Abstract
T cell activation, a key early event in the adaptive immune response, is subject to elaborate transcriptional control. In the present study, we examined how the activities of eight major transcription factor (TF) families are integrated to shape the epigenome of naive and activated CD4 and CD8 T cells. By leveraging extensive polymorphisms in evolutionarily divergent mice, we identified the ‘heavy lifters’ positively influencing chromatin accessibility. Members of Ets, Runx and TCF/Lef TF families occupied the vast majority of accessible chromatin regions, acting as ‘housekeepers’, ‘universal amplifiers’ and ‘placeholders’, respectively, at sites that maintained or gained accessibility upon T cell activation. In addition, a small subset of strongly induced immune response genes displayed a noncanonical TF recruitment pattern. Our study provides a key resource and foundation for the understanding of transcriptional and epigenetic regulation in T cells and offers a new perspective on the hierarchical interactions between critical TFs.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All next-generation sequencing data generated in this paper were deposited in the Gene Expression Omnibus under SuperSeries accession nos, GSE166718 and GSE184006. Processed data are included with the manuscript as supplementary tables. The B6 reference genome (GRCm38) was downloaded from the National Center for Biotechnology Information. Files CAST_EiJ.mgp.v5.snps.dbSNP142.vcf and CAST_EiJ.mgp.v5.indels.dbSNP142.normed.vcf with SNP and indel genetic variants between B6 and Cast genomes were obtained from the Mouse Genome Project53. Gene annotations were downloaded from the ENSEMBL database at ftp.ensembl.org/ensembl/pub/release-83/gtf or obtained from GENCODE v.M5 (ref. 60). Experimentally determined Mus musculus motifs identified by CHiP-seq, PBM, SELEX or HocoMoco were retrieved from the CIS-BP database57.
Code availability
The customized Shell and Python code used in the present study is publicly available at the MSKCC website: http://cbio.mskcc.org/public/Leslie/zhongy/downloads/index8.html
References
Kaech, S. M. & Cui, W. Transcriptional control of effector and memory CD8+ T cell differentiation. Nat. Rev. Immunol. 12, 749–761 (2012).
Zhu, J., Yamane, H. & Paul, W. E. Differentiation of effector CD4 T cell populations (*). Annu. Rev. Immunol. 28, 445–489 (2010).
McLane, L. M., Abdel-Hakeem, M. S. & Wherry, E. J. CD8 T cell exhaustion during chronic viral infection and cancer. Annu. Rev. Immunol. 37, 457–495 (2019).
Mackay, L. K. & Kallies, A. Transcriptional regulation of tissue-resident lymphocytes. Trends Immunol. 38, 94–103 (2017).
Milner, J. J. & Goldrath, A. W. Transcriptional programming of tissue-resident memory CD8. Curr. Opin. Immunol. 51, 162–CD169 (2018).
Vaquerizas, J. M., Kummerfeld, S. K., Teichmann, S. A. & Luscombe, N. M. A census of human transcription factors: function, expression and evolution. Nat. Rev. Genet. 10, 252–263 (2009).
Gray, P. A. et al. Mouse brain organization revealed through direct genome-scale TF expression analysis. Science 306, 2255–2257 (2004).
Badis, G. et al. Diversity and complexity in DNA recognition by transcription factors. Science 324, 1720–1723 (2009).
Jolma, A. et al. DNA-binding specificities of human transcription factors. Cell 152, 327–339 (2013).
Gerstein, M. B. et al. Architecture of the human regulatory network derived from ENCODE data. Nature 489, 91–100 (2012).
Lee, T. I. & Young, R. A. Transcriptional regulation and its misregulation in disease. Cell 152, 1237–1251 (2013).
Spitz, F. & Furlong, E. E. Transcription factors: from enhancer binding to developmental control. Nat. Rev. Genet. 13, 613–626 (2012).
Chang, J. T., Wherry, E. J. & Goldrath, A. W. Molecular regulation of effector and memory T cell differentiation. Nat. Immunol. 15, 1104–1115 (2014).
Hosokawa, H. & Rothenberg, E. V. How transcription factors drive choice of the T cell fate. Nat. Rev. Immunol. https://doi.org/10.1038/s41577-020-00426-6 (2020).
van der Veeken, J. et al. The transcription factor Foxp3 shapes regulatory T cell identity by tuning the activity of trans-acting intermediaries. Immunity 53, 971–984.e975 (2020).
van der Veeken, J. et al. Natural genetic variation reveals key features of epigenetic and transcriptional memory in virus-specific CD8 T cells. Immunity 50, 1202–1217.e1207 (2019).
Hota, S. K. & Bruneau, B. G. ATP-dependent chromatin remodeling during mammalian development. Development 143, 2882–2897 (2016).
Polansky, J. K. et al. Methylation matters: binding of Ets-1 to the demethylated Foxp3 gene contributes to the stabilization of Foxp3 expression in regulatory T cells. J. Mol. Med. 88, 1029–1040 (2010).
Wotton, D., Ghysdael, J., Wang, S., Speck, N. A. & Owen, M. J. Cooperative binding of Ets-1 and core binding factor to DNA. Mol. Cell Biol. 14, 840–850 (1994).
Kim, W. Y. et al. Mutual activation of Ets-1 and AML1 DNA binding by direct interaction of their autoinhibitory domains. EMBO J. 18, 1609–1620 (1999).
Wang, C. Y. et al. Activation of the granulocyte–macrophage colony-stimulating factor promoter in T cells requires cooperative binding of Elf-1 and AP-1 transcription factors. Mol. Cell Biol. 14, 1153–1159 (1994).
Bassuk, A. G. & Leiden, J. M. A direct physical association between ETS and AP-1 transcription factors in normal human T cells. Immunity 3, 223–237 (1995).
Pognonec, P., Boulukos, K. E., Gesquière, J. C., Stéhelin, D. & Ghysdael, J. Mitogenic stimulation of thymocytes results in the calcium-dependent phosphorylation of c-ets-1 proteins. EMBO J. 7, 977–983 (1988).
Rabault, B. & Ghysdael, J. Calcium-induced phosphorylation of ETS1 inhibits its specific DNA binding activity. J. Biol. Chem. 269, 28143–28151 (1994).
Cowley, D. O. & Graves, B. J. Phosphorylation represses Ets-1 DNA binding by reinforcing autoinhibition. Genes Dev. 14, 366–376 (2000).
Foulds, C. E., Nelson, M. L., Blaszczak, A. G. & Graves, B. J. Ras/mitogen-activated protein kinase signaling activates Ets-1 and Ets-2 by CBP/p300 recruitment. Mol. Cell Biol. 24, 10954–10964 (2004).
Bories, J. C. et al. Increased T-cell apoptosis and terminal B-cell differentiation induced by inactivation of the Ets-1 proto-oncogene. Nature 377, 635–638 (1995).
Muthusamy, N., Barton, K. & Leiden, J. M. Defective activation and survival of T cells lacking the Ets-1 transcription factor. Nature 377, 639–642 (1995).
Mélet, F., Motro, B., Rossi, D. J., Zhang, L. & Bernstein, A. Generation of a novel Fli-1 protein by gene targeting leads to a defect in thymus development and a delay in Friend virus-induced erythroleukemia. Mol. Cell Biol. 16, 2708–2718 (1996).
Kim, C. J. et al. The transcription factor Ets1 suppresses T follicular helper type 2 cell differentiation to halt the onset of systemic lupus erythematosus. Immunity 49, 1034–1048.e1038 (2018).
Anderson, M. K., Hernandez-Hoyos, G., Diamond, R. A. & Rothenberg, E. V. Precise developmental regulation of Ets family transcription factors during specification and commitment to the T cell lineage. Development 126, 3131–3148 (1999).
Yamada, T., Park, C. S., Mamonkin, M. & Lacorazza, H. D. Transcription factor ELF4 controls the proliferation and homing of CD8+ T cells via the Krüppel-like factors KLF4 and KLF2. Nat. Immunol. 10, 618–626 (2009).
Luo, C. T. et al. Ets transcription factor GABP controls T cell homeostasis and immunity. Nat. Commun. 8, 1062 (2017).
Hollenhorst, P. C., McIntosh, L. P. & Graves, B. J. Genomic and biochemical insights into the specificity of ETS transcription factors. Annu. Rev. Biochem. 80, 437–471 (2011).
Thompson, C. B. et al. cis-acting sequences required for inducible interleukin-2 enhancer function bind a novel Ets-related protein, Elf-1. Mol. Cell Biol. 12, 1043–1053 (1992).
Wang, C. Y., Petryniak, B., Ho, I. C., Thompson, C. B. & Leiden, J. M. Evolutionarily conserved Ets family members display distinct DNA binding specificities. J. Exp. Med. 175, 1391–1399 (1992).
Hollenhorst, P. C., Shah, A. A., Hopkins, C. & Graves, B. J. Genome-wide analyses reveal properties of redundant and specific promoter occupancy within the ETS gene family. Genes Dev. 21, 1882–1894 (2007).
Hollenhorst, P. C. et al. DNA specificity determinants associate with distinct transcription factor functions. PLoS Genet. 5, e1000778 (2009).
Giese, K., Kingsley, C., Kirshner, J. R. & Grosschedl, R. Assembly and function of a TCR alpha enhancer complex is dependent on LEF-1-induced DNA bending and multiple protein–protein interactions. Genes Dev. 9, 995–1008 (1995).
Verbaro, D. J., Sakurai, N., Kim, B., Shinkai, Y. & Egawa, T. Cutting edge: the histone methyltransferase G9a is required for silencing of helper T lineage-associated genes in proliferating CD8 T cells. J. Immunol. 200, 3891–3896 (2018).
Lazarevic, V. et al. T-bet represses TH17 differentiation by preventing Runx1-mediated activation of the gene encoding RORγt. Nat. Immunol. 12, 96–104 (2011).
Zhang, F., Meng, G. & Strober, W. Interactions among the transcription factors Runx1, RORgammat and Foxp3 regulate the differentiation of interleukin 17-producing T cells. Nat. Immunol. 9, 1297–1306 (2008).
Zheng, Y. et al. Role of conserved non-coding DNA elements in the Foxp3 gene in regulatory T-cell fate. Nature 463, 808–812 (2010).
Shin, B. et al. Runx1 and Runx3 drive progenitor to T-lineage transcriptome conversion in mouse T cell commitment via dynamic genomic site switching. Proc. Natl Acad. Sci. USA https://doi.org/10.1073/pnas.2019655118 (2021)
Taniuchi, I. et al. Differential requirements for Runx proteins in CD4 repression and epigenetic silencing during T lymphocyte development. Cell 111, 621–633 (2002).
Egawa, T., Tillman, R. E., Naoe, Y., Taniuchi, I. & Littman, D. R. The role of the Runx transcription factors in thymocyte differentiation and in homeostasis of naive T cells. J. Exp. Med. 204, 1945–1957 (2007).
Levanon, D. et al. Transcriptional repression by AML1 and LEF-1 is mediated by the TLE/Groucho corepressors. Proc. Natl Acad. Sci. USA 95, 11590–11595 (1998).
Setoguchi, R. et al. Repression of the transcription factor Th-POK by Runx complexes in cytotoxic T cell development. Science 319, 822–825 (2008).
Johnson, J. L. et al. Lineage-determining transcription factor TCF-1 initiates the epigenetic identity of T cells. Immunity 48, 243–257.e210 (2018).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Skene, P. J. & Henikoff, S. An efficient targeted nuclease strategy for high-resolution mapping of DNA binding sites. eLife https://doi.org/10.7554/eLife.21856 (2017).
Liu, N. et al. Direct promoter repression by BCL11A controls the fetal to adult hemoglobin switch. Cell 173, 430–442.e417 (2018).
Keane, T. M. et al. Mouse genomic variation and its effect on phenotypes and gene regulation. Nature 477, 289–294 (2011).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Li, Q., Brown, J. B., Huang, H. & Bickel, P. J. Measuring reproducibility of high-throughput experiments. Ann. Appl. Stat. 5, 1752–1779 (2011).
Weirauch, M. T. et al. Determination and inference of eukaryotic transcription factor sequence specificity. Cell 158, 1431–1443 (2014).
Grant, C. E., Bailey, T. L. & Noble, W. S. FIMO: scanning for occurrences of a given motif. Bioinformatics 27, 1017–1018 (2011).
Link, V. M., Romanoski, C. E., Metzler, D. & Glass, C. K. MMARGE: motif mutation analysis for regulatory genomic elements. Nucleic Acids Res. 46, 7006–7021 (2018).
Frankish, A. et al. GENCODE reference annotation for the human and mouse genomes. Nucleic Acids Res. 47, D766–D773 (2019).
Granja, J. M. et al. ArchR is a scalable software package for integrative single-cell chromatin accessibility analysis. Nat. Genet. 53, 403–411 (2021).
Acknowledgements
We thank all members of the Rudensky lab for discussions and technical assistance. We thank the NIH tetramer core for providing critical reagents. The present study was supported by NIH grants (nos. R01AI034206, U54 CA209975) and NIH/National Cancer Institute (NCI, grant no. P30 CA008748), AACR-Bristol-Myers Squibb Immuno-oncology Research Fellowship (no. 19-40-15-PRIT), National Natural Science Foundation of China (grant no. 32170883 to Y.Z.), the Ludwig Center at MSKCC and the Parker Institute for Cancer Immunotherapy. A.Y.R. is an investigator with the Howard Hughes Medical Institute. We acknowledge the use of the MSKCC Single Cell Research Initiative and the Integrated Genomics Operation Core, funded by the NIH/NCI Cancer Center Support (grant no. P30 CA008748), Cycle for Survival, and the Marie-Josée and Henry R. Kravis Center for Molecular Oncology.
Author information
Authors and Affiliations
Contributions
J.v.d.V. and A.Y.R. designed the study. J.v.d.V. performed the experiments and analyzed the data. Y.Z. and C.S.L. designed the allele-specific analysis pipeline. Y.Z. performed the computational analysis of sequencing data. S.K.W. and Y.P. analyzed the scATAC-seq data. J.v.d.V. and A.Y.R. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
A.Y.R. is an SAB member, has equity in Sonoma Biotherapeutics and Vedanta Biosciences, and is a co-inventor or has IP licensed to Takeda that is unrelated to the content of the present study. The remaining authors declare no competing interests.
Additional information
Peer review information.Peer review reports are available. L. A. Dempsey was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 RNA-seq and ATAC-seq of naive and activated CD4 and CD8 T cells.
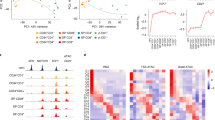
a: Example of gating strategy for cell isolation from the spleen. Live lymphocytes were identified based on FSC-A/SSC-A. Doublets were gated out and CD4 and CD8 T cells were identified as TCRβ+CD4+ and TCRβ+CD8+, respectively. Naive CD4 and CD8 T cells used as input for RNA-seq, ATAC-seq, and CUT&RUN experiments were isolated from uninfected mice as CD44+CD62L− cells. Bulk activated CD4 and CD8 T cells used as input for CUT&RUN experiments were isolated as CD44+ cells from LCMV Armstrong infected mice on day 7-8 post-infection (p.i.). LCMV-specific activated and memory CD4 and CD8 T cells used as input for ATAC-seq and RNA-seq experiments were isolated based on tetramer staining on day 7 or day 60 p.i. b: RNA-seq and ATAC-seq reads at the Cd4, Cd8a, Sell and Cd44 loci in naive and activated CD4 and CD8 T cells. c-e: Effect of TF binding motif variants on chromatin accessibility measured as ΔMean in naive and activated CD4 T cells (A), naive CD4 and CD8 T cells (B), activated CD4 and CD8 T cells (C). Data points are colored by TF family and scaled according to the –log10 p-value of a two-sided t-test comparing the mean allelic ratios between peaks with stronger matches on the B6 vs Cast allele. The most significant p-value obtained across the two cell types (naive and activated CD8) was used for scaling. Motifs with <50 variant-containing motif occurrences were excluded from the plot. f: Mean RNA-seq counts for Tbx21, Eomes, Tcf7 and Lef1 in naive and activated CD4 and CD8 T cells.
Extended Data Fig. 2 Association between gene expression and chromatin accessibility changes.
a: Correlation between changes in chromatin accessibility and gene expression in activated vs naive CD8 T cells. Showing individual ATAC-seq peaks linked to the nearest gene (left) or the sum of counts across all ATAC-seq peaks linked to the same gene (right). b: Fold change in gene expression in activated vs naive CD8 T cells (x-axis) plotted against the fold change in gene expression in memory vs naive CD8 T cells (y-axis). A subset of genes whose expression is transiently induced upon T cell activation is highlighted in red (left). Aggregated chromatin accessibility and gene expression changes for this gene set (right). c: Gene ontology (GO) term enrichment analysis on genes highlighted in panel B.
Extended Data Fig. 3 RNA-seq and ATAC-seq changes at Runx1, TCF1, and Ets1-bound genes.
a-b: Gene tracks of RNA-seq, ATAC-seq and CUT&RUN coverage at the Bcl2 and Arhgap30 loci in naive and activated CD8 T cells. c: Track examples showing the effect of selected Ets1 motif variants on allelic bias in chromatin accessibility and TF binding in activated CD8 T cells from (B6/Cast) F1 mice. Vertical bars mark the position of genetic variants, with colors indicating the fraction of variant-containing reads coming from either the B6 (black) or Cast (red) allele. A genetic variant in the Camk2b locus results in a stronger Ets1 motif match on the B6 allele and a weaker match on the Cast allele and is associated with B6-specific chromatin accessibility and TF binding. The opposite pattern is observed at the Dnah8 locus. d: Gene expression changes at genes (>100 reads in at least one cell type) nearest to Ets1-bound and -unbound ATAC-seq peaks. e: Mean expression of genes nearest to Ets1-bound and -unbound ATAC-seq peaks.
Extended Data Fig. 4 TF and cofactor binding at Ets1-bound and –unbound chromatin regions.
a: Gene tracks of RNA-seq, ATAC-seq and CUT&RUN coverage at the Gzmb and Tbx21 loci in naive and activated CD8 T cells. b: Fraction of peaks bound by Brg1 at Ets1-unbound ATAC-seq peaks occupied by distinct combinations of activation-induced TFs. Data points are scaled according to the number of peaks in each set. c: Effect of TF binding motif variants on Brg1 occupancy at Ets1-bound (Ets1+) and Ets1-unbound (Ets1−) sites measured as ΔMean in activated CD8 T cells. Data points are colored by TF family and scaled according to the –log10 p-value of a t-test comparing the allelic ratios between peaks with stronger motif matches on the B6 vs Cast allele. The most significant p-value obtained across the two cell types (naive and activated CD8) was used for scaling.
Extended Data Fig. 5 TF binding at TCF1-bound peak sets in naive and activated CD4 and CD8 T cells.
a-c: Gene tracks of RNA-seq, ATAC-seq and CUT&RUN read coverage at the Il7r, Pde3b and Klrg1 loci in naive and activated CD4 and CD8 T cells. d: Fraction of TCF1-bound sites and TCF1up sites containing specific TF-binding motifs. e: Median motif match p-value at sites bound at TCF1-bound sites and TCF1up sites. f: Percentage of TCF1+ Ets1− peaks and TCF1up peaks bound by each activation-induced TF.
Extended Data Fig. 6 Single cell ATAC-seq analysis of T cell subsets during acute LCMV infection.
a: Pseudo-bulk scATAC-seq tracks showing chromatin accessibility at CD4 and CD8 loci across splenic cell populations in LCMV infected (B6/Cast) F1 mice. b: Effect of genetic variation in TF binding motifs on allele-specific ATAC-seq counts from bulk cell or single cell experiments. Data points are colored by TF family and scaled according to the -log10 p-value of a two-sided t-test comparing the allelic ratios between peaks with stronger matches on B6 vs Cast Allele. c: Violin plots and box plots showing accessibility of indicated peak groups across T cell subsets. The center line in the box plots represents the median, the box limits represent the 25th and 75th percentiles, and the whiskers are the minimum/maximum values within 1.5 times the interquartile range. n = the number of cells in each cell annotation (420 Naive CD4, 344 Naive CD8, 234 Mem CD4, 231 Mem CD8, 198 Treg, 1321 Tfh, 652 Th1, 2583 Effector CD8).
Supplementary information
Rights and permissions
About this article
Cite this article
Zhong, Y., Walker, S.K., Pritykin, Y. et al. Hierarchical regulation of the resting and activated T cell epigenome by major transcription factor families. Nat Immunol 23, 122–134 (2022). https://doi.org/10.1038/s41590-021-01086-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-021-01086-x
This article is cited by
-
Runx factors launch T cell and innate lymphoid programs via direct and gene network-based mechanisms
Nature Immunology (2023)
-
Chronic lymphocytic leukemia presence impairs antigen-specific CD8+ T-cell responses through epigenetic reprogramming towards short-lived effectors
Leukemia (2023)
-
Parsing the transcription factors governing T cell immunity
Nature Immunology (2022)
-
The m1A modification of tRNAs: a translational accelerator of T-cell activation
Cellular & Molecular Immunology (2022)
-
TCF-1: a maverick in T cell development and function
Nature Immunology (2022)