cre+Airefl/fl) mice were transferred with naïve OT-II CD4+ T cells and subsequently injected with HKCA-OVA four times during a single week. b, Statistical analysis of the frequencies of OT-II T cells subtypes (n = 4, mean ± SD, two-tailed Student’s t-test). c, GO enrichment analysis of upregulated differentially expressed genes from RorcGFP+ OT-II T cells vs Non-proliferating OT-II T cells comparison. d, Graphical model summarizing the role of Aire+ ILC3s in the induction of Candida-specific TH17 response. While the immediate immune response to C. albicans infection is dominated by neutrophils, monocytes and macrophages, Aire+ ILC3s become essential in the later phase, as they facilitate clonal expansion of the primed TH17 cell clones in the LN. The expanded candida-specific TH17 clones subsequently limit C. albicans overgrowth at mucosal surfaces and its dissemination into epithelial tissues; e, Scheme illustrating the putative mechanism through which Aire+ ILC3 cells induce the expansion of candida-specific TH17 clones.
Supplementary information
Supplementary Information
Supplementary Tables 1 and 2.
Rights and permissions
About this article
Cite this article
Dobeš, J., Ben-Nun, O., Binyamin, A. et al. Extrathymic expression of Aire controls the induction of effective TH17 cell-mediated immune response to Candida albicans. Nat Immunol 23, 1098–1108 (2022). https://doi.org/10.1038/s41590-022-01247-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-022-01247-6
This article is cited by
-
The emerging family of RORγt+ antigen-presenting cells
Nature Reviews Immunology (2024)
-
The roles of lncRNAs in Th17-associated diseases, with special focus on JAK/STAT signaling pathway
Clinical and Experimental Medicine (2023)
-
Extrathymic Aire primes Candida-specific TH17 cells
Nature Immunology (2022)
-
How regulatory T cells are primed to aid tolerance of gut bacteria
Nature (2022)
-
Novel antigen-presenting cell imparts Treg-dependent tolerance to gut microbiota
Nature (2022)