Abstract
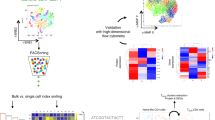
T cell memory relies on the generation of antigen-specific progenitors with stem-like properties. However, the identity of these progenitors has remained unclear, precluding a full understanding of the differentiation trajectories that underpin the heterogeneity of antigen-experienced T cells. We used a systematic approach guided by single-cell RNA-sequencing data to map the organizational structure of the human CD8+ memory T cell pool under physiological conditions. We identified two previously unrecognized subsets of clonally, epigenetically, functionally, phenotypically and transcriptionally distinct stem-like CD8+ memory T cells. Progenitors lacking the inhibitory receptors programmed death-1 (PD-1) and T cell immunoreceptor with Ig and ITIM domains (TIGIT) were committed to a functional lineage, whereas progenitors expressing PD-1 and TIGIT were committed to a dysfunctional, exhausted-like lineage. Collectively, these data reveal the existence of parallel differentiation programs in the human CD8+ memory T cell pool, with potentially broad implications for the development of immunotherapies and vaccines.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Publicly available scRNA-seq data were retrieved from the Gene Expression Omnibus via accession code GSE120575. Microarray data from YFV-17D-specific CD8+ T cells were retrieved from the Gene Expression Omnibus via accession code GSE26347. Gene sets of interest were retrieved from the Molecular Signatures Database (http://www.broadinstitute.org/gsea/msigdb/index.jsp). The ATAC-seq data reported in this paper are available on request. The bulk RNA-seq and scRNA-seq data reported in this paper have been deposited in the Gene Expression Omnibus under accession code GSE147398. The TCR-seq data reported in this paper have been deposited at the European Bioinformatic Institute under accession code E-MTAB-8892. All other data that support the findings of this study are available on request from the corresponding author.
Code availability
Scripts used to analyze the ATAC-seq data are available at https://github.com/luglilab/SP018_CD8_Galletti_et_al. All other codes are available on request.
References
Kaech, S. M., Wherry, E. J. & Ahmed, R. Effector and memory T-cell differentiation: implications for vaccine development. Nat. Rev. Immunol. 2, 251–262 (2002).
Gattinoni, L., Speiser, D. E., Lichterfeld, M. & Bonini, C. T memory stem cells in health and disease. Nat. Med. 23, 18–27 (2017).
Lugli, E., Galletti, G., Boi, S. K. & Youngblood, B. A. Stem, effector, and hybrid states of memory CD8+ T cells. Trends Immunol. 41, 17–28 (2020).
Gattinoni, L. et al. A human memory T cell subset with stem cell-like properties. Nat. Med. 17, 1290–1297 (2011).
Biasco, L. et al. In vivo tracking of T cells in humans unveils decade-long survival and activity of genetically modified T memory stem cells. Sci. Transl. Med. 7, 273ra13 (2015).
Mahnke, Y. D., Brodie, T. M., Sallusto, F., Roederer, M. & Lugli, E. The who’s who of T-cell differentiation: human memory T-cell subsets. Eur. J. Immunol. 43, 2797–2809 (2013).
Graef, P. et al. Serial transfer of single-cell-derived immunocompetence reveals stemness of CD8+ central memory T cells. Immunity 41, 116–126 (2014).
Wherry, E. J. T cell exhaustion. Nat. Immunol. 12, 492–499 (2011).
Angelosanto, J. M., Blackburn, S. D., Crawford, A. & Wherry, E. J. Progressive loss of memory T cell potential and commitment to exhaustion during chronic viral infection. J. Virol. 86, 8161–8170 (2012).
Schietinger, A. et al. Tumor-specific T cell dysfunction is a dynamic antigen-driven differentiation program initiated early during tumorigenesis. Immunity 45, 389–401 (2016).
Philip, M. et al. Chromatin states define tumour-specific T cell dysfunction and reprogramming. Nature 545, 452–456 (2017).
Sen, D. R. et al. The epigenetic landscape of T cell exhaustion. Science 354, 1165–1169 (2016).
Im, S. J. et al. Defining CD8+ T cells that provide the proliferative burst after PD-1 therapy. Nature 537, 417–421 (2016).
Leong, Y. A. et al. CXCR5+ follicular cytotoxic T cells control viral infection in B cell follicles. Nat. Immunol. 17, 1187–1196 (2016).
Utzschneider, D. T. et al. T cell factor 1-expressing memory-like CD8+ T cells sustain the immune response to chronic viral infections. Immunity 45, 415–427 (2016).
Brummelman, J. et al. High-dimensional single cell analysis identifies stem-like cytotoxic CD8+ T cells infiltrating human tumors. J. Exp. Med. 215, 2520–2535 (2018).
He, R. et al. Follicular CXCR5-expressing CD8+ T cells curtail chronic viral infection. Nature 537, 412–428 (2016).
Sade-Feldman, M. et al. Defining T cell states associated with response to checkpoint immunotherapy in melanoma. Cell 175, 998–1013.e20 (2018).
Siddiqui, I. et al. Intratumoral Tcf1+PD-1+CD8+ T cells with stem-like properties promote tumor control in response to vaccination and checkpoint blockade immunotherapy. Immunity 50, 195–211.e10 (2019).
Miller, B. C. et al. Subsets of exhausted CD8+ T cells differentially mediate tumor control and respond to checkpoint blockade. Nat. Immunol. 20, 326–336 (2019).
Becht, E. et al. Dimensionality reduction for visualizing single-cell data using UMAP. Nat. Biotechnol. 37, 38–44 (2018).
Dusseaux, M. et al. Human MAIT cells are xenobiotic-resistant, tissue-targeted, CD161hi IL-17-secreting T cells. Blood 117, 1250–1259 (2011).
Lugli, E. et al. Superior T memory stem cell persistence supports long-lived T cell memory. J. Clin. Invest. 123, 594–599 (2013).
Oberdoerffer, S. et al. Regulation of CD45 alternative splicing by heterogeneous ribonucleoprotein, hnRNPLL. Science 321, 686–691 (2008).
Lugli, E. et al. Identification, isolation and in vitro expansion of human and nonhuman primate T stem cell memory cells. Nat. Protoc. 8, 33–42 (2013).
Stephen, T. L. et al. SATB1 expression governs epigenetic repression of PD-1 in tumor-reactive T cells. Immunity 46, 51–64 (2017).
Yao, C. et al. Single-cell RNA-seq reveals TOX as a key regulator of CD8+ T cell persistence in chronic infection. Nat. Immunol. 20, 890–901 (2019).
Alfei, F. et al. TOX reinforces the phenotype and longevity of exhausted T cells in chronic viral infection. Nature 571, 265–269 (2019).
Scott, A. C. et al. TOX is a critical regulator of tumour-specific T cell differentiation. Nature 571, 270–274 (2019).
Khan, O. et al. TOX transcriptionally and epigenetically programs CD8+ T cell exhaustion. Nature 571, 211–218 (2019).
Wang, X. et al. TOX promotes the exhaustion of antitumor CD8+ T cells by preventing PD1 degradation in hepatocellular carcinoma. J. Hepatol. 71, 731–741 (2019).
Seo, H. et al. TOX and TOX2 transcription factors cooperate with NR4A transcription factors to impose CD8+ T cell exhaustion. Proc. Natl Acad. Sci. USA 116, 12410–12415 (2019).
Li, J., He, Y., Hao, J., Ni, L. & Dong, C. High levels of Eomes promote exhaustion of anti-tumor CD8+ T cells. Front. Immunol. 9, 2981 (2018).
Giordano, M. et al. Molecular profiling of CD8 T cells in autochthonous melanoma identifies Maf as driver of exhaustion. EMBO J. 34, 2042–2058 (2015).
Gattinoni, L. et al. Wnt signaling arrests effector T cell differentiation and generates CD8+ memory stem cells. Nat. Med. 15, 808–813 (2009).
Kondo, T. et al. Notch-mediated conversion of activated T cells into stem cell memory-like T cells for adoptive immunotherapy. Nat. Commun. 8, 15338 (2017).
Widjaja, C. E. et al. Proteasome activity regulates CD8+ T lymphocyte metabolism and fate specification. J. Clin. Invest. 127, 3609–3623 (2017).
Yang, Z. Z. et al. TGF-β upregulates CD70 expression and induces exhaustion of effector memory T cells in B-cell non-Hodgkin’s lymphoma. Leukemia 28, 1872–1884 (2014).
Vodnala, S. K. et al. T cell stemness and dysfunction in tumors are triggered by a common mechanism. Science 363, eaau0135 (2019).
Henning, A. N., Roychoudhuri, R. & Restifo, N. P. Epigenetic control of CD8+ T cell differentiation. Nat. Rev. Immunol. 18, 340–356 (2018).
Doering, T. A. et al. Network analysis reveals centrally connected genes and pathways involved in CD8+ T cell exhaustion versus memory. Immunity 37, 1130–1144 (2012).
Thommen, D. S. et al. A transcriptionally and functionally distinct PD-1+ CD8+ T cell pool with predictive potential in non-small-cell lung cancer treated with PD-1 blockade. Nat. Med. 24, 994–1004 (2018).
Utzschneider, D. T. et al. High antigen levels induce an exhausted phenotype in a chronic infection without impairing T cell expansion and survival. J. Exp. Med. 213, 1819–1834 (2016).
Fuertes Marraco, S. A. et al. Long-lasting stem cell–like memory CD8+ T cells with a naive-like profile upon yellow fever vaccination. Sci. Transl. Med. 7, 282ra48 (2015).
Akondy, R. S. et al. Origin and differentiation of human memory CD8 T cells after vaccination. Nature 552, 362–367 (2017).
Price, D. A. et al. Avidity for antigen shapes clonal dominance in CD8+ T cell populations specific for persistent DNA viruses. J. Exp. Med. 202, 1349–1361 (2005).
Blank, C. U. et al. Defining ‘T cell exhaustion’. Nat. Rev. Immunol. 19, 665–674 (2019).
Pauken, K. E. et al. Epigenetic stability of exhausted T cells limits durability of reinvigoration by PD-1 blockade. Science 354, 1160–1165 (2016).
Ghoneim, H. E. et al. De novo epigenetic programs inhibit PD-1 blockade-mediated T cell rejuvenation. Cell 170, 142–157.e19 (2017).
West, E. E. et al. Tight regulation of memory CD8+ T cells limits their effectiveness during sustained high viral load. Immunity 35, 285–298 (2011).
Lugli, E. et al. IL-15 delays suppression and fails to promote immune reconstitution in virally suppressed chronically SIV-infected macaques. Blood 118, 2520–2529 (2011).
Roberto, A. et al. Role of naive-derived T memory stem cells in T-cell reconstitution following allogeneic transplantation. Blood 125, 2855–2864 (2015).
Falcone, L. & Casucci, M. Exploiting secreted luciferases to monitor tumor progression in vivo. Methods Mol. Biol. 1393, 105–111 (2016).
Brummelman, J. et al. Development, application and computational analysis of high-dimensional fluorescent antibody panels for single-cell flow cytometry. Nat. Protoc. 14, 1946–1969 (2019).
Newell, E. W. et al. Combinatorial tetramer staining and mass cytometry analysis facilitate T-cell epitope mapping and characterization. Nat. Biotechnol. 31, 623–629 (2013).
Bakker, A. H. et al. Conditional MHC class I ligands and peptide exchange technology for the human MHC gene products HLA-A1, -A3, -A11, and -B7. Proc. Natl Acad. Sci. USA 105, 3825–3830 (2008).
Simoni, Y. et al. Bystander CD8+ T cells are abundant and phenotypically distinct in human tumour infiltrates. Nature 557, 575–579 (2018).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Leek, J. T., Johnson, W. E., Parker, H. S., Jaffe, A. E. & Storey, J. D. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 28, 882–883 (2012).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 17, 10–12 (2011).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Capper, R. et al. The nature of telomere fusion and a definition of the critical telomere length in human cells. Genes Dev. 21, 2495–2508 (2007).
Shugay, M. et al. Towards error-free profiling of immune repertoires. Nat. Methods 11, 653–655 (2014).
Bolotin, D. A. et al. MiXCR: software for comprehensive adaptive immunity profiling. Nat. Methods 12, 380–381 (2015).
Shugay, M. et al. VDJtools: unifying post-analysis of T cell receptor repertoires. PLoS Comput. Biol. 11, e1004503 (2015).
Acknowledgements
The authors thank G. Natoli (European Institute of Oncology, Milan) for assistance with the ATAC-seq protocol, R. Roychoudhuri (University of Cambridge) and M. Iannacone (San Raffaele Scientific Institute, Milan) for critical discussions, and G. Cugini and G. Colombo (Humanitas Clinical and Research Center, Milan) for the provision of lymph node samples. This work was funded by the European Research Council (ERC-2014-STG PERSYST no. 640511 to E.L.) and by the Associazione Italiana per la Ricerca sul Cancro (AIRC IG 20676 to E.L.). Additional support was provided by the Associazione Italiana per la Ricerca sul Cancro (AIRC IG 21567 to D.M.), the Italian Ministry of Health (Bando Ricerca Finalizzata PE-2016-02363915 to D.M.), the Intramural Research Fund of the Humanitas Clinical and Research Center (5 ×1000 2019 Program to D.M.) and Cancer Research UK (C17199/A18246/A29202 to D.M.B.). G.G., G.D.S., S.P. and E.S. were supported by Fellowships from the Fondazione Italiana per la Ricerca sul Cancro-Associazione Italiana per la Ricerca sul Cancro (FIRC-AIRC). A.N.D. and M.M. were supported by the Ministry of Education, Youth, and Sports of the Czech Republic (CEITEC 2020 LQ1601). D.A.P. was supported by a Wellcome Trust Senior Investigator Award (100326/Z/12/Z). D.M.C. was supported by the Ministry of Health of the Russian Federation (0908300057056). The purchase of a FACSSymphony A5 was defrayed in part by a grant from the Italian Ministry of Health (Agreement 82/2015).
Author information
Authors and Affiliations
Contributions
G.G. and E.L. conceived the study; G.G., G.D.S., E.M.C.M., S.P., C.M., T.M.B., A.N.D., M.M., E.S., G.A., F.D.P., V.Z., A.S., B.C., F.S.C., A.A., C.P., S.P., L.G., R.E.J., D.M.B., E.G., S.L.-L. and K.L. performed experiments; G.G., G.D.S., E.M.C.M., S.P., T.M.B., A.N.D., D.M.B., D.M.C., E.W.N., M.C. and E.L. analyzed data; D.M., S.K.B., B.A.Y. and D.A.P. provided critical expertise and resources; E.L. supervised the study; G.G., D.A.P. and E.L. wrote the manuscript. All authors contributed intellectually and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The Laboratory of Translational Immunology receives reagents in kind as part of a collaborative research agreement with BD Biosciences (Italy). L.G. and E.L. are inventors on a patent describing methods for the generation and isolation of TSCM cells. E.L. has a consulting agreement with Achilles Therapeutics. L.G. has consulting agreements with Lyell Immunopharma and Advaxis Immunotherapies. E.W.N. is a cofounder and advisor for ImmunoScape. The other authors have no competing interests.
Additional information
Peer review information Peer reviewer reports are available. Zoltan Fehervari was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–5.
Supplementary Table 1
Samples used in this study.
Supplementary Table 2
Differentially expressed genes identified by scRNA-seq.
Supplementary Table 3
Differentially expressed genes between clusters C2 and C6 from scRNA-seq.
Supplementary Table 4
Differentially expressed genes from bulk RNA-seq plus enrichment analysis with publicly available data.
Supplementary Table 5
Differentially expressed genes between activated TSTEM and TPEX cells.
Supplementary Table 6
Flow cytometry reagents.
Supplementary Table 7
Mass cytometry reagents.
Rights and permissions
About this article
Cite this article
Galletti, G., De Simone, G., Mazza, E.M.C. et al. Two subsets of stem-like CD8+ memory T cell progenitors with distinct fate commitments in humans. Nat Immunol 21, 1552–1562 (2020). https://doi.org/10.1038/s41590-020-0791-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-020-0791-5
This article is cited by
-
Advancements in understanding chicken coccidiosis: from Eimeria biology to innovative control strategies
One Health Advances (2024)
-
Cystine deprivation triggers CD36-mediated ferroptosis and dysfunction of tumor infiltrating CD8+ T cells
Cell Death & Disease (2024)
-
The effect of metformin on senescence of T lymphocytes
Immunity & Ageing (2023)
-
GITR and TIGIT immunotherapy provokes divergent multicellular responses in the tumor microenvironment of gastrointestinal cancers
Genome Medicine (2023)
-
CD28/PD1 co-expression: dual impact on CD8+ T cells in peripheral blood and tumor tissue, and its significance in NSCLC patients' survival and ICB response
Journal of Experimental & Clinical Cancer Research (2023)