Abstract
Reduced infiltration of anti-tumor lymphocytes remains a major cause of tumor immune evasion and is correlated with poor cancer survival. Here, we found that upregulation of regulator of G protein signaling (RGS)1 in helper TH1 cells and cytotoxic T lymphocytes (CTLs) reduced their trafficking to and survival in tumors and was associated with shorter survival of patients with breast and lung cancer. RGS1 was upregulated by type II interferon (IFN)–signal transducer and activator of transcription (STAT)1 signaling and impaired trafficking of circulating T cells to tumors by inhibiting calcium influx and suppressing activation of the kinases ERK and AKT. RGS1 knockdown in adoptively transferred tumor-specific CTLs significantly increased their infiltration and survival in breast and lung tumor grafts and effectively inhibited tumor growth in vivo, which was further improved when combined with programmed death ligand (PD-L)1 checkpoint inhibition. Our findings reveal RGS1 is important for tumor immune evasion and suggest that targeting RGS1 may provide a new strategy for tumor immunotherapy.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
Curtis, Kao and the Cancer Genome Atlas datasets were obtained from the Oncomine Platform (https://www.oncomine.org). Data supporting the findings of this study are available from the corresponding authors upon reasonable request. Source data are provided with this paper.
References
Gu-Trantien, C. et al. CD4+ follicular helper T cell infiltration predicts breast cancer survival. J. Clin. Invest. 123, 2873–2892 (2013).
Ruffell, B., DeNardo, D. G., Affara, N. I. & Coussens, L. M. Lymphocytes in cancer development: polarization towards pro-tumor immunity. Cytokine Growth Factor Rev. 21, 3–10 (2010).
Su, S. et al. Blocking the recruitment of naive CD4+ T cells reverses immunosuppression in breast cancer. Cell Res. 27, 461–482 (2017).
Coussens, L. M., Zitvogel, L. & Palucka, A. K. Neutralizing tumor-promoting chronic inflammation: a magic bullet? Science 339, 286–291 (2013).
Palucka, A. K. & Coussens, L. M. The basis of oncoimmunology. Cell 164, 1233–1247 (2016).
Nagarsheth, N., Wicha, M. S. & Zou, W. Chemokines in the cancer microenvironment and their relevance in cancer immunotherapy. Nat. Rev. Immunol. 17, 559–572 (2017).
Curiel, T. J. et al. Specific recruitment of regulatory T cells in ovarian carcinoma fosters immune privilege and predicts reduced survival. Nat. Med. 10, 942–949 (2004).
Huang, D. et al. NKILA lncRNA promotes tumor immune evasion by sensitizing T cells to activation-induced cell death. Nat. Immunol. 19, 1112–1125 (2018).
Franciszkiewicz, K., Boissonnas, A., Boutet, M., Combadiere, C. & Mami-Chouaib, F. Role of chemokines and chemokine receptors in shaping the effector phase of the antitumor immune response. Cancer Res. 72, 6325–6332 (2012).
De Vries, L., Zheng, B., Fischer, T., Elenko, E. & Farquhar, M. G. The regulator of G protein signaling family. Annu. Rev. Pharmacol. Toxicol. 40, 235–271 (2000).
Ross, E. M. & Wilkie, T. M. GTPase-activating proteins for heterotrimeric G proteins: regulators of G protein signaling (RGS) and RGS-like proteins. Annu. Rev. Biochem. 69, 795–827 (2000).
Xie, Z., Chan, E. C. & Druey, K. M. R4 regulator of G protein signaling (RGS) proteins in inflammation and immunity. AAPS J. 18, 294–304 (2016).
Agenes, F., Bosco, N., Mascarell, L., Fritah, S. & Ceredig, R. Differential expression of regulator of G-protein signalling transcripts and in vivo migration of CD4+ naive and regulatory T cells. Immunology 115, 179–188 (2005).
Dangaj, D. et al. Cooperation between constitutive and inducible chemokines enables T cell engraftment and immune attack in solid tumors. Cancer Cell 35, 885–900 (2019).
Harlin, H. et al. Chemokine expression in melanoma metastases associated with CD8+ T-cell recruitment. Cancer Res. 69, 3077–3085 (2009).
Chatterjee, T. K. & Fisher, R. A. Cytoplasmic, nuclear, and Golgi localization of RGS proteins. Evidence for N-terminal and RGS domain sequences as intracellular targeting motifs. J. Biol. Chem. 275, 24013–24021 (2000).
Denecke, B., Meyerdierks, A. & Bottger, E. C. RGS1 is expressed in monocytes and acts as a GTPase-activating protein for G-protein-coupled chemoattractant receptors. J. Biol. Chem. 274, 26860–26868 (1999).
Chen, D. S. & Mellman, I. Oncology meets immunology: the cancer–immunity cycle. Immunity 39, 1–10 (2013).
Hu, J. et al. T-cell homing therapy for reducing regulatory T cells and preserving effector T-cell function in large solid tumors. Clin. Cancer Res. 24, 2920–2934 (2018).
Rizvi, N. A. et al. Activity and safety of nivolumab, an anti-PD-1 immune checkpoint inhibitor, for patients with advanced, refractory squamous non-small-cell lung cancer (CheckMate 063): a phase 2, single-arm trial. Lancet Oncol. 16, 257–265 (2015).
Larkin, J. et al. Combined nivolumab and ipilimumab or monotherapy in untreated melanoma. N. Engl. J. Med. 373, 23–34 (2015).
Emens, L. A. et al. Long-term clinical outcomes and biomarker analyses of atezolizumab therapy for patients with metastatic triple-negative breast cancer: a phase 1 study. JAMA Oncol. 5, 74–82 (2019).
Schmid, P. et al. Atezolizumab and nab-paclitaxel in advanced triple-negative breast cancer. N. Engl. J. Med. 379, 2108–2121 (2018).
Youn, H. D. & Liu, J. O. Cabin1 represses MEF2-dependent Nur77 expression and T cell apoptosis by controlling association of histone deacetylases and acetylases with MEF2. Immunity 13, 85–94 (2000).
Sistigu, A. et al. Cancer cell–autonomous contribution of type I interferon signaling to the efficacy of chemotherapy. Nat. Med. 20, 1301–1309 (2014).
Nolz, J. C., Starbeck-Miller, G. R. & Harty, J. T. Naive, effector and memory CD8 T-cell trafficking: parallels and distinctions. Immunotherapy 3, 1223–1233 (2011).
Hong, J. X., Wilson, G. L., Fox, C. H. & Kehrl, J. H. Isolation and characterization of a novel B cell activation gene. J. Immunol. 150, 3895–3904 (1993).
Moratz, C., Hayman, J. R., Gu, H. & Kehrl, J. H. Abnormal B-cell responses to chemokines, disturbed plasma cell localization, and distorted immune tissue architecture in Rgs1−/− mice. Mol. Cell. Biol. 24, 5767–5775 (2004).
Gibbons, D. L. et al. Cutting edge: regulator of G protein signaling-1 selectively regulates gut T cell trafficking and colitic potential. J. Immunol. 187, 2067–2071 (2011).
Patel, J. et al. RGS1 regulates myeloid cell accumulation in atherosclerosis and aortic aneurysm rupture through altered chemokine signalling. Nat. Commun. 6, 6614 (2015).
Liu, J. et al. The IRENA lncRNA converts chemotherapy-polarized tumor-suppressing macrophages to tumor-promoting phenotypes in breast cancer. Nat. Cancer 2, 457–473 (2021).
Ma, Y. et al. Anticancer chemotherapy-induced intratumoral recruitment and differentiation of antigen-presenting cells. Immunity 38, 729–741 (2013).
Restifo, N. P., Dudley, M. E. & Rosenberg, S. A. Adoptive immunotherapy for cancer: harnessing the T cell response. Nat. Rev. Immunol. 12, 269–281 (2012).
Pockaj, B. A. et al. Localization of 111indium-labeled tumor infiltrating lymphocytes to tumor in patients receiving adoptive immunotherapy. Augmentation with cyclophosphamide and correlation with response. Cancer 73, 1731–1737 (1994).
Bernhard, H. et al. Adoptive transfer of autologous, HER2-specific, cytotoxic T lymphocytes for the treatment of HER2-overexpressing breast cancer. Cancer Immunol. Immunother. 57, 271–280 (2008).
Hong, M. et al. Chemotherapy induces intratumoral expression of chemokines in cutaneous melanoma, favoring T-cell infiltration and tumor control. Cancer Res. 71, 6997–7009 (2011).
Kershaw, M. H. et al. Redirecting migration of T cells to chemokine secreted from tumors by genetic modification with CXCR2. Hum. Gene Ther. 13, 1971–1980 (2002).
John, L. B. et al. Anti-PD-1 antibody therapy potently enhances the eradication of established tumors by gene-modified T cells. Clin. Cancer Res. 19, 5636–5646 (2013).
Su, S. et al. CD10+GPR77+ cancer-associated fibroblasts promote cancer formation and chemoresistance by sustaining cancer stemness. Cell 172, 841–856 (2018).
Chen, J. et al. CCL18 from tumor-associated macrophages promotes breast cancer metastasis via PITPNM3. Cancer Cell 19, 541–555 (2011).
Iellem, A. et al. Unique chemotactic response profile and specific expression of chemokine receptors CCR4 and CCR8 by CD4+CD25+ regulatory T cells. J. Exp. Med. 194, 847–853 (2001).
Poznansky, M. C. et al. Active movement of T cells away from a chemokine. Nat. Med. 6, 543–548 (2000).
He, S. et al. MIP-3α and MIP-1α rapidly mobilize dendritic cell precursors into the peripheral blood. J. Leukoc. Biol. 84, 1549–1556 (2008).
Bystry, R. S., Aluvihare, V., Welch, K. A., Kallikourdis, M. & Betz, A. G. B cells and professional APCs recruit regulatory T cells via CCL4. Nat. Immunol. 2, 1126–1132 (2001).
Bottcher, J. P. et al. NK cells stimulate recruitment of cDC1 into the tumor microenvironment promoting cancer immune control. Cell 172, 1022–1037 (2018).
Stanford, M. M. & Issekutz, T. B. The relative activity of CXCR3 and CCR5 ligands in T lymphocyte migration: concordant and disparate activities in vitro and in vivo. J. Leukoc. Biol. 74, 791–799 (2003).
Islam, S. A. et al. Mouse CCL8, a CCR8 agonist, promotes atopic dermatitis by recruiting IL-5+ TH2 cells. Nat. Immunol. 12, 167–177 (2011).
Lin, Y. P., Cheng, Y. J., Huang, J. Y., Lin, H. C. & Yang, B. C. Zap70 controls the interaction of talin with integrin to regulate the chemotactic directionality of T-cell migration. Mol. Immunol. 47, 2022–2029 (2010).
Su, S. et al. Immune checkpoint inhibition overcomes ADCP-induced immunosuppression by macrophages. Cell 175, 442–457 (2018).
Acknowledgements
This work was supported by grants from the National Key Research and Development Program of China (2017YFA0106300 (S.S.), 2016YFC1302300 (E.S.)), the Natural Science Foundation of China (81621004 (E.S.), 81720108029 (E.S.), 81930081 (E.S.), 91940305 (E.S.), 81802643 (D.H.), 82002786 (L.Y.), 91942309 (S.S.) and 92057210 (S.S.)), the Guangdong Basic and Applied Basic Research Foundation (2018A030310085 (D.H.), 2020A1515011458 (D.H.) and 2021A1515010238 (X.C.)), the Clinical Innovation Research Program of Bioland Laboratory (2018GZR0201004 (E.S.)), the Guangdong Science and Technology Department (2020B1212060018 (E.S.) and 2020B1212030004 (E.S.)), the Guangzhou Science Technology and Innovation Commission (201803040015 (E.S.)), the Program for Guangdong Introducing Innovative and Entrepreneurial Teams (2019BT02Y198 (E.S.)), the Science and Technology Program of Guangzhou (202103000070 (S.S.)) and the China Postdoctoral Science Foundation (2020M683106 (X.C.)). The research was partly supported by the Fountain-Valley Life Sciences Fund of the University of Chinese Academy of Sciences Education Foundation.
Author information
Authors and Affiliations
Contributions
D.H., S.S. and E.S. conceived ideas and designed experiments. S.S. and E.S. conducted experiments. D.H., X.C., X.Z., L.L., J.L., Y.X., Y.L., Q.O., J.C. and L.Y. performed experiments and acquired data. D.H. and X.Z. analyzed data. H.Y., Q.L., F.S. and E.S. provided patient samples for clinical data analysis and the PDX model. D.H., S.S. and E.S. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Immunology thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available. Zoltan Fehervari was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Low infiltration of CTLs and TH1 was associated with impaired recruitment.
a, Kaplan-Meier survival curve of breast cancer patients with CD4highCXCR3high, CD4highTBX21high, CD4highGPR44(PTGDR2)high, CD4highGATA3high or CD8highIFNGhigh signature in Curtis, TCGA or Kao breast cancer online database. Two-tailed log rank P values are shown. b, Proportion of TH1 (CD3+ CD4+ CXCR3+), TH2 (CD3+ CD4+ CRTH2+) cells and CTLs (CD3+ CD8+ CD45RO+) in peripheral blood and tumor tissues of breast cancer patients was determined by flow cytometry. Numbers indicate the percentages of TH1, TH2 cells or CTLs (mean ± s.e.m., n = 3). TH1: P = 0.0018, TH2: P = 0.0117, CTL: P = 0.0004 compared with the proportion of T cell subsets in peripheral blood by two-tailed Students’ t-test). c, TH1 cells were isolated from the peripheral blood (PB) of breast cancer patients with high TI-TH1 (P3), low TI-TH1 (P4) or healthy donors, labeled with CFSE (green), and overlaid on breast tumor slices. Then the tumor slices were fixed and stained for CD4 (white) and T-bet (red). Representative images of migrated T cells are shown (n = 5). Yellow arrows denote the migrated TH1 cells, while white arrows denote tumor-infiltrating TH1 cells. Scale bar, 50 µm. d, TH2 cells were isolated from the peripheral blood of breast cancer patients with high TI-TH2 cells (P5), low TI-TH2 cells (P6) or healthy donors, labeled with CFSE, and overlaid on breast tumor slices. Quantification of number of adherent CFSE+ T cells are shown (mean ± s.e.m). Each data point is the mean of triplicate from each human individual and data are representative of breast cancer patients with high (n = 3) or low (n = 3) TI-TH2 cells, or healthy donors (n = 3) with similar results. The exact P values are listed in the graph. P values were obtained by two-tailed one-way ANOVA with Tukey’s multiple comparisons test.
Extended Data Fig. 2 The levels of chemokines and their corresponding receptors are equivalent in the tumor with high and low TI-TH1/ CTL infiltration.
a, Relative quantification of CCL3, CCL4, CCL5, CXCL9, CXCL10, CXCL11, CCL22 and CXCL12 mRNA in breast cancer tissues with low and high TI-TH1/ CTLs were examined by RT-qPCR (mean±s.e.m., n = 24; Breast cancer tissues with low TI-TH1 cells: n = 15; Breast cancer tissues with high TI-TH1 cells: n = 9; Breast cancer tissues with low TI- CTLs: n = 15; Breast cancer tissues with high TI-CTLs: n = 9). b, Representative plots of CXCR3, CCR4, CCR5 and CXCR4 expression in TH1 cells and CTLs isolated from breast cancer with low and high TI-TH1/CTLs, were determined by flow cytometry. The lower bar graph corresponds to the mean fluorescence intensity (MFI) (mean±s.e.m., n = 5).
Extended Data Fig. 3 RGS1 upregulation in CTLs and TH1 cells correlates with the weak recruitment ability.
a, Representative images and the quantification of RGS1, RGS2 and RGS16 in naïve and effector T cells (mean±s.e.m., n = 6). Scale bar, 5 μm. b, Expression of RGS1, RGS2 and RGS16 was performed by RT-qPCR and normalized to GAPDH in TH1, TH2 and effector CD8+ T cells (CTLs) isolated from peripheral blood of healthy donors (mean±s.e.m., n = 5). c, Representative histograms of RGS1 expression in TH1, TH2 and effector CD8+ T cells isolated from peripheral blood of healthy donors, determined by flow cytometry. The bars correspond to the mean fluorescence intensity (MFI) of RGS1 (mean±s.e.m. × 104, n = 5). d, Expression of RGS1, RGS2 and RGS16 was performed by RT-qPCR and normalized to GAPDH in TH1 and CTLs isolated from breast cancer with low or high TI-TH1/CTLs (mean±s.e.m., n = 4). e, First row: automated identification of the cells co-stained with CD8 and CD45RO by Imaris software to built CD8+CD45RO+ cells (yellow). Second row: automated identification of the cells co-stained with CD4 and T-bet by Imaris software to built CD4+T-bet+ cells (yellow). The mean intensity of RGS1 in these cells were analyzed by ImageJ. Scale bar, 50 µm. f, Representative histograms of RGS1 expression in TH1, TH2 and CTLs isolated from peripheral blood of breast cancer patients, determined by flow cytometry. The bars correspond to MFI of RGS1 (mean±s.e.m. × 104, n = 5). g, CTL/TH1 infiltration is positively associated with the apoptosis of breast cancer cells (TUNEL+ %) (n = 46, Pearson’s correlation coefficient R and two-tailed P value). h, CTL infiltration, but not TH1, is negatively correlated to the proliferation of tumor cells (Ki-67+ %) (n = 46, Pearson’s correlation coefficient R and two-tailed P value). The exact P values are listed in the graph (a, c, d, f, g, h). ****P < 0.0001 (b, f). P values were determined by two-tailed Student’s t-test (a, d) and two-tailed one-way ANOVA with Tukey’s multiple comparison test (b, c, f).
Extended Data Fig. 4 RGS1 impairs CTL and TH1 recruitment in NSCLC patients.
a,b, Representative immunofluorescent staining for RGS1 expression of CTLs (a) and TH1 cells (b) infiltrated in tumors and lymph nodes (LN) indicated by arrows in human non-small cell lung cancer (NSCLC) with low or high TI-CTL/TH1 (CD4/CD8-white, T-bet/CD45RO-red, RGS1-green) (n = 74). Asterisk indicates the location of higher magnification images at right. Scale bar, 50 μm. The mean fluorescent intensity of RGS1 in CTLs (a) and TH1 cells (b) is shown on the right, analyzed by Image J software (mean±s.e.m., low TI-TH1, n = 37; high TI-TH1, n = 37; low TI-CTL, n = 37; high TI-CTL, n = 37). The exact P values are listed in the graph. c,d, Representative histograms of RGS1 expression in CTLs (c) and TH1 (d) isolated from LN, peripheral blood (PB), and tumors (T) of patients with low or high TI-CTL/TH1, determined by flow cytometry. The bars correspond to the mean fluorescent intensity (MFI) of RGS1 (mean±s.e.m. × 104, n = 5). *P < 0.05, **P < 0.01. The exact P values are as follow: P = 0.0070 (CTL in LN), 0.0133 (CTL in PB), 0.0062 (CTL in T), 0.0106 (TH1 in LN), 0.0055 (TH1 in PB), 0.0294 (TH1 in T). e, High RGS1 expression in CTLs or TH1 cells in LN of NSCLC patients, determined by mean fluorescent intensity, is associated with less tumor-infiltrating CTLs or TH1 cells (n = 74) (Pearson’s correlation coefficient R and two-tailed P values). f, Kaplan-Meier survival curves for OS in NSCLC patients with high (n = 37) and low (n = 37) RGS1 expression in CTLs or TH1 cells in LN, determined by the median value of MFI evaluated by immunofluorescent staining. The exact P values are listed in the graph (a-b, c, d, f, g, h). ****P < 0.0001 (b, f). P values were determined by two-tailed Students’ t-test (a-d).
Extended Data Fig. 5 RGS1 has no effect on the proliferation, cytokine secretion, and phenotype of tumor-specific T cells.
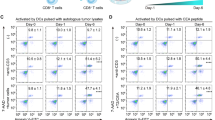
a, Schematic model demonstrating the experimental set up of Transwell migration assay for analyzing the chemotaxis of T cells towards chemokines or tumor conditioned media (CM). b,c, Transwell migration assay for the chemotaxis of CTLs (b) and TH1 (c) with indicated treatment towards PBS, CXCL9 (100ng/ml) or tumor CM (n = 3). The migrated T cells were recorded and counted using time-lapse videomicroscopy at 2-min intervals for 1 h. ****P < 0.0001, **: CTL: CXCL9: shRGS1-1: P = 0.0022, shRGS1-2: P = 0.0024 compared with shGFP-transduced T cells. d,e, CTLs and TH1 cells were isolated from peripheral blood of NSCLC patients, transduced with shvec, shGFP, shRGS1-1, shRGS1-2, labeled with CFSE (green), and overlaid on autologous NSCLC tumor slices. The number of CFSE+ T cells that migrated into the tumor slice was measured by immunofluorescence microscopy. Shown are the number of CFSE+ cells/high power field for each condition (d) and representative images (e) (mean±s.e.m., n = 4. ****P < 0.0001 compared with shvec). Scale bar, 50 µm. f, CFSE-labeled CD8+ or CD4+ T cells were incubated with tumor lysate-pulsed DCs and the proliferation were assessed by flow cytometry. Numbers denote the percentage of cells undergoing at least one cellular division (mean±s.e.m., n = 3). g,h, Tumor-specific CTLs with indicated treatment were analyzed by flow cytometry. Numbers indicate the percentage of gated CD8+ cells stained for perforin, granzyme B, interferon gamma (g), CD69 and CD45RO (h) (mean ± s.e.m., n = 3). i,j, Tumor-specific CD4+ T cells with indicated treatment were analyzed by flow cytometry. Numbers indicate the percentage of gated CD4+ cells stained for interferon gamma (i) or T-bet (j) (mean±s.e.m., n = 3). P values were determined by two-tailed one-way ANOVA with Dunnett’s multiple comparison test (b, c, d).
Extended Data Fig. 6 RGS1 inhibits GPCR-mediated intracellular signaling events.
a, The quantification of the bands of Fig. 4a, determined by Image J, is shown by the fold change relative to 5% input (mean±s.e.m., n = 3). ***P = 0.0002 (RGS1), **P = 0.0014 (CXCR3), P = 0.0012 (CCR4), *P = 0.0135 (CXCR4). b, The levels of cAMP were examined in T cells with indicated treatment and the bars showed the fold change compared with shvec-transduced CTLs treated with PBS (mean±s.e.m., n = 3). ****P < 0.0001 compared with shvec-transduced T cells with indicated treatment. c, The quantification of the bands of Fig. 4d, determined by Image J and normalized to GAPDH, is shown (mean±s.e.m., n = 3). ** P < 0.01, *** P < 0.001, **** P < 0.0001 compared with shvec-transduced CTLs with indicated treatment. The exact P values are follow:RGS1: PBS-shRGS1-1: P = 0.0051, PBS-shRGS1-2: P = 0.0038, CXCL12-shRGS1-1: P = 0.0002, CXCL12-shRGS1-2: P = 0.0004. d, CTLs were isolated from peripheral blood of healthy donors and transduced with shRGS1-1, shRGS1-2, shGFP or shvec, followed by CXCL9 treatment. The RGS1 expression, the phosphorylation and total levels of ERK and Akt in CTLs with indicated treatment were determined by Western blotting (n = 3). The quantification of the bands, determined by Image J and normalized to GAPDH, is shown (mean ± s.e.m., n = 3). *P < 0.05, **P < 0.01, ****P < 0.0001 compared with shvec-transduced CTLs with indicated treatment. The exact P values are follow:RGS1: PBS-shRGS1-1: P = 0.0133, PBS-shRGS1-2: P = 0.0433, CXCL9-shRGS1-1: P = 0.0014, CXCL12-shRGS1-2: P = 0.0065. P values were determined by two-tailed two-way ANOVA with Dunnett’s multiple comparison tests (a-d).
Extended Data Fig. 7 Adoptively transferred CTLs with Rgs1 knockdown reveals enhanced anti-tumor effects in mouse breast and lung cancer models.
a,b, Mice bearing MMTV-PyMT tumors and LLC1 tumor grafts were intravenously injected with the tumor-specific CTLs transduced with shRgs1-1, shRgs1-2, shGFP or shvec and labeled with CFSE (green). a, The tumor grafts were imaged 2 days after T-cell transfusion by intravital two-photon microscopy. The blood vessels were indicated by i.v. injection of Rhodamine-dextrane (red) (n = 6 per group). Scale bar, 100 μm. b, MMTV-PyMT tumors and LLC1 tumor grafts infiltrating CTLs were isolated 7 days after T cell transfusion and analyzed by flow cytometry. Bars correlate to the percentages of gated CD8+ cells stained for CFSE, CD44, CD69, perforin, granzyme B or IFN-γ (mean±s.d.; n = 5 per group). MMTV-PyMT tumor: CFSE: *P = 0.0124 (shvec). IFN-γ: *** P = 0.0003 (shRgs1-1), ** P = 0.0017 (shRgs1-2). Perforin: *** P = 0.0002 (shRgs1-2). ****P < 0.001 compared with tumors without CTLs transfusion. #### P < 0.0001 compared with tumors receiving shvec-transduced CTL transfusion. LLC: CD44: ***P = 0.0006 (shRgs1-2). CD69: ** P = 0.0012 (shRgs1-2). Perforin: ** P = 0.0026 (shRgs1-1). ****P < 0.001 compared with tumors without CTLs transfusion. ####P < 0.0001 compared with tumors receiving shvec-transduced CTL transfusion. c, Quantification of Fig. 5d (mean±s.d.; n = 6 per group). %ID/g, the percentage of injected dose per gram of tissue. ****P < 0.0001. P values were determined by two-tailed one-way ANOVA with Dunnett’s multiple comparison test (b, c).
Extended Data Fig. 8 Adoptively transferred CTLs with RGS1 knockdown reveals enhanced anti-tumor effects in breast cancer PDXs.
a, Representative ER−PR−HER2− PDX is shown in comparison with the original tumor samples, which was immunostained for ER, PR, HER2 and PD-L1 (n = 4 cases of PDX). Scale bar, 50 μm. b, Quantification of Fig. 6b (mean±s.e.m.; n = 4 cases of PDX; ****P < 0.0001). c,d, Representative immunofluorescence images for TUNEL/CD8 (c) or Ki-67/CD8 (d) co-staining in the harvested PDXs (n = 3 cases of PDX per group). White arrows indicate transferred CTLs. Scale bar, 50 μm. The bars at the bottom correspond to the percentages of apoptotic (c) or proliferative (d) CTLs (mean±s.e.m., n = 3 cases of PDX per group). *P = 0.0189 compared with shvec-transduced CTLs. e,f, PDX-infiltrating CTLs were isolated 7 days after T cell transfusion. e, Representative histograms of apoptosis of PDX-infiltrating CTLs, as determined by flow cytometry. Numerical values denote the percentages of cleaved caspase-3+ cells in gated CD8+ T cells (mean±s.e.m., n = 4 cases of PDX). **P = 0.0013 compared with shvec-transduced CTLs. f, The proliferation of transferred CD8+ T cells were assessed by flow cytometry for CFSE+ cells. Numbers denote the percentage of cells undergoing at least one cellular division (mean±s.e.m., n = 4 cases of PDX). g, PDX growth rates of SYMH133, SYMH137 and SYMH145 were determined by tumor volume (mean±s.d., n = 3 mice per group per PDX case). SYMH133: **P = 0.0064; SYMH145: *P = 0.0248 ((-) vs shvec-CTL/αPD-L1), 0.0108 (shRGS1-CTL/IgG vs shRGS1-CTL/αPD-L1); ****P < 0.0001. h, Quantification of Fig. 6f (mean±s.e.m., n = 4 cases of PDX). The exact P values are listed in the graph. ****P < 0.0001. P values were determined by two-tailed one-way ANOVA with Dunnett’s multiple comparison test (b-e) or two-tailed two-way ANOVA with Tukey’s multiple comparison tests (g, h).
Extended Data Fig. 9 IFN-STAT1 signaling in CTLs and TH1 cells upregulates RGS1 and inhibits T cell trafficking.
a, Representative immunofluorescent staining for IFNG expression in cancer cells (CK+), macrophages (CD68+), DC (CD11c+), T cells (CD3+) and natural killer cells (CD56+) in breast tumor tissues with low or high IFNG production (IFNGlow: n = 7, IFNGhigh: n = 5). Asterisk indicates the location of higher magnification images at right. Scale bar, 50 μm. b, Representative plots of flow cytometry analysis of STAT1 phosphorylation (p-STAT1) in the whole tumors (left: grey: isotype, red: p-STAT1) and gated p-STAT1+ cells for different markers staining. Numbers indicate the percentages of different cell types in p-STAT1+ cells (mean±s.e.m.; IFNGlow: n = 7, IFNGhigh: n = 5). c,d, 4T1 cells transduced with sgStat1 (Stat1KO) or a control sgRNA (WT) were inoculated into the fat pad of BALB/c mice, respectively. c, Tumors were collected for protein extraction and the phosphorylation and total levels of Stat1 were determined by Western blotting. Three independent experiments were performed and yielded similar results. α-Tubulin, loading control. d, Tumors were collected 2 days after T cell transfusion and analyzed by flow cytometry. Representative plots of the infiltration of macrophages (F4/80+CD11b+) and DCs (CD11c+) and the p-STAT1 levels in these cells. Numbers indicate the percentages of macrophages and DCs in the tumors and the percentages of p-Stat1+ cells in gated cells (mean ± s.d.; n = 3 per group). *P = 0.0232, **P = 0.0026, ***P = 0.001, ****P < 0.0001 compared with WT grafts by two-tailed Student’s t-test. e, Representative histograms of Rgs1 expression in CD8+ T cells isolated from the spleens of mice bearing 4T1 tumor grafts and transduced with shRgs1, sgStat1 or control lentivirus (shGFP+ctrl-sgRNA), determined by flow cytometry. Numbers in the plots indicate the MFI of Rgs1 (mean±s.d. ×103, n = 3, ****P < 0.0001 compared with ctrl-CTLs by two-tailed one-way ANOVA with Dunnett’s multiple comparison test). f, The scheme of the mechanism underlying RGS1 expression and contribution to the trafficking of CTLs and TH1 cells.
Supplementary information
Supplementary Information
Supplementary Figs. 1 and 2 and Tables 1–5.
Supplementary Video 1
Silencing RGS1 in CTLs enhanced the chemotactic response to CXCL9 and tumor CM. Freshly isolated CTLs transduced with shGFP (stained with CFSE, green), shRGS1-1 (stained with Hoechst 33342, blue) and shRGS1-2 (stained with CellTracker Orange CMTMR Dye, orange) were added to the top chamber, and culture medium without chemokines (PBS), with CXCL9 (100 ng ml−1) or tumor CM was added to the lower chamber of the transwell apparatus. Migrated T cells were recorded and counted using time-lapse videomicroscopy at 2-min intervals for 1 h (n = 3). Quantification of migrated T cells is shown in Extended Data Fig. 5b.
Supplementary Video 2
Silencing RGS1 in TH1 cells enhanced the chemotactic response to CXCL9 and tumor CM. Freshly isolated human TH1 cells transduced with shGFP (stained with CFSE, green), shRGS1-1 (stained with Hoechst 33342, blue) and shRGS1-2 (stained with CellTracker Orange CMTMR Dye, orange) were added to the top chamber, and culture medium without chemokines (PBS), with CXCL9 (100 ng ml−1) or tumor CM was added to the lower chamber of the transwell apparatus. Migrated T cells were recorded and counted using time-lapse videomicroscopy at 2-min intervals for 1 h (n = 3). Quantification of migrated T cells is shown in Extended Data Fig. 5c.
Supplementary Video 3
Silencing RGS1 in CTLs promoted directional migration toward tumor CM in a μ-slide chemotactic chamber. CTLs were transduced with shGFP (stained with CFSE, green), shRGS1-1 (stained with Hoechst 33342, blue) or shRGS1-2 (stained with CellTracker Orange CMTMR Dye, orange) and monitored for migration in a μ-slide chemotaxis chamber with tumor CM (n = 6). Time-lapse videomicroscopy was employed to record the migration of cells at 2-min intervals for 6 h. Migration tracks of the cells are shown in Fig. 3d.
Supplementary Video 4
Silencing RGS1 in TH1 cells promoted directional migration toward tumor CM in a μ-slide chemotactic chamber. TH1 cells were transduced with shGFP (stained with CFSE, green), shRGS1-1 (stained with Hoechst 33342, blue) or shRGS1-2 (stained with CellTracker Orange CMTMR Dye, orange) and monitored for migration in a μ-slide chemotaxis chamber with tumor CM (n = 6). Time-lapse videomicroscopy was employed to record the migration of cells at 2-min intervals for 6 h. Migration tracks of the cells are shown in Fig. 3d.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 4
Unprocessed immunoblots.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 7
Statistical source data.
Source Data Fig. 7
Unprocessed EMSA blots.
Source Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 6
Unprocessed immunoblots.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 9
Unprocessed immunoblots.
Rights and permissions
About this article
Cite this article
Huang, D., Chen, X., Zeng, X. et al. Targeting regulator of G protein signaling 1 in tumor-specific T cells enhances their trafficking to breast cancer. Nat Immunol 22, 865–879 (2021). https://doi.org/10.1038/s41590-021-00939-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-021-00939-9
This article is cited by
-
Tumour circular RNAs elicit anti-tumour immunity by encoding cryptic peptides
Nature (2024)
-
Function and regulation of RGS family members in solid tumours: a comprehensive review
Cell Communication and Signaling (2023)
-
Regulator of G protein signaling protein 6 alleviates acute lung injury by inhibiting inflammation and promoting cell self-renewal in mice
Cellular & Molecular Biology Letters (2023)
-
Anti-PD-1 therapy achieves favorable outcomes in HBV-positive non-liver cancer
Oncogenesis (2023)
-
Regulator of G protein signaling-1 regulates immune infiltration and macrophage polarization in clear cell renal cell carcinoma
International Urology and Nephrology (2023)