Abstract
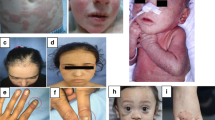
We report a pleiotropic disease due to loss-of-function mutations in RHBDF2, the gene encoding iRHOM2, in two kindreds with recurrent infections in different organs. One patient had recurrent pneumonia but no colon involvement, another had recurrent infectious hemorrhagic colitis but no lung involvement and the other two experienced recurrent respiratory infections. Loss of iRHOM2, a rhomboid superfamily member that regulates the ADAM17 metalloproteinase, caused defective ADAM17-dependent cleavage and release of cytokines, including tumor-necrosis factor and amphiregulin. To understand the diverse clinical phenotypes, we challenged Rhbdf2−/− mice with Pseudomonas aeruginosa by nasal gavage and observed more severe pneumonia, whereas infection with Citrobacter rodentium caused worse inflammatory colitis than in wild-type mice. The fecal microbiota in the colitis patient had characteristic oral species that can predispose to colitis. Thus, a human immunodeficiency arising from iRHOM2 deficiency causes divergent disease phenotypes that can involve the local microbial environment.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The RNA-seq data have been deposited to the GEO database under the accession number GSE184877. WGS/WES data for the kindreds of P1 and P2 were submitted to the National Center for Biotechnology Information (NCBI) database of Genotypes and Phenotypes (dbGaP) (accession no., phs002478.v1.p1). Source data are provided with this paper.
References
Williams, R. A., Mamotte, C. D. & Burnett, J. R. Phenylketonuria: an inborn error of phenylalanine metabolism. Clin. Biochem. Rev. 29, 31–41 (2008).
Freeman, M. The rhomboid-like superfamily: molecular mechanisms and biological roles. Annu. Rev. Cell Dev. Biol. 30, 235–254 (2014).
Lambrecht, B. N., Vanderkerken, M. & Hammad, H. The emerging role of ADAM metalloproteinases in immunity. Nat. Rev. Immunol. 18, 745–758 (2018).
Dulloo, I., Muliyil, S. & Freeman, M. The molecular, cellular and pathophysiological roles of iRhom pseudoproteases. Open Biol. 9, 190003 (2019).
Adrain, C., Zettl, M., Christova, Y., Taylor, N. & Freeman, M. Tumor necrosis factor signaling requires iRhom2 to promote trafficking and activation of TACE. Science 335, 225–228 (2012).
McIlwain, D. R. et al. iRhom2 regulation of TACE controls TNF-mediated protection against Listeria and responses to LPS. Science 335, 229–232 (2012).
Cavadas, M. et al. Phosphorylation of iRhom2 controls stimulated proteolytic shedding by the metalloprotease ADAM17/TACE. Cell Rep. 21, 745–757 (2017).
Grieve, A. G. et al. Phosphorylation of iRhom2 at the plasma membrane controls mammalian TACE-dependent inflammatory and growth factor signalling. eLife 6, e23968 (2017).
Hosur, V., Farley, M. L., Burzenski, L. M., Shultz, L. D. & Wiles, M. V. ADAM17 is essential for ectodomain shedding of the EGF-receptor ligand amphiregulin. FEBS Open Bio 8, 702–710 (2018).
Zaiss, D. M. W., Gause, W. C., Osborne, L. C. & Artis, D. Emerging functions of amphiregulin in orchestrating immunity, inflammation, and tissue repair. Immunity 42, 216–226 (2015).
Christova, Y., Adrain, C., Bambrough, P., Ibrahim, A. & Freeman, M. Mammalian iRhoms have distinct physiological functions including an essential role in TACE regulation. EMBO Rep. 14, 884–890 (2013).
Li, X. et al. iRhoms 1 and 2 are essential upstream regulators of ADAM17-dependent EGFR signaling. Proc. Natl Acad. Sci. USA 112, 6080–6085 (2015).
Su, A. I. et al. A gene atlas of the mouse and human protein-encoding transcriptomes. Proc. Natl Acad. Sci. USA 101, 6062–6067 (2004).
Issuree, P. D. et al. iRHOM2 is a critical pathogenic mediator of inflammatory arthritis. J. Clin. Invest. 123, 928–932 (2013).
Qing, X. et al. iRhom2 promotes lupus nephritis through TNF-ɑ and EGFR signaling. J. Clin. Invest. 128, 1397–1412 (2018).
Chenxu, G. et al. iRhom2 loss alleviates renal injury in long-term PM2.5-exposed mice by suppression of inflammation and oxidative stress. Redox Biol. 19, 147–157 (2018).
Kim, J. H. et al. Role of iRhom2 in intestinal ischemia-reperfusion-mediated acute lung injury. Sci. Rep. 8, 3797 (2018).
Luo, W. W. et al. iRhom2 is essential for innate immunity to DNA viruses by mediating trafficking and stability of the adaptor STING. Nat. Immunol. 17, 1057–1066 (2016).
Luo, W. W. et al. iRhom2 is essential for innate immunity to RNA virus by antagonizing ER- and mitochondria-associated degradation of VISA. PLoS Pathog. 13, e1006693 (2017).
Blaydon, D. C. et al. RHBDF2 mutations are associated with tylosis, a familial esophageal cancer syndrome. Am. J. Hum. Genet. 90, 340–346 (2012).
Saarinen, S. et al. Analysis of a Finnish family confirms RHBDF2 mutations as the underlying factor in tylosis with esophageal cancer. Fam. Cancer 11, 525–528 (2012).
Brooke, M. A. et al. iRHOM2-dependent regulation of ADAM17 in cutaneous disease and epidermal barrier function. Hum. Mol. Genet. 23, 4064–4076 (2014).
Hosur, V. et al. Rhbdf2 mutations increase its protein stability and drive EGFR hyperactivation through enhanced secretion of amphiregulin. Proc. Natl Acad. Sci. USA 111, E2200–E2209 (2014).
Hosur, V., Low, B. E., Shultz, L. D. & Wiles, M. V. Genetic deletion of amphiregulin restores the normal skin phenotype in a mouse model of the human skin disease tylosis. Biol. Open 6, 1174–1179 (2017).
Yong, P. F. et al. An update on the hyper-IgE syndromes. Arthritis Res. Ther. 14, 228 (2012).
Al-Shaikhly, T. & Ochs, H. D. Hyper IgE syndromes: clinical and molecular characteristics. Immunol. Cell Biol. 97, 368–379 (2019).
Grimbacher, B. et al. Genetic linkage of hyper-IgE syndrome to chromosome 4. Am. J. Hum. Genet. 65, 735–744 (1999).
Vassalli, P. The pathophysiology of tumor necrosis factors. Annu. Rev. Immunol. 10, 411–452 (1992).
Black, R. A. et al. A metalloproteinase disintegrin that releases tumour-necrosis factor-ɑ from cells. Nature 385, 729–733 (1997).
Moss, M. L. et al. Cloning of a disintegrin metalloproteinase that processes precursor tumour-necrosis factor-ɑ. Nature 385, 733–736 (1997).
Zunke, F. & Rose-John, S. The shedding protease ADAM17: physiology and pathophysiology. Biochim. Biophys. Acta Mol. Cell Res. 1864, 2059–2070 (2017).
Dreymueller, D. et al. Smooth muscle cells relay acute pulmonary inflammation via distinct ADAM17/ErbB axes. J. Immunol. 192, 722–731 (2014).
Aversa, G., Punnonen, J. & de Vries, J. E. The 26-kD transmembrane form of tumor necrosis factor ɑ on activated CD4+ T cell clones provides a costimulatory signal for human B cell activation. J. Exp. Med. 177, 1575–1585 (1993).
Hudson, D. M. et al. P3h3-null and Sc65-null mice phenocopy the collagen lysine under-hydroxylation and cross-linking abnormality of Ehlers–Danlos syndrome type VIA. J. Biol. Chem. 292, 3877–3887 (2017).
Tian, Y. et al. Cytokine secretion requires phosphatidylcholine synthesis. J. Cell Biol. 181, 945–957 (2008).
Sadikot, R. T., Blackwell, T. S., Christman, J. W. & Prince, A. S. Pathogen–host interactions in Pseudomonas aeruginosa pneumonia. Am. J. Respir. Crit. Care Med. 171, 1209–1223 (2005).
Berasain, C. & Avila, M. A. Amphiregulin. Semin. Cell Dev. Biol. 28, 31–41 (2014).
Liang, C. C., Park, A. Y. & Guan, J. L. In vitro scratch assay: a convenient and inexpensive method for analysis of cell migration in vitro. Nat. Protoc. 2, 329–333 (2007).
Atarashi, K. et al. Ectopic colonization of oral bacteria in the intestine drives TH1 cell induction and inflammation. Science 358, 359–365 (2017).
Schirmer, M. et al. Compositional and temporal changes in the gut microbiome of pediatric ulcerative colitis patients are linked to disease course. Cell Host Microbe 24, 600–610 e604 (2018).
Collins, J. W. et al. Citrobacter rodentium: infection, inflammation and the microbiota. Nat. Rev. Microbiol. 12, 612–623 (2014).
Lee, A., Fox, J. G., Otto, G. & Murphy, J. A small animal model of human Helicobacter pylori active chronic gastritis. Gastroenterology 99, 1315–1323 (1990).
Marchetti, M. et al. Development of a mouse model of Helicobacter pylori infection that mimics human disease. Science 267, 1655–1658 (1995).
Higgins, L. M., Frankel, G., Douce, G., Dougan, G. & MacDonald, T. T. Citrobacter rodentium infection in mice elicits a mucosal Th1 cytokine response and lesions similar to those in murine inflammatory bowel disease. Infect. Immun. 67, 3031–3039 (1999).
Poholek, C. H. et al. Noncanonical STAT3 activity sustains pathogenic Th17 proliferation and cytokine response to antigen. J. Exp. Med. 217, e20191761 (2020).
Lee, P. W. et al. IL-23R-activated STAT3/STAT4 is essential for Th1/Th17-mediated CNS autoimmunity. JCI Insight 2, e91663 (2017).
Zheng, Y. et al. TNFɑ promotes Th17 cell differentiation through IL-6 and IL-1β produced by monocytes in rheumatoid arthritis. J. Immunol. Res. 2014, 385352 (2014).
Fremond, C. et al. Membrane TNF confers protection to acute mycobacterial infection. Respir. Res. 6, 136 (2005).
Pfeffer, K. et al. Mice deficient for the 55 kd tumor necrosis factor receptor are resistant to endotoxic shock, yet succumb to L. monocytogenes infection. Cell 73, 457–467 (1993).
Rothe, J. et al. Mice lacking the tumour necrosis factor receptor 1 are resistant to TNF-mediated toxicity but highly susceptible to infection by Listeria monocytogenes. Nature 364, 798–802 (1993).
Cook, S. A. et al. HEM1 deficiency disrupts mTORC2 and F-actin control in inherited immunodysregulatory disease. Science 369, 202–207 (2020).
Crepin, V. F., Collins, J. W., Habibzay, M. & Frankel, G. Citrobacter rodentium mouse model of bacterial infection. Nat. Protoc. 11, 1851–1876 (2016).
Ozen, A. et al. Broadly effective metabolic and immune recovery with C5 inhibition in CHAPLE disease. Nat. Immunol. 22, 128–139 (2021).
Segata, N. et al. Metagenomic microbial community profiling using unique clade-specific marker genes. Nat. Methods 9, 811–814 (2012).
Knights, D. et al. Bayesian community-wide culture-independent microbial source tracking. Nat. Methods 8, 761–763 (2011).
Messier, E. M., Mason, R. J. & Kosmider, B. Efficient and rapid isolation and purification of mouse alveolar type II epithelial cells. Exp. Lung Res. 38, 363–373 (2012).
Lee, J. S. et al. Immunophenotyping of COVID-19 and influenza highlights the role of type I interferons in development of severe COVID-19. Sci. Immunol. 5, eabd1554 (2020).
Liao, Y., Wang, J., Jaehnig, E. J., Shi, Z. & Zhang, B. WebGestalt 2019: gene set analysis toolkit with revamped UIs and APIs. Nucleic Acids Res. 47, W199–W205 (2019).
Saraiva, L. R. et al. Hierarchical deconstruction of mouse olfactory sensory neurons: from whole mucosa to single-cell RNA-seq. Sci. Rep. 5, 18178 (2015).
Chen, J., Bardes, E. E., Aronow, B. J. & Jegga, A. G. ToppGene Suite for gene list enrichment analysis and candidate gene prioritization. Nucleic Acids Res. 37, W305–W311 (2009).
Acknowledgements
This work was supported by the Division of Intramural Research, National Institute of Allergy and Infectious Diseases, NIH, and the Sidra Medicine Internal Research Fund (grant nos. SDR400013, SDR200070). S.K. was supported by the Japan Research Foundation for Clinical Pharmacology. J.M.F. was supported by the Postdoctoral Research Associate Training Program of the National Institute of General Medical Sciences. A.D.W. was supported by the Emory University M.D./Ph.D. Program, the National Institutes of Health M.D./Ph.D. Partnerships Program, the National Institutes of Health Oxford-Cambridge Scholars Program and the International Biomedical Research Alliance. We thank K. Huang, S. Xirasagar, D. Hurt and other members of the Bioinformatics and Computational Biosciences Branch (BCBB), NIAID, and Y. Zhang for variant assessment and bioinformatics support. We also thank the Sidra Medicine pathology team, notably W. Mifsud, and the Sidra Medicine Genomics and Bioinformatics cores’ teams, notably S. Lorenz, L. Mathew, L. Liu, K. Wang, F. Vempalli and G. Mubarak, for technical support. We thank H. Su, J. Milner, J. Ravell and Y. Zhang for invaluable editorial and scientific feedback. We are grateful to Merck Sharp & Dohme Corp., a subsidiary of Merck & Co., Inc., Kenilworth, NJ, USA, for generous support.
Author information
Authors and Affiliations
Contributions
S.K., J.M.F., H.M.R.-M. and R.K. performed experiments and contributed to the study design, overall review and writing of the manuscript. I.V.-C., A.A.-S., Y.Y., L.Z., J.Z., A.D.W., X.J., T.K.F., A.Y.P., A.J.O., A.K.C., M.M., E.H.A.M., R.H., L.R.S., S.G., A.A.A.-S., E.F., H.H.L., A.F.F. and Y.B. performed experiments and analyzed and interpreted data. H.M., A.P.S., E.S., E.C., E.K.-A., S.B. and A.O. managed and oversaw care of the patients. A.O. and B.L. participated in the study design and coordination. M.J.L. contributed to the study design, overall review and writing of the manuscript and coordinated the overall direction of the study. All authors have read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
H.H.L. and E.F. were employees of Merck Sharp & Dohme Corp., a subsidiary of Merck & Co., Inc., Kenilworth, NJ, USA. All other authors have no competing interests.
Peer review information
Nature Immunology thanks Alain Fischer and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. L. A. Dempsey was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Disease and RHBDF expression.
(a) (left) Thoracic computed tomography (CT) scan in healthy control. (right) Photographs of the mucosa from lower G.I. endoscopy in healthy control. (b) RNA−seq data (GTEx Portal https://www.gtexportal.org) showing RHBDF2 expression across organs. (c) RHBDF1 expression in peripheral blood cell-types. Data are derived from the single cell RNA-seq dataset from the GEO database, accession number GSE149689. The clusters were projected into a 2-dimensional space using uniform manifold approximation and projection (UMAP) and identified using (left panel) canonical markers, and (right panel) RHBDF1 expression in individual cells. Color intensity indicates the expression level.
Extended Data Fig. 2 iRHOM2 deficiency impairs ADAM17-dependent TNF shedding.
(a) Quantitative RT-PCR (Q-PCR) of ADAM17 mRNA isolated from healthy control (HC) and patient (P) T cells. β-actin, GAPDH, or 18 S served as the endogenous control. Data of three independent experiments. (b) Western blot of TNF and Na + /K + ATPase Alpha 1 expression in membrane fractions of patient and healthy T cells stimulated with anti-CD3 for four hours. (c) Membrane TNF on live-gated CD14+ monocytes from HCs and P1 either untreated or stimulated with lipopolysaccharide (LPS) for 4 hours measured as in Fig. 1 (n = 4 for HC group). (d) Flow cytometry as in b, for HC cells either with or without treating with control or iRhom2 (RHBDF2)-targeting small interfering RNAs (siRNAs) as indicated. (e) Knockdown and knockout efficiency are shown by Q-PCR of RHBDF2 using RNA isolated from T cells. β−actin was the endogenous control. RQ, relative quantitation (n = 3 per group). (f) Knockout efficiency shown by western blot of ADAM17 and β-actin loading control using T cells from HC. Asterisk, pro-ADAM17; arrowhead, mature ADAM17. (g) mTNF (mean fluorescence intensity) MFI for T cells from HC and Ps stimulated with anti-CD3 antibody. Data of three independent experiments. (h) mTNF MFI for HC T cells treated with control sgRNA and iRHOM2 sgRNA and the stimulated with 10 μg/ml anti-CD3 antibody for 4 hours (n = 3 per group). (i) Q-PCR of RHBDF2 mRNA isolated from T cells treated with negative (control) or wild-type RHBDF2 coding sequence. RNA was isolated 5 days after lentiviral transduction. β−actin was the endogenous control (n = 3 per group). (j) Flow cytometry dot plots of cell side scatter (SSC) mTNF expression (percentage given in the expression gate) by live CD2+ HC T cells transduced with empty or iRHOM2 overexpressing lentiviral vector and then stimulated with anti-CD3 (10 μg/ml) for 4 hours. (k) Flow cytometric mTNF measurements from isolated cell populations from either WT or Rhbdf2−/− mice (Gating strategy was shown in Extended Data Fig. 6); Error bars represent standard error of the mean (n = 6 per group). (l) CD4 T cells (upper panel) and naïve CD8 T cells (lower panel) from HC and STAT3-deficient HIES patients stimulated with PMA (20 ng/ml) and ionomycin (750 ng/ml) or dimethyl sulfoxide (DMSO) vehicle for 4 hours and analyzed. Scatter plots were generated by gating on live CD3+CD4+, CD3+CD8+, CD4+, or CD8+ T cells (n = 2 per group). All data are mean ± s.d. and were analyzed by two-tailed, unpaired Student’s t-test (g, h, k).
Extended Data Fig. 3 Cytokine and mRNA changes associated with iRHOM2 deficiency.
(a) Heat map of multiplex measurements of 48 cytokine levels in HC plasma using the Luminex system. (b) Volcano plot of whole blood RNA-seq showing upregulated genes (red) and down regulated genes (blue) in the patient sample from iRHOM2 deficient patients compared to HC. (c) GSEA based on the patient whole blood RNA-seq data in comparison to healthy control. NES; normalized enrichment score. (d) Table of ADAM17 regulated molecules extracted from whole blood RNA-seq. (e) Volcano plot of gene expression with fold difference between log2 normalized expression shown in Fig. 5d versus −log10 adjusted P-value in whole blood. Vertical grey lines indicate fold changes, with a cut off ±2. The horizontal line represents a p-value of 1−10. (f) Similar with Fig. 5f, volcano plot of gene expression before and after 3 hours stimulation in T cells from 4 HC (left panel) and 4 patients (right panel). Red dots represent upregulated mRNAs and blue dots indicate downregulated mRNAs. (g) Hierarchical clustering of stimulated T cells RNA-seq data in the different time points using HC samples. Time is shown on the x-axis and gene expression levels are shown on the y-axis. Mean expression was used from 4 HC samples. Statistical significance was calculated by using the Wald test for hypothesis testing (b, e, f).
Extended Data Fig. 4 Loss of iRHOM2 in lung epithelial cells affects ADAM17 maturation/activity.
(a) Flow cytometry dot plots confirming the enrichment of mouse non-hematopoietic CD326+ epithelial cells isolated from total lung cells from two wild type (WT) or Rhbdf2−/− (KO) mice by magnetic separation. CD45, hematopoietic cell marker; CD326, epithelial cell marker. (b) Western blot as in Fig. 4 g, of A549 lung tumor cells treated with the indicated siRNA. (c) Q-PCR of RHBDF1, RHBDF2 and ADAM17 in A549 cells treated with the indicated siRNAs or control RNA for 48 hours. RQ, relative quantitation. Data of three independent experiments (n = 3 per group). (d) Q-PCR as in (c), in A549 cells (n = 3 per group). (e) Q-PCR as in (c), in H1299 cells (n = 3 per group). (f) Q-PCR as in (c), in PC9 cells (n = 3 per group). (g) Q-PCR of AREG, HBEGF and TGFA in A549 cells knocked out of indicated targets (n = 3 per group). (h) Wound healing assay as in Fig. 4j, in knockout of indicated targets in A549 cells (n = 3 per group). (i) Wound healing assay as in Fig. 4j, in knockout H1299 cells either with or without HB-EGF (n = 9 per group). (j) Wound healing assay as in Fig. 4i, in knockout A549 cells (n = 9 per group). (k) Wound healing assay as in Fig. 4i, in knockout PC9 cells (n = 9 per group). All data are mean ± s.d. and were analyzed by two-tailed, unpaired Student’s t-test (h-k).
Extended Data Fig. 5 Loss of iRHOM2 in colon epithelial cells is compensated for by iRHOM1 expression.
(a) Scatter plot of principal coordinates analysis axis one (PC1) and axis two (PC2) performed on North American healthy control and patients’ microbiome profiles. Individual data points are colored according to patients and HC. (b) Composition of microbes in patient stool organized by identified families of bacteria (c) Crypt length in colons of mice with and without infection by C. rodentium (n = 3 for each WT and KO groups). (d) The number of goblet cells in 500 µm2 in colons of mice (n = 3 for each WT and KO groups). (e) Q-PCR of RHBDF1 and RHBDF2 in SW620 cells treated with the indicated sgRNAs or control RNA. 18 S was the endogenous control. RQ, relative quantitation (n = 3 per group). All data are mean ± s.d. and were analyzed by two-tailed, unpaired Student’s t-test (c-e).
Extended Data Fig. 6 Gating strategy for detecting T cells, eosinophils, neutrophils, and macrophages.
Scatter plots of gating strategy are shown.
Supplementary information
Source data
Source Data Fig. 1
Unprocessed immunoblot image.
Source Data Fig. 2
Unprocessed immunoblot image.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Unprocessed immunoblot image.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Unprocessed immunoblot image.
Source Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 2
Unprocessed immunoblot image.
Source Data Extended Data Fig. 2
Statistical source data for Fig. 6g.
Source Data Extended Data Fig. 4
Unprocessed immunoblot image.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Rights and permissions
About this article
Cite this article
Kubo, S., Fritz, J.M., Raquer-McKay, H.M. et al. Congenital iRHOM2 deficiency causes ADAM17 dysfunction and environmentally directed immunodysregulatory disease. Nat Immunol 23, 75–85 (2022). https://doi.org/10.1038/s41590-021-01093-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-021-01093-y