Abstract
The diversity of regulatory T (Treg) cells in health and in disease remains unclear. Individuals with colorectal cancer harbor a subpopulation of RORγt+ Treg cells with elevated expression of β-catenin and pro-inflammatory properties. Here we show progressive expansion of RORγt+ Treg cells in individuals with inflammatory bowel disease during inflammation and early dysplasia. Activating Wnt–β-catenin signaling in human and murine Treg cells was sufficient to recapitulate the disease-associated increase in the frequency of RORγt+ Treg cells coexpressing multiple pro-inflammatory cytokines. Binding of the β-catenin interacting partner, TCF-1, to DNA overlapped with Foxp3 binding at enhancer sites of pro-inflammatory pathway genes. Sustained Wnt–β-catenin activation induced newly accessible chromatin sites in these genes and upregulated their expression. These findings indicate that TCF-1 and Foxp3 together limit the expression of pro-inflammatory genes in Treg cells. Activation of β-catenin signaling interferes with this function and promotes the disease-associated RORγt+ Treg phenotype.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
RNA-seq, ChIP–seq and ATAC-seq datasets have been deposited in the Gene Expression Omnibus under accession code GSE139960. All raw data and source data that support the findings of this study are available from the corresponding authors upon request.
References
Kuswanto, W. et al. Poor repair of skeletal muscle in aging mice reflects a defect in local, interleukin-33-dependent accumulation of regulatory T cells. Immunity 44, 355–367 (2016).
Ito, M. et al. Brain regulatory T cells suppress astrogliosis and potentiate neurological recovery. Nature 565, 246–250 (2019).
Zhu, J., Yamane, H. & Paul, W. E. Differentiation of effector CD4 T cell populations. Annu. Rev. Immunol. 28, 445–489 (2010).
Levine, A. G. et al. Stability and function of regulatory T cells expressing the transcription factor T-bet. Nature 546, 421–425 (2017).
Yu, F., Sharma, S., Edwards, J., Feigenbaum, L. & Zhu, J. Dynamic expression of transcription factors T-bet and GATA-3 by regulatory T cells maintains immunotolerance. Nat. Immunol. 16, 197–206 (2015).
Ohnmacht, C. et al. The microbiota regulates type 2 immunity through RORγt+ T cells. Science 349, 989–993 (2015).
Sefik, E. et al. Individual intestinal symbionts induce a distinct population of RORγ+ regulatory T cells. Science 349, 993–997 (2015).
Yang, B. H. et al. Foxp3+ T cells expressing RORγt represent a stable regulatory T cell effector lineage with enhanced suppressive capacity during intestinal inflammation. Mucosal Immunol. 9, 444–457 (2016).
Samstein, R. M. et al. Foxp3 exploits a pre-existent enhancer landscape for regulatory T cell lineage specification. Cell 151, 153–166 (2012).
Fu, W. et al. A multiply redundant genetic switch ‘locks in’ the transcriptional signature of regulatory T cells. Nat. Immunol. 13, 972–980 (2012).
Delacher, M. et al. Genome-wide DNA-methylation landscape defines specialization of regulatory T cells in tissues. Nat. Immunol. 18, 1160–1172 (2017).
Kwon, H. K., Chen, H. M., Mathis, D. & Benoist, C. Different molecular complexes that mediate transcriptional induction and repression by FoxP3. Nat. Immunol. 18, 1238–1248 (2017).
Xing, S. et al. Tcf1 and Lef1 are required for the immunosuppressive function of regulatory T cells. J. Exp. Med. 216, 847–866 (2019).
Emmanuel, A. O. et al. TCF-1 and HEB cooperate to establish the epigenetic and transcription profiles of CD4+CD8+ thymocytes. Nat. Immunol. 19, 1366–1378 (2018).
Choi, Y. S. et al. LEF-1 and TCF-1 orchestrate TFH differentiation by regulating differentiation circuits upstream of the transcriptional repressor Bcl6. Nat. Immunol. 16, 980–990 (2015).
Zhou, X. et al. Differentiation and persistence of memory CD8+ T cells depend on T cell factor 1. Immunity 33, 229–240 (2010).
Scott, A. C. et al. TOX is a critical regulator of tumour-specific T cell differentiation. Nature 571, 270–274 (2019).
MacDonald, B. T., Tamai, K. & He, X. Wnt/β-catenin signaling: components, mechanisms and diseases. Dev. Cell 17, 9–26 (2009).
Lovatt, M. & Bijlmakers, M.J. Stabilisation of β-catenin downstream of T cell receptor signalling. PloS ONE 5, e12794 (2010).
van Loosdregt, J. et al. Canonical Wnt signaling negatively modulates regulatory T cell function. Immunity 39, 298–310 (2013).
Sumida, T. et al. Activated β-catenin in Foxp3+ regulatory T cells links inflammatory environments to autoimmunity. Nat. Immunol. 19, 1391–1402 (2018).
Mittal, D., Gubin, M. M., Schreiber, R. D. & Smyth, M. J. New insights into cancer immunoediting and its three component phases–elimination, equilibrium and escape. Curr. Opin. Immunol. 27, 16–25 (2014).
Wing, J. B., Tanaka, A. & Sakaguchi, S. Human FOXP3+ regulatory T cell heterogeneity and function in autoimmunity and cancer. Immunity 50, 302–316 (2019).
Salama, P. et al. Tumor-infiltrating FOXP3+ T regulatory cells show strong prognostic significance in colorectal cancer. J. Clin. Oncol. 27, 186–192 (2009).
deLeeuw, R. J., Kost, S. E., Kakal, J. A. & Nelson, B. H. The prognostic value of FoxP3+ tumor-infiltrating lymphocytes in cancer: a critical review of the literature. Clin. Cancer Res. 18, 3022–3029 (2012).
Sinicrope, F. A. et al. Intraepithelial effector CD3+/regulatory FoxP3+ T cell ratio predicts a clinical outcome of human colon carcinoma. Gastroenterology 137, 1270–1279 (2009).
Saito, T. et al. Two FoxP3+CD4+ T cell subpopulations distinctly control the prognosis of colorectal cancers. Nat. Med. 22, 679–684 (2016).
Feagins, L. A., Souza, R. F. & Spechler, S. J. Carcinogenesis in IBD: potential targets for the prevention of colorectal cancer. Nat. Rev. Gastroenterol. Hepatol. 6, 297–305 (2009).
Terzic, J., Grivennikov, S., Karin, E. & Karin, M. Inflammation and colon cancer. Gastroenterology 138, 2101–2114 (2010).
Blatner, N. R. et al. Expression of RORγt marks a pathogenic regulatory T cell subset in human colon cancer. Sci. Transl. Med. 4, 164ra159 (2012).
Keerthivasan, S. et al. β-Catenin promotes colitis and colon cancer through imprinting of pro-inflammatory properties in T cells. Sci. Transl. Med. 6, 225ra228 (2014).
Miyara, M. et al. Functional delineation and differentiation dynamics of human CD4+ T cells expressing the FoxP3 transcription factor. Immunity 30, 899–911 (2009).
Pratama, A., Schnell, A., Mathis, D. & Benoist, C. Developmental and cellular age direct conversion of CD4+ T cells into RORγ+ or Helios+ colon Treg cells. J. Exp. Med. 217, e20190428 (2019).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Kanehisa, M., Furumichi, M., Tanabe, M., Sato, Y. & Morishima, K. KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 45, D353–D361 (2017).
Tosolini, M. et al. Clinical impact of different classes of infiltrating T cytotoxic and helper cells (TH1, TH2, Treg, TH17) in patients with colorectal cancer. Cancer Res. 71, 1263–1271 (2011).
Beurel, E., Michalek, S. M. & Jope, R. S. Innate and adaptive immune responses regulated by glycogen synthase kinase-3 (GSK3). Trends Immunol. 31, 24–31 (2010).
Wei, L., Laurence, A., Elias, K. M. & O’Shea, J. J. IL-21 is produced by TH17 cells and drives IL-17 production in a STAT3-dependent manner. J. Biol. Chem. 282, 34605–34610 (2007).
O’Shea, J. J. & Paul, W. E. Mechanisms underlying lineage commitment and plasticity of helper CD4+ T cells. Science 327, 1098–1102 (2010).
Kovalovsky, D. et al. β-catenin/Tcf determines the outcome of thymic selection in response to αβTCR signaling. J. Immunol. 183, 3873–3884 (2009).
Moser, A. R. et al. ApcMin: a mouse model for intestinal and mammary tumorigenesis. Eur. J. Cancer 31A, 1061–1064 (1995).
Chassaing, B., Aitken, J. D., Malleshappa, M. & Vijay-Kumar, M. Dextran sulfate sodium-induced colitis in mice. Curr. Protoc. Immunol. 104, 15.25.1–15.25.14 (2014).
Tanaka, T. et al. Dextran sodium sulfate strongly promotes colorectal carcinogenesis in ApcMin/+ mice: inflammatory stimuli by dextran sodium sulfate results in development of multiple colonic neoplasms. Int. J. Cancer 118, 25–34 (2006).
Brunkow, M. E. et al. Disruption of a new forkhead/winged-helix protein, scurfin, results in the fatal lymphoproliferative disorder of the scurfy mouse. Nat. Genet. 27, 68–73 (2001).
Franckaert, D. et al. Promiscuous Foxp3-cre activity reveals a differential requirement for CD28 in Foxp3+ and Foxp3− T cells. Immunol. Cell Biol. 93, 417–423 (2015).
Kitagawa, Y. et al. Guidance of regulatory T cell development by Satb1-dependent super-enhancer establishment. Nat. Immunol. 18, 173–183 (2017).
Subramanian, A. et al. Gene-set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Zitvogel, L., Ayyoub, M., Routy, B. & Kroemer, G. Microbiome and anticancer immunosurveillance. Cell 165, 276–287 (2016).
Voloshanenko, O. et al. Wnt secretion is required to maintain high levels of Wnt activity in colon cancer cells. Nat. Commun. 4, 2610 (2013).
Moparthi, L. & Koch, S. Wnt signaling in intestinal inflammation. Differentiation 108, 24–32 (2019).
Turner, J. R. Intestinal mucosal barrier function in health and disease. Nat. Rev. Immunol. 9, 799–809 (2009).
Konig, J. et al. Human intestinal barrier function in health and disease. Clin. Transl. Gastroenterol. 7, e196 (2016).
Pham, T. H., Okada, T., Matloubian, M., Lo, C. G. & Cyster, J. G. S1P1 receptor signaling overrides retention mediated by Gαi-coupled receptors to promote T cell egress. Immunity 28, 122–133 (2008).
Hoerl, A. E. & Kennard, R. W. Ridge regression: biased estimation for nonorthogonal problems. Technometrics 12, 55–67 (1970).
Therneau, T.M. A Package for Survival Analysis in S version 2.38. https://CRAN.R-project.org/ (2015).
Friedman, J. H., Hastie, T. & Tibshirani, R. Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. 33, 1–22 (2010).
Kassambara, A., Kosinski, M., Biecek, P. & Fabian, S. Package ‘survminer’. https://cran.r-project.org/web/packages/survminer/survminer.pdf 2017.
Harada, N. et al. Intestinal polyposis in mice with a dominant stable mutation of the β-catenin gene. EMBO J. 18, 5931–5942 (1999).
Rubtsov, Y. P. et al. Regulatory T cell-derived interleukin-10 limits inflammation at environmental interfaces. Immunity 28, 546–558 (2008).
Gounari, F. et al. Loss of adenomatous polyposis coli gene function disrupts thymic development. Nat. Immunol. 6, 800–809 (2005).
Atarashi, K. et al. TH17 cell induction by adhesion of microbes to intestinal epithelial cells. Cell 163, 367–380 (2015).
Dose, M. et al. β-Catenin induces T cell transformation by promoting genomic instability. Proc. Natl Acad. Sci. USA 111, 391–396 (2014).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Zhang, Y. et al. Model-based analysis of ChIP–seq (MACS). Genome Biol. 9, R137 (2008).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 7, 562–578 (2012).
Reich, M. et al. GenePattern 2.0. Nat. Genet. 38, 500–501 (2006).
Shen, L., Shao, N., Liu, X. & Nestler, E. ngs.plot: quick mining and visualization of next-generation sequencing data by integrating genomic databases. BMC Genomics 15, 284 (2014).
Freese, N. H., Norris, D. C. & Loraine, A. E. Integrated genome browser: visual analytics platform for genomics. Bioinformatics 32, 2089–2095 (2016).
Tripathi, S. et al. Meta- and orthogonal integration of influenza ‘OMICs’ data defines a role for UBR4 in virus budding. Cell Host Microbe 18, 723–735 (2015).
Acknowledgements
We thank clinical research coordinators B. Putz and A. Duong for facilitating the patient sample supply, S. Keerthivasan for introducing relevant mouse strains to the laboratory, and M. -L. Alegre for helpful advice on the manuscript. We also want to thank the Flow Cytometry Core (UCFlow), Human Tissue Resource Center and the Animal Resources Center at the University of Chicago. This work was supported by National Institutes of Health (NIH) grants R21AI076720 and R01AI147652, a Digestive Diseases Research Core Center pilot award (NIDDK P30DK42086) and an ASH bridge grant (to F.G.), and grants R01AI108682 (to F.G. and K.K.) and R01CA160436 (to K.K). Further support came from the Praespero autoimmunity fund (to F.G. and K.K.) and the Chicago Biomedical Consortium (to F.G.). A.O.E. was supported by an NIH minority supplement. P.S.M. was supported by T32 HL07605 Institutional NRSA and is currently an LLS fellow. J.Q. was supported by an AAI Careers in Immunology Fellowship. M.O. is a T32HD007009 award recipient.
Author information
Authors and Affiliations
Contributions
J.Q., F.G. and K.K. designed research and interpreted data. J.Q., S.A., L.H., J.W., A.M., M.O., P.S.M., A.O.E., A.O. and M.K. performed experiments and analyzed data. S.B.M., A.T.P., R.S., J.P. and C.R.W. analyzed data. J.Q., F.G. and K.K. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Immunology thanks Vincenzo Bronte and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available. L. A. Dempsey was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
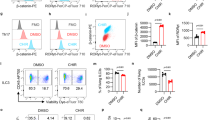
Extended Data Fig. 1 RORγt+ Treg cells producing pro-inflammatory cytokines expand in the PB/colonic mucosa of IBD patients.
Flow cytometric gating scheme to identify Treg cells in PB a. Live CD3+CD4+CD25+ Foxp3+ cells were assessed for CD127 expression and CD127−CD3+CD4+CD25+Foxp3+ Treg cells were gated for the analyses in b-f. b. Dotplots of RORγt versus CD4 expression in PB Treg cells of representative HD (HD21), and IBD (IMB29), and IBD/Dys (IMB17) patients c. Relative frequencies of RORγt+ cells within PB Treg cells. d. Representative β-catenin expression in RORγt+/RORγt− PB Treg cells. e. Cumulative β-catenin expression in RORγt+/RORγt− PB Treg cells (MFI). f. IL-17, IFN-γ, and TNF production in RORγt+ PB Treg cells before and after stimulation by PMA/ionomycin as indicated. b-f. two-sided unpaired t-test. Number of samples in c,e,f. HD(n = 16), IBD(n = 15), IBD_Dys(n = 17) g. Flow cytometric gating scheme assessing pro-inflammatory cytokine production by RORγt+ and RORγt− PB Treg populations (CD127−CD3+CD4+CD25+Foxp3+) after PMA/ionomycin stimulation. h. (left) Flow cytometric gating scheme for assessing pro-inflammatory cytokine production versus Helios expression in PB Treg cells, (right) Cumulative frequencies of Helios+ Treg cells in HD(n = 5), IBD(n = 5) and IBD/Dys(n = 4) patients, and IL-17, IFN-γ, and TNF expression in Helios+ and Helios− PB Treg cells of the same samples (two-sided unpaired t-test). i. Flow cytometric gating scheme for assessing IL-17 production by CD3+CD4+Foxp3+ RORγt+ and RORγT− colonic mucosa Treg cells.
Extended Data Fig. 2 The expression of Th17 and Treg cell signature genes in the TCGA CRC dataset indicate a prognostic relevance.
a–d. Analysis of the TCGA CRC cohort: a. Spearman correlation of average z-scores for the Th17_UP versus Treg_UP (blue, r = 0.7836, p < 0.001) signatures in CRC patients (n = 524). b. Ridge regression regularization and, c. cross validation of the deployed Cox proportional-hazards regression on the CRC clinical data extracted from TCGA. d. Histogram of Th17 gene-based score distribution through the CRC patient cohort (median = −0.033). If not stated differently data are represented as median + /− SEM and statistical testing is depicted as *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001, ****P ≤ 0.0001.
Extended Data Fig. 3 tSNE projection of flow cytometric data identifies unique populations within RORγt+ Treg cells in the APC/DSS IBD-associated CRC model.
Strategy for tSNE projection of a, splenic and b, colon-tissue resident Treg populations exemplarily shown for the activation marker flow-cytometric panel: Treg populations (gated on as live/dead stain negative, CD4+CD8−Foxp3+) from all samples of all treatment groups were concatenated into one fcs file and the tSNE analysis was performed on the concatenated file (settings used for FlowJo tSNE Plugin Tool: Iterations = 1000, Perplexity = 30, ETA/Learning Rate = 100). RORγt positive and negative fractions of Treg cells are indicated in red and green in the tSNE landscapes, respectively. Different RORγt+ Treg populations were identified within in the tSNE plot and histogram expression profiles of identified RORγt+ Treg populations are shown.
Extended Data Fig. 4 Treg cell specific up-regulation of β-catenin leads to a severe scurfy-like phenotype in mice.
a, Kaplan-Mayer survival curve of Foxp3Cre(0/+) wild-type (Cre) and Foxp3Cre(0/+) Ctnnb1fl(ex3) (CAT) male mice. b, Representative picture of the scurfy-like phenotype of a CAT mouse compared to a Cre mouse (26 d old). c, Representative picture of the pathologic enlargement of peripheral LNs and splenomegaly in a CAT mouse compared to a Cre litter mate control mouse. d, H&E-stainings of paraffin sections from organs of a representative 21 d old male CAT mouse and a Cre litter mate. Enlargement of secondary lymphoid organs (SPL – spleen & pLNs – peripheral lymph nodes) and reduction of the thymic cortex (THY – thymus, arrows) in CAT mice. CAT mice show severe immune infiltrates in lung and liver (middle panels, arrows), and eosinophil infiltration in the small intestine (SI) and colon (arrows). Size are provided in the images. At least 4 independent CAT and Cre litter mate mice were analysed. e, Frequency of CD3+, CD3+CD4+, and CD3+CD8+ T cell numbers in peripheral lymphoid organs of 3-4 weeks old Cre and CAT mice as determined by flow cytometric analysis. Cre_SPL(n = 7), Cre_mLN(n = 7), Cre_pLN(n = 7), CAT_SPL(n = 10), CAT_mLN(n = 8), p(CAT_mLN)=10, data are represented as median + /− SEM and statistical testing is depicted as two-sided unpaired t-tests with *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001, ****P ≤ 0.0001.
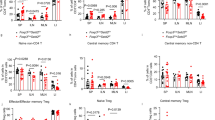
Extended Data Fig. 5 Treg cell specific stabilization of β-catenin results in severe systemic T cell activation and an altered Treg cell phenotype.
Activation status of a, CD3+CD4+ conventional T cells and, b, CD3+CD8+ CTLs was assessed via flow cytometric staining for CD25, CD44, CD69, CD62L, and Ki67 in peripheral lymphoid organs of 3-4 weeks old Foxp3Cre(0/+) wild-type (Cre) and Foxp3Cre(0/+) Ctnnb1fl(ex3) (CAT) male mice (CAT(n = 5), WT(n = 5)). c, Frequencies of total Treg cells within viable cells in peripheral lymphoid organs and the thymus (Cre_SPL(n = 5), Cre_mLN(n = 5), Cre_pLN(n = 5), Cre_THY(n = 5), CAT_SPL(n = 10), CAT_mLN(n = 10), CAT_pLN(n = 10), CAT_THY(n = 5)). d, Expression of Neuropilin in Cre and CAT Treg cells depicted as representative flow plots and cumulative column plots throughout peripheral lymphoid organs (Cre_SPL(n = 5), Cre_mLN(n = 5), Cre_pLN(n = 5), CAT_SPL(n = 4), CAT_mLN(n = 3), CAT_THY(n = 5)). Data are represented as median + /− SEM and statistical testing is depicted as two-sided unpaired t-tests with *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001, ****P ≤ 0.0001.
Extended Data Fig. 6 The chemokine receptor CCR9 is upregulated in β-cateninhi Treg cells.
a, Representative histograms for flow cytometric characterization of CCR9 expression in the natural chimera female heterozygote CD3+CD4+CD25+Foxp4+ Treg cells comparing YFP+ (Cre+) to YFP− (Cre−) populations in Foxp3Cre(+/−) Ctnnb1fl(ex3) (CAT) and Foxp3Cre(+/−) (Cre) chimeras in MLN and Spleen, as indicated. (FMO = fluorescence minus one negative staining control). b, Quantification of CCR9 MFI respresented as bar graphs in MLN (Cre(n = 5), CAT(n = 5)). Data are represented as median + /− SEM and statistical testing is depicted as one way Anova multiple comparisons with *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001, ****P ≤ 0.0001.
Extended Data Fig. 7 TCF-1 binds gene loci that are involved in T cell activation and survival in wild-type Treg cells.
a, Pie chart of the distribution of TCF-1 binding in genomic regions (P = promoter, PE = poised enhancer, AE = active enhancer) identified by K-means-clustering. Peak numbers (p), corresponding gene numbers (g), and relative percentages (%) for each genomic region. b, Heat maps centered on TCF-1 binding in indicated genomic regions (± 1.5 kb) and enrichment of histone marks (H3K4me1, H3K4me3, H3K27me3, H3K27Ac), chromatin accessibility (ATAC), and DNA methylation (MBD). c, Enrichment histograms of TCF-1 binding, histone marks, chromatin accessibility, and DNA methylation marks at TCF-1 bound sites (± 1.5 kb) in the indicated genomic regions. d, De novo transcription-factor-binding motif analysis (HOMER) of TCF-1-bound sites for the indicated genomic regions. Most, significantly enriched motifs and corresponding p values are listed. e, Functional pathways enriched for TCF-1-bound genes in the indicated genomic regions. Pathways and statistical enrichment were determined using Metascape (http://www.metascape.org).
Extended Data Fig. 8 Foxp3 preferentially binds accessible chromatin and its consensus motif within different regulatory elements of genes in Treg cells.
a, Pie chart of the distribution of Foxp3 binding in genomic regions (P = promoter, PE = poised enhancer, AE = active enhancer) identified by K-means-clustering. Peak numbers (p), corresponding gene numbers (g), and relative percentages (%) for each genomic region. b, Heat maps centered on Foxp3 binding in indicated genomic regions (± 1.5 kb) and enrichment of histone marks (H3K4me1, H3K4me3, H3K27me3, H3K27Ac), chromatin accessibility (ATAC), and DNA methylation (MBD). c, Enrichment histograms of Foxp3 binding, histone marks, chromatin accessibility, and DNA methylation marks at Foxp3 bound sites (± 1.5 kb) in the indicated genomic regions. d, De novo transcription-factor-binding motif analysis (HOMER) of TCF-1-bound sites for the indicated genomic regions. Most, significantly enriched motifs and corresponding p values are listed. e, Functional pathways enriched for TCF-1-bound genes in the indicated genomic regions. Pathways and statistical enrichment were determined using Metascape (http://www.metascape.org).
Extended Data Fig. 9 Wnt/β-catenin activation does not significantly alter the Treg_UP signature and enhances the expression of leukocyte migration signature genes in Treg cells.
RNA expression heat maps (n = 3 biological replicates for each genotype, FPKMs) showing genes of GSEA analysis for the Treg_UP, TH17_UP, and leukocyte migration signature displayed in Fig. 8d of CAT versus Cre Treg cells.
Supplementary information
Rights and permissions
About this article
Cite this article
Quandt, J., Arnovitz, S., Haghi, L. et al. Wnt–β-catenin activation epigenetically reprograms Treg cells in inflammatory bowel disease and dysplastic progression. Nat Immunol 22, 471–484 (2021). https://doi.org/10.1038/s41590-021-00889-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-021-00889-2
This article is cited by
-
The dynamic shifts of IL-10-producing Th17 and IL-17-producing Treg in health and disease: a crosstalk between ancient "Yin-Yang" theory and modern immunology
Cell Communication and Signaling (2024)
-
TRPM8 inhibits substance P release from primary sensory neurons via PKA/GSK-3beta to protect colonic epithelium in colitis
Cell Death & Disease (2024)
-
Dysregulation of Wnt/β-catenin signaling contributes to intestinal inflammation through regulation of group 3 innate lymphoid cells
Nature Communications (2024)
-
Unravelling the Road to Recovery: Mechanisms of Wnt Signalling in Spinal Cord Injury
Molecular Neurobiology (2024)
-
Regulatory T cells in the face of the intestinal microbiota
Nature Reviews Immunology (2023)