Abstract
Evasion of host immunity is a hallmark of cancer; however, mechanisms linking oncogenic mutations and immune escape are incompletely understood. Through loss-of-function screening of 1,001 tumor suppressor genes, we identified death-associated protein kinase 3 (DAPK3) as a previously unrecognized driver of anti-tumor immunity through the stimulator of interferon genes (STING) pathway of cytosolic DNA sensing. Loss of DAPK3 expression or kinase activity impaired STING activation and interferon (IFN)-β-stimulated gene induction. DAPK3 deficiency in IFN-β-producing tumors drove rapid growth and reduced infiltration of CD103+CD8α+ dendritic cells and cytotoxic lymphocytes, attenuating the response to cancer chemo-immunotherapy. Mechanistically, DAPK3 coordinated post-translational modification of STING. In unstimulated cells, DAPK3 inhibited STING K48-linked poly-ubiquitination and proteasome-mediated degradation. After cGAMP stimulation, DAPK3 was required for STING K63-linked poly-ubiquitination and STING–TANK-binding kinase 1 interaction. Comprehensive phospho-proteomics uncovered a DAPK3-specific phospho-site on the E3 ligase LMO7, critical for LMO7–STING interaction and STING K63-linked poly-ubiquitination. Thus, DAPK3 is an essential kinase for STING activation that drives tumor-intrinsic innate immunity and tumor immune surveillance.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Screening results are presented in Supplementary Table 1 and phospho-proteomics results in Supplementary Table 2. Mass spectrometry proteome and phosphoproteome data were deposited in MassIVE (identifier PXD023639) and ProteomeXchange (identifier PXD023637). Uncropped immunoblot images are provided in the manuscript. Source data are provided with this paper. Additional data will be made available from the corresponding author upon reasonable request.
References
Zitvogel, L., Galluzzi, L., Kepp, O., Smyth, M. J. & Kroemer, G. Type I interferons in anticancer immunity. Nat. Rev. Immunol. 15, 405–414 (2015).
Vanpouille-Box, C., Demaria, S., Formenti, S. C. & Galluzzi, L. Cytosolic DNA sensing in organismal tumor control. Cancer Cell 34, 361–378 (2018).
Kwon, J. & Bakhoum, S. F. The cytosolic DNA-sensing cGAS–STING pathway in cancer. Cancer Discov. 10, 26–39 (2020).
Chen, Q., Sun, L. & Chen, Z. J. Regulation and function of the cGAS–STING pathway of cytosolic DNA sensing. Nat. Immunol. 17, 1142–1149 (2016).
Corrales, L., McWhirter, S. M., Dubensky, T. W. Jr. & Gajewski, T. F. The host STING pathway at the interface of cancer and immunity. J. Clin. Investig. 126, 2404–2411 (2016).
Woo, S. R. et al. STING-dependent cytosolic DNA sensing mediates innate immune recognition of immunogenic tumors. Immunity 41, 830–842 (2014).
Harding, S. M. et al. Mitotic progression following DNA damage enables pattern recognition within micronuclei. Nature 548, 466–470 (2017).
Mackenzie, K. J. et al. cGAS surveillance of micronuclei links genome instability to innate immunity. Nature 548, 461–465 (2017).
Ahn, J. et al. Inflammation-driven carcinogenesis is mediated through STING. Nat. Commun. 5, 5166 (2014).
Yang, H., Wang, H., Ren, J., Chen, Q. & Chen, Z. J. cGAS is essential for cellular senescence. Proc. Natl Acad. Sci. USA 114, E4612–E4620 (2017).
Wang, Z. et al. cGAS/STING axis mediates a topoisomerase II inhibitor-induced tumor immunogenicity. J. Clin. Invest. 129, 4850–4862 (2019).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Brognard, J., Zhang, Y. W., Puto, L. A. & Hunter, T. Cancer-associated loss-of-function mutations implicate DAPK3 as a tumor-suppressing kinase. Cancer Res. 71, 3152–3161 (2011).
Lio, C. W. et al. cGAS-STING signaling regulates initial innate control of cytomegalovirus infection. J. Virol. 90, 7789–7797 (2016).
Farag, A. K. & Roh, E. J. Death-associated protein kinase (DAPK) family modulators: current and future therapeutic outcomes. Med. Res. Rev. 39, 349–385 (2019).
Fang, J. et al. Attenuation of EPO-dependent erythroblast formation by death-associated protein kinase-2. Blood 112, 886–890 (2008).
Willemsen, J. et al. Phosphorylation-Dependent Feedback Inhibition of RIG-I by DAPK1 Identified by Kinome-wide siRNA Screening. Mol. Cell 65, 403–415.e8 (2017).
Kocher, B. A., White, L. S. & Piwnica-Worms, D. DAPK3 suppresses acini morphogenesis and is required for mouse development. Mol. Cancer Res. 13, 358–367 (2014).
Hosoba, K. et al. Phosphorylation of myosin II regulatory light chain by ZIP kinase is responsible for cleavage furrow ingression during cell division in mammalian cultured cells. Biochem. Biophys. Res. Commun. 459, 686–691 (2015).
Demaria, O. et al. STING activation of tumor endothelial cells initiates spontaneous and therapeutic antitumor immunity. Proc. Natl Acad. Sci. USA 112, 15408–15413 (2015).
Mitchison, T. J., Pineda, J., Shi, J. & Florian, S. Is inflammatory micronucleation the key to a successful anti-mitotic cancer drug? Open Biol. 7, 170182 (2017).
Zierhut, C. et al. The cytoplasmic DNA sensor cGAS promotes mitotic cell death. Cell 178, 302–315.e23 (2019).
Lohard, S. et al. STING-dependent paracriny shapes apoptotic priming of breast tumors in response to anti-mitotic treatment. Nat. Commun. 11, 259 (2020).
Luo, W. W. et al. iRhom2 is essential for innate immunity to DNA viruses by mediating trafficking and stability of the adaptor STING. Nat. Immunol. 17, 1057–1066 (2016).
Graves, P. R., Winkfield, K. M. & Haystead, T. A. Regulation of zipper-interacting protein kinase activity in vitro and in vivo by multisite phosphorylation. J. Biol. Chem. 280, 9363–9374 (2005).
Okamoto, M. et al. Identification of death-associated protein kinases inhibitors using structure-based virtual screening. J. Med. Chem. 52, 7323–7327 (2009).
Zhang, J., Hu, M. M., Wang, Y. Y. & Shu, H. B. TRIM32 protein modulates type I interferon induction and cellular antiviral response by targeting MITA/STING protein for K63-linked ubiquitination. J. Biol. Chem. 287, 28646–28655 (2012).
Tsuchida, T. et al. The ubiquitin ligase TRIM56 regulates innate immune responses to intracellular double-stranded DNA. Immunity 33, 765–776 (2010).
Liu, S. et al. Phosphorylation of innate immune adaptor proteins MAVS, STING, and TRIF induces IRF3 activation. Science 347, aaa2630 (2015).
Tsuchiya, Y., Jounai, N., Takeshita, F., Ishii, K. J. & Mizuguchi, K. Ligand-induced ordering of the C-terminal tail primes STING for phosphorylation by TBK1. EBioMedicine 9, 87–96 (2016).
Zhao, B. et al. A conserved PLPLRT/SD motif of STING mediates the recruitment and activation of TBK1. Nature 569, 718–722 (2019).
Tanaka, Y. & Chen, Z. J. STING specifies IRF3 phosphorylation by TBK1 in the cytosolic DNA signaling pathway. Sci. Signal. 5, ra20 (2012).
Lapek, J. D. Jr., Lewinski, M. K., Wozniak, J. M., Guatelli, J. & Gonzalez, D. J. Quantitative temporal viromics of an inducible HIV-1 model yields insight to global host targets and phospho-dynamics associated with protein Vpr. Mol. Cell. Proteomics 16, 1447–1461 (2017).
Burch, L. R., Scott, M., Pohler, E., Meek, D. & Hupp, T. Phage-peptide display identifies the interferon-responsive, death-activated protein kinase family as a novel modifier of MDM2 and p21WAF1. J. Mol. Biol. 337, 115–128 (2004).
Arif, A., Chatterjee, P., Moodt, R. A. & Fox, P. L. Heterotrimeric GAIT complex drives transcript-selective translation inhibition in murine macrophages. Mol. Cell. Biol. 32, 5046–5055 (2012).
Schmid, J. A. & Birbach, A. IκB kinase β (IKKβ/IKK2/IKBKB)—a key molecule in signaling to the transcription factor NF-κB. Cytokine Growth Factor Rev. 19, 157–165 (2008).
Nehru, V., Almeida, F. N. & Aspenström, P. Interaction of RhoD and ZIP kinase modulates actin filament assembly and focal adhesion dynamics. Biochem. Biophys. Res. Commun. 433, 163–169 (2013).
Tang, H. W. et al. Atg1-mediated myosin II activation regulates autophagosome formation during starvation-induced autophagy. EMBO J. 30, 636–651 (2011).
Ni, G., Konno, H. & Barber, G. N. Ubiquitination of STING at lysine 224 controls IRF3 activation. Sci. Immunol. 2, eaah7119 (2017).
Yang, L. et al. UBXN3B positively regulates STING-mediated antiviral immune responses. Nat. Commun. 9, 2329 (2018).
Mullard, A. Can innate immune system targets turn up the heat on ‘cold’ tumours? Nat. Rev. Drug Discov. 17, 3–5 (2018).
Mallipeddi, R. et al. Reduced expression of insulin-like growth factor-binding protein-3 (IGFBP-3) in squamous cell carcinoma complicating recessive dystrophic epidermolysis bullosa. J. Invest. Dermatol. 122, 1302–1309 (2004).
Bi, J. et al. Downregulation of ZIP kinase is associated with tumor invasion, metastasis and poor prognosis in gastric cancer. Int. J. Cancer 124, 1587–1593 (2009).
Song, S. et al. Decreased expression of STING predicts poor prognosis in patients with gastric cancer. Sci. Rep. 7, 39858 (2017).
Puto, L. A. & Reed, J. C. Daxx represses RelB target promoters via DNA methyltransferase recruitment and DNA hypermethylation. Genes Dev. 22, 998–1010 (2008).
Seo, G. J. et al. TRIM56-mediated monoubiquitination of cGAS for cytosolic DNA sensing. Nat. Commun. 9, 613 (2018).
Wang, Q. et al. The E3 ubiquitin ligase AMFR and INSIG1 bridge the activation of TBK1 kinase by modifying the adaptor STING. Immunity 41, 919–933 (2014).
Kawai, T. & Akira, S. Signaling to NF-κB by Toll-like receptors. Trends Mol. Med. 13, 460–469 (2007).
Balka, K. R. et al. TBK1 and IKKε act redundantly to mediate STING-induced NF-κB responses in myeloid cells. Cell Rep. 31, 107492 (2020).
Pokatayev, V. et al. Homeostatic regulation of STING protein at the resting state by stabilizer TOLLIP. Nat. Immunol. 21, 158–167 (2020).
Dhanwani, R. et al. Cellular sensing of extracellular purine nucleosides triggers an innate IFN-β response. Sci. Adv. 6, eaba3688 (2020).
Zhao, M., Sun, J. & Zhao, Z. TSGene: a web resource for tumor suppressor genes. Nucleic Acids Res. 41, D970–D976 (2013).
Sharma, S. & Rao, A. RNAi screening: tips and techniques. Nat. Immunol. 10, 799–804 (2009).
Sharma, S. et al. An siRNA screen for NFAT activation identifies septins as coordinators of store-operated Ca2+ entry. Nature 499, 238–242 (2013).
Wang, Y. et al. Reversed-phase chromatography with multiple fraction concatenation strategy for proteome profiling of human MCF10A cells. Proteomics 11, 2019–2026 (2011).
Elias, J. E. & Gygi, S. P. Target-decoy search strategy for increased confidence in large-scale protein identifications by mass spectrometry. Nat. Methods 4, 207–214 (2007).
Wozniak, J. M. & Gonzalez, D. J. PTMphinder: an R package for PTM site localization and motif extraction from proteomic datasets. PeerJ 7, e7046 (2019).
Xu, S., Feng, Y. & Zhao, S. Proteins with evolutionarily hypervariable domains are associated with immune response and better survival of basal-like breast cancer patients. Comput. Struct. Biotechnol. J. 17, 430–440 (2019).
Cerami, E. et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2, 401–404 (2012).
Gao, J. et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. 6, pl1 (2013).
Acknowledgements
We thank J. Huang (LJI) for support for the HUVEC siRNA screen; S. Shresta (LJI) for Ifnar1-null mice; C. Benedict (LJI) for MEFs, NHDFs and hCMV; N. P. Restifo (NCI) for MCA205; J. Schlom (NCI) for MC38; S. Schoenberger (LJI) for B16F10; C. C. Hedrick (LJI) for LLC-RFP; Y. C. Liu (LJI) for pEF-neo-HA-Ub; D. Zajonc (LJI) for pGEX-4T-2; J. Day and K. Tanguay (LJI) for shRNA and siRNA constructs, and siRNA screen support; D. Freeman, B. McDonald and R. El Morabiti (LJI) for hCMV propagation; M. Diep, K. Foos and M. Kaur (LJI) for technical assistance; Z. Mikulski (LJI Microscopy Core Facility) for confocal microscopy support; A. Sethi (LJI Bioinformatics Core Facility) for pathway analysis support; LJI Flow Cytometry Core Facility for cell sorting (FACSAria-II Cell Sorter; supported by the Shared Instrumentation Grant (SIG) program no. S10 RR027366); and D. Araujo (LJI) for proofreading the manuscript. This work was supported by National Institutes of Health (NIH) grant nos. R01CA199376, U01DE028227 and U54CA260591 (S.S.); and NIH grant no. S10OD020025 and grant no. R01ES027595 (M.J.). C.-W.J.L. was supported by a Cancer Research Institute (CRI) Irvington Postdoc Fellowship. A.C. was supported by the UCSD Microbial Sciences Initiative Graduate Research Fellowship and by the UCSD Graduate Training Program in Cellular and Molecular Pharmacology, through an institutional training grant from the National Institute of General Medical Sciences, grant no. T32 GM007752.
Author information
Authors and Affiliations
Contributions
M.T. designed, optimized and performed in vitro and in vivo experiments and the siRNA screen in THP1-Blue ISG cells. C.-W.J.L. optimized and performed the siRNA screen in HUVECs and generated L929 reporter cells. A.C. optimized and performed phospho-proteomics in THP1-Blue ISG cells under the supervision of D.J.G. M.S. optimized and performed mass spectrometry analysis in HEK293T and L929 lysates and supported phospho-proteomics under the supervision of M.M. F.A. provided support for bioinformatics analyses. M.J. provided support for mass spectrometry studies. S.S. provided overall direction and supervision. M.T. and S.S. wrote the manuscript with input from co-authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Immunology thanks Katherine A. Fitzgerald, Hong-Bing Shu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available. L. A. Dempsey was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 DAPK3 is a positive regulator of STING signaling and some TLR pathways.
a, Schematic representation of the RNAi screen for IRF3 nuclear translocation. b, Immunostaining of IRF3 in HUVEC stimulated with poly (dA:dT) (1 μg/ml) (right panel) for 3 h. Scale bar, 100 μm. c, d, qRT-PCR of DAPK1/Dapk1, DAPK2/Dapk2, and DAPK3/Dapk3 in (c) human and (d) mouse cell lines. #, Not detected. e, Images of IRF3 localization in L929-mRuby-hIRF3 stimulated with poly (dA;dT) (1 μg/ml) for 3 h. Scale bar, 100 μm. f, g, (f) qRT-PCR of IFNB1 and (g) immunoblot of HUVEC transfected with indicated siRNA. h, i, (h) qRT-PCR of Ifnb1 and (i) immunoblot of BMDM transfected with indicated siRNA. (f, h) Cells were stimulated with poly (dA:dT) (0.2 μg/ml) for 4 h. j, k, (j) qRT-PCR of Il6 in L929-mRuby-hIRF3 and (k) IL6 inTHP1-Blue ISG transduced with indicated shRNA and stimulated as indicated in Fig. 1g-l. l, m, qRT-PCR of (l) Ifnb1 in L929-mRuby-hIRF3 and (m) IFNB1 inTHP1-Blue ISG transduced with indicated shRNA and transfected with poly (I:C)(LMW) and poly (I:C)(HMW)(0.1 μg/ml for L929-mRuby-hIRF3, 0.5 μg/ml for THP1-Blue ISG). n, o, qRT-PCR of (n) IFNB1 and (o) IL6 in THP1-Blue ISG transduced with indicated shRNA and stimulated with FSL-1(100 ng/ml), naked poly (I:C)(LMW)(10 μg/ml), or LPS(100 ng/ml) for 4 h. p, q, Immunoblot of (p) THP1-Blue ISG and (q) L929-mRuby-hIRF3 transduced with indicated shRNA. Data in (b-e, g, i, p, q) are representative or (f, h, j-o) the mean of three independent experiments. Values represent mean ± s.d. *P < 0.05, **P < 0.01, and ***P < 0.001. Statistical comparisons were conducted using two-tailed t-test (f, h, j-o).
Extended Data Fig. 2 Association of DAPK3 with outcomes in human cancer.
Kaplan-Meier survival analysis of pancreatic adenocarcinoma, uterine corpus endometrial carcinoma, and esophageal carcinoma comparing the top (high) and bottom (low) tertiles of patients with respect to DAPK3 expression levels as reported by TCGA data portal. Statistical comparisons were conducted using two-sided log-rank test58.
Extended Data Fig. 3 Teniposide and paclitaxel induce micronuclei formation and anti-tumor immunity to B16F10 tumors in a type I IFN signaling-dependent manner.
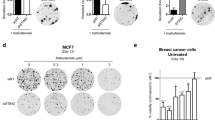
a, b, (Left) Immunoblot and (right) in vitro cell growth of (a) MCA205 and (b) B16F10 transduced with indicated shRNA. c, Flow cytometry of tumor-infiltrating CD8+T cells and CD103+CD8α+DCs in MCA205 tumor suspensions isolated from WT or Ifnar1-KO mice on Day 6 (n = 6 per group). d, Confocal fluorescence microscopy of MCA205 stably expressing cGAS-Clover. Scale bar, 10 μm. e, qRT-PCR of Ifnb1 in unstimulated MCA205 and B16F10 transduced with indicated shRNA. f, (Left) Immunoblot and (right) qRT-PCR of Ifnb1 in shDapk3#1-transduced MCA205 ectopically expressing V5-tagged DAPK3(WT) or DAPK3(D161A). Cells were stimulated with 2′,3′-cGAMP, 3′,3′-cGAMP, c-di-GMP (20 μg/ml for all three agonists) or DMXAA (50 μg/ml) for 4 h. g, Confocal fluorescence microscopy of B16F10 stably expressing cGAS-Clover. Cells were treated with teniposide (10 μM) for 24 h or paclitaxel (100 nM) for 72 h. Scale bar, 10 μm. h, (Left) Apoptosis measured in shRNA-transduced B16F10 treated with teniposide (10 μM) for 24 h or (right) paclitaxel (100 nM) for 48 h. i, Tumor volume of B16F10 subcutaneously transplanted into WT and Ifnar1-KO mice and treated with teniposide or paclitaxel (n = 6 for vehicle, n = 7 for teniposide and paclitaxel). Tumor size on Day 15 is represented (right panel). Data are representative (a-d, g-i) or the mean (a, b, e, f) of three independent experiments. Values represented mean ± s.d. *P < 0.05, **P < 0.01, and ***P < 0.001. Statistical comparisons were conducted using two-tailed t-test (a-c, e, f, h, i).
Extended Data Fig. 4 Flow cytometry gating strategy for tumor-infiltrating leukocytes.
Tumor single cell suspensions were stained with different fluorophore-conjugated antibodies and analyzed by flow cytometry.
Extended Data Fig. 5 DAPK3 does not directly phosphorylate STING or TBK1.
a-d, (a) qRT-PCR of Sting1 and Dapk3, (b) immunoblot, (c) IRF3 nuclear translocation, and (d) p65 nuclear translocation in L929-mRuby-hIRF3 transduced with indicated shRNA stimulated with poly (dA:dT) (0.5 μg/ml) or VACV70 (2 μg/ml) for 3 h. e, Immunoblot of L929-mRuby-hIRF3 transfected with indicated siRNA. f, Immunoblot of L929-mRuby-hIRF3 transduced with indicated shRNA stimulated with VACV70 (2 μg/ml) for 2 h and 4 h. g, Immunoblot of HUVEC stably expressing V5-tagged DAPK3(D161A), DAPK3(T180A), or luciferase. Cells were infected at MOI = 5, 2, or 1. h, Immunoblot of THP1-Blue ISG stably expressing V5-tagged DAPK3(WT) or DAPK3(D161A). i, qRT-PCR of Ifnb1 in L929-mRuby-hIRF3 pre-treated with DAPK inhibitors for 3 h prior to 2′,3′-cGAMP stimulation (10 μg/ml) for 4 h. j, qRT-PCR of IFNB1 in THP1-Blue ISG pre-treated with DAPK inhibitors (50 μM) for 6 h prior to 2′,3′-cGAMP or c-di-GMP stimulation (10 μg/ml for both) for 4 h. k, l, In vitro kinase assay of (k) GST-tagged human STING C-terminus (aa 149-379) and (l) GST-tagged human TBK1(K38M). Peptides were incubated with GST-tagged DAPK3 or TBK1 in the presence of [γ-32P] ATP. Data in (b, e-h, k, l) are representative or (a, c, d, i, j) mean of three independent experiments. Values represent mean ± s.d. *P < 0.05, **P < 0.01, and ***P < 0.001. Statistical comparisons were conducted using two-tailed t-test (a, c, d, i, j).
Extended Data Fig. 6 DAPK3 is not involved in STING trafficking from ER to Golgi.
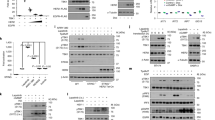
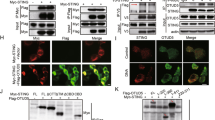
a, Confocal fluorescence microscopy of THP1-Blue ISG transduced with indicated shRNA and unstimulated or stimulated with 2′,3′-cGAMP (25 μg/ml) for 3 h. Scale bar, 15 μm. b, (Upper) Co-localization of STING/Calreticulin and (lower) STING/GM130 analyzed using Image J software. Data are pooled from three independent experiments (n > 1,500 cells for unstimulated 32 images and cGAMP-stimulated 73 images). c, (Upper) Confocal fluorescence microscopy of THP1-Blue ISG stably expressing GFP-tagged DAPK3(WT) unstimulated or stimulated with 2′,3′-cGAMP (50 μg/ml) for 3 h. Localization of GFP-DAPK3, STING, and TBK1. (Lower) Co-localization of GFP-DAPK3/TBK1, GFP-DAPK3/STING, and TBK1/STING was analyzed using Image J software. Data are pooled from three independent experiments (n > 1,500 cells for unstimulated and cGAMP-stimulated 70 images). Scale bars, 15 μm. d, Schematic representation of human STING mutants. e, (Upper) Immunoprecipitation and immunoblot of HEK293T transfected with plasmid encoding HA-tagged human STING (WT, 1-379), phospho-deficient mutant (3S-3A), or C-terminal deletion mutant (aa 1-340) unstimulated or stimulated with 2′,3′-cGAMP (5 μg/ml) for 2 h, and (lower) immunoblot of whole cell lysates (WCL). Values represented as mean ± s.d. Data in (a, c, e) are representative of three independent experiments.
Extended Data Fig. 7 Phosphorylation of TRIP12 on S312 or TRIM56 on T442 are not involved in STING K63-linked poly ubiquitination.
a, Primary RNAi screen of E3 ligases in THP1-Blue ISG transfected with indicated siRNA. SEAP activity was measured after normalization with CellTiter-Glo. Black; siControl, Blue; previously reported E3 ligases for K63-linked poly-ubiquitination of STING, Red; positive control (for example siSTING1 and siTBK1). siTRIM56 value was used for determining cut-off. b, Secondary RNAi screen of E3 ligases in THP1-Blue ISG transfected with indicated siRNA. qRT-PCR of IFNB1 was performed. #; candidates for subsequent analysis. c, d, In vitro kinase assay of (c) GST-tagged human TRIP12 peptide (aa 260-360) and (d) GST-tagged human TRIM56 peptide (aa 400-500). Peptides were incubated with GST-tagged DAPK3 or TBK1 in the presence of [γ-32P] ATP. e, Schematic representation of human TRIP12 mutants. f, (Upper) Immunoprecipitation and immunoblot of HEK293T transfected with plasmids encoding HA-tagged human STING and V5-tagged human TRIP12 (WT) or phospho-deficient TRIP12 (S312A), and (lower) immunoblot of whole cell lysates (WCL). g, (Upper) Immunoprecipitation and immunoblot of HEK293T transfected with plasmids encoding 3×Flag-tagged human STING, HA-tagged Ub(K63O), and V5-tagged human TRIP12(WT), phospho-deficient TRIP12(S312A), or HECT domain-deficient TRIP12(ΔHECT), and (lower) immunoblot of WCL. h, (Upper) Immunoprecipitation and immunoblot of HEK293T transfected with plasmids encoding 3×Flag-tagged human STING, HA-tagged Ub(K63O), and V5-tagged human TRIM56(WT), phospho-deficient TRIM56(T442A), or enzyme-inactive TRIM56(C24S), and (lower) immunoblot of WCL. Data in (c, d, f-h) are representative of three independent experiments. Values represent mean ± s.d. *P < 0.05, **P < 0.01, and ***P < 0.001 (compared to siControl) (a, b). Statistical comparisons were conducted using two-tailed t-test (a, b).
Extended Data Fig. 8 DAPK3, LMO7, and TRIP12 are highly mutated in human cancers.
a-c, Genomic alterations of (a) DAPK3, (b) LMO7 and (c) TRIP12 in human cancers from cBioportal.
Extended Data Fig. 9 LMO7 and TRIP12 are positive regulators of STING-IFNβ signaling in THP1 and HUVEC.
a, In vitro kinase assay of GST-tagged human LMO7 (aa 360-460). Peptides were incubated with GST-tagged DAPK3 or TBK1 in the presence of [γ-32P] ATP. b, (Upper) Immunoprecipitation and immunoblot of HEK293T transduced with indicated shRNA prior to transfection with plasmids encoding 3×Flag-tagged human STING, HA-tagged Ub(K63O), and V5-tagged human LMO7(WT) and (lower) immunoblot of whole cell lysates (WCL). c, (Upper) Immunoblot of THP1-Blue ISG transduced with two distinct shLMO7 or (lower) shTRIP12 sequences. d, (Upper) Immunoprecipitation and immunoblot of THP1-Blue ISG transduced with indicated shRNA and stimulated with 2′,3′-cGAMP (10 μg/ml) for 3 h and 6 h, and (lower) immunoblot of WCL. e, f, Immunoblot of THP1-Blue ISG transduced with two distinct (e) shLMO7 or (f) shTRIP12 sequences and stimulated with 2′,3′-cGAMP (10 μg/ml) for 3 h and 6 h. g, h, Immunoblot of (g) THP1-Blue ISG and (h) HUVEC transfected with indicated siRNA. i, j, qRT-PCR of IFNB1 and CXCL10 in (i) THP1-Blue ISG and (j) HUVEC transfected with indicated siRNA stimulated with VACV70 (2 μg/ml), 2′,3′-cGAMP (10 μg/ml), and c-di-GMP (10 μg/ml). Data in (a-h) are representative or (i, j) mean of three independent experiments. Values represent mean ± s.d. *P < 0.05, **P < 0.01, and ***P < 0.001. Statistical comparisons were conducted using two-tailed t-test (i, j).
Extended Data Fig. 10 Schematic model of the DAPK3-STING axis.
In unstimulated cells (L929 and MCA205), DAPK3 maintains steady-state STING levels by inhibiting STING K48-linked poly-ubiquitination and proteasome-mediated degradation. In DNA-stimulated cells (THP1), DAPK3 promotes STING activation by phosphorylating the E3 ligase LMO7 at S863, enabling LMO7-STING interaction, STING K63-linked poly-ubiquitination, and recruitment of TBK1.
Supplementary information
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 3
Unprocessed immunoblot images.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 4
Unprocessed immunoblot images.
Source Data Fig. 5
Unprocessed immunoblot images.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 6
Unprocessed immunoblot images.
Source Data Fig. 7
Statistical source data.
Source Data Fig. 7
Unprocessed immunoblot images.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 1
Unprocessed immunoblot images.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 3
Unprocessed immunoblot images.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 5
Unprocessed immunoblot images.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 6
Unprocessed immunoblot images.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 7
Unprocessed immunoblot images.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 9
Unprocessed immunoblot images.
Rights and permissions
About this article
Cite this article
Takahashi, M., Lio, CW.J., Campeau, A. et al. The tumor suppressor kinase DAPK3 drives tumor-intrinsic immunity through the STING–IFN-β pathway. Nat Immunol 22, 485–496 (2021). https://doi.org/10.1038/s41590-021-00896-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-021-00896-3
This article is cited by
-
O-GlcNAc of STING mediates antiviral innate immunity
Cell Communication and Signaling (2024)
-
Harnessing innate immune pathways for therapeutic advancement in cancer
Signal Transduction and Targeted Therapy (2024)
-
ABLIM1, a novel ubiquitin E3 ligase, promotes growth and metastasis of colorectal cancer through targeting IĸBα ubiquitination and activating NF-ĸB signaling
Cell Death & Differentiation (2024)
-
PES1 reduces CD8+ T cell infiltration and immunotherapy sensitivity via interrupting ILF3-IL15 complex in esophageal squamous cell carcinoma
Journal of Biomedical Science (2023)
-
ESCRT-dependent STING degradation inhibits steady-state and cGAMP-induced signalling
Nature Communications (2023)