Abstract
Adoptive cell therapies using genetically engineered T cell receptor or chimeric antigen receptor T cells are emerging forms of immunotherapy that redirect T cells to specifically target cancer. However, tumor antigen heterogeneity remains a key challenge limiting their efficacy against solid cancers. Here, we engineered T cells to secrete the dendritic cell (DC) growth factor Fms-like tyrosine kinase 3 ligand (Flt3L). Flt3L-secreting T cells expanded intratumoral conventional type 1 DCs and substantially increased host DC and T cell activation when combined with immune agonists poly (I:C) and anti-4-1BB. Importantly, combination therapy led to enhanced inhibition of tumor growth and the induction of epitope spreading towards antigens beyond those recognized by adoptively transferred T cells in solid tumor models of T cell receptor and chimeric antigen receptor T cell therapy. Our data suggest that augmenting endogenous DCs is a promising strategy to overcome the clinical problem of antigen-negative tumor escape following adoptive cell therapy.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The data generated or analyzed to support the findings of this study are available in this article and from the corresponding author upon request without restrictions. Source data for Figs. 1–7 and Extended Data Figs. 1–9 are presented with the paper.
References
Mardiana, S., Solomon, B. J., Darcy, P. K. & Beavis, P. A. Supercharging adoptive T cell therapy to overcome solid tumor-induced immunosuppression. Sci. Transl. Med. 11, eaaw2293 (2019).
Mardiana, S., Lai, J., House, I. G., Beavis, P. A. & Darcy, P. K. Switching on the green light for chimeric antigen receptor T-cell therapy. Clin. Transl. Immunology 8, e1046 (2019).
June, C. H., O’Connor, R. S., Kawalekar, O. U., Ghassemi, S. & Milone, M. C. CAR T cell immunotherapy for human cancer. Science 359, 1361–1365 (2018).
Majzner, R. G. & Mackall, C. L. Tumor antigen escape from CAR T-cell therapy. Cancer Discov. 8, 1219–1226 (2018).
Jackson, H. J. & Brentjens, R. J. Overcoming antigen escape with CAR T-cell therapy. Cancer Discov. 5, 1238–1240 (2015).
Salmon, H. et al. Expansion and activation of CD103+ dendritic cell progenitors at the tumor site enhances tumor responses to therapeutic PD-L1 and BRAF inhibition. Immunity 44, 924–938 (2016).
Roberts, E. W. et al. Critical role for CD103+/CD141+ dendritic cells bearing CCR7 for tumor antigen trafficking and priming of T cell immunity in melanoma. Cancer Cell 30, 324–336 (2016).
Broz, M. L. et al. Dissecting the tumor myeloid compartment reveals rare activating antigen-presenting cells critical for T cell immunity. Cancer Cell 26, 638–652 (2014).
Xu, Y. et al. Closely related T-memory stem cells correlate with in vivo expansion of CAR.CD19-T cells and are preserved by IL-7 and IL-15. Blood 123, 3750–3759 (2014).
Alizadeh, D. et al. IL15 enhances CAR-T cell antitumor activity by reducing mTORC1 activity and preserving their stem cell memory phenotype. Cancer Immunol. Res. 7, 759–772 (2019).
Gattinoni, L., Klebanoff, C. A. & Restifo, N. P. Paths to stemness: building the ultimate antitumour T cell. Nat. Rev. Cancer 12, 671–684 (2012).
Brentjens, R. J. et al. Safety and persistence of adoptively transferred autologous CD19-targeted T cells in patients with relapsed or chemotherapy refractory B-cell leukemias. Blood 118, 4817–4828 (2011).
Wang, L. X. et al. Tumor ablation by gene-modified T cells in the absence of autoimmunity. Cancer Res. 70, 9591–9598 (2010).
Schlitzer, A. et al. Identification of cDC1- and cDC2-committed DC progenitors reveals early lineage priming at the common DC progenitor stage in the bone marrow. Nat. Immunol. 16, 718–728 (2015).
Grajales-Reyes, G. E. et al. Batf3 maintains autoactivation of Irf8 for commitment of a CD8ɑ+ conventional DC clonogenic progenitor. Nat. Immunol. 16, 708–717 (2015).
Sichien, D. et al. IRF8 transcription factor controls survival and function of terminally differentiated conventional and plasmacytoid dendritic cells, respectively. Immunity 45, 626–640 (2016).
Guilliams, M. et al. Unsupervised high-dimensional analysis aligns dendritic cells across tissues and species. Immunity 45, 669–684 (2016).
Durai, V. et al. Cryptic activation of an Irf8 enhancer governs cDC1 fate specification. Nat. Immunol. 20, 1161–1173 (2019).
Binnewies, M. et al. Unleashing type-2 dendritic cells to drive protective antitumor CD4+ T cell immunity. Cell 177, 556–571.e16 (2019).
Bottcher, J. P. et al. NK cells stimulate recruitment of cDC1 into the tumor microenvironment promoting cancer immune control. Cell 172, 1022–1037.e14 (2018).
Ruffell, B. et al. Macrophage IL-10 blocks CD8+ T cell-dependent responses to chemotherapy by suppressing IL-12 expression in intratumoral dendritic cells. Cancer Cell 26, 623–637 (2014).
Marigo, I. et al. T cell cancer therapy requires CD40-CD40L activation of tumor necrosis factor and inducible nitric-oxide-synthase-producing dendritic cells. Cancer Cell 30, 651 (2016).
Sade-Feldman, M. et al. Defining T cell states associated with response to checkpoint immunotherapy in melanoma. Cell 175, 998–1013.e20 (2018).
Siddiqui, I. et al. Intratumoral Tcf1+PD-1+CD8+ T cells with stem-like properties promote tumor control in response to vaccination and checkpoint blockade immunotherapy. Immunity 50, 195–211.e10 (2019).
Fuertes, M. B. et al. Host type I IFN signals are required for antitumor CD8+ T cell responses through CD8ɑ+ dendritic cells. J. Exp. Med. 208, 2005–2016 (2011).
Hammerich, L. et al. Systemic clinical tumor regressions and potentiation of PD1 blockade with in situ vaccination. Nat. Med. 25, 814–824 (2019).
Segal, N. H. et al. Phase I study of single-agent utomilumab (PF-05082566), a 4-1BB/CD137 agonist, in patients with advanced cancer. Clin. Cancer Res. 24, 1816–1823 (2018).
Mardiana, S. et al. A multifunctional role for adjuvant anti-4-1BB therapy in augmenting antitumor response by chimeric antigen receptor T cells. Cancer Res. 77, 1296–1309 (2017).
Sanchez-Paulete, A. R. et al. Cancer immunotherapy with immunomodulatory anti-CD137 and anti-PD-1 monoclonal antibodies requires BATF3-dependent dendritic cells. Cancer Discov. 6, 71–79 (2016).
Canale, F. P. et al. CD39 expression defines cell exhaustion in tumor-infiltrating CD8+ T cells—response. Cancer Res. 78, 5175 (2018).
John, L. B. et al. Anti-PD-1 antibody therapy potently enhances the eradication of established tumors by gene-modified T cells. Clin. Cancer Res. 19, 5636–5646 (2013).
Beavis, P. A. et al. Targeting the adenosine 2A receptor enhances chimeric antigen receptor T cell efficacy. J. Clin. Invest. 127, 929–941 (2017).
Michie, J. et al. Antagonism of IAPs enhances CAR T-cell efficacy. Cancer Immunol. Res. 7, 183–192 (2019).
Rafiq, S. et al. Targeted delivery of a PD-1-blocking scFv by CAR-T cells enhances anti-tumor efficacy in vivo. Nat. Biotechnol. 36, 847–856 (2018).
Koneru, M., Purdon, T. J., Spriggs, D., Koneru, S. & Brentjens, R. J. IL-12 secreting tumor-targeted chimeric antigen receptor T cells eradicate ovarian tumors. Oncoimmunology 4, e994446 (2015).
Newick, K. et al. Augmentation of CAR T-cell trafficking and antitumor efficacy by blocking protein kinase A localization. Cancer Immunol. Res. 4, 541–551 (2016).
Chang, Z. L. et al. Rewiring T-cell responses to soluble factors with chimeric antigen receptors. Nat. Chem. Biol. 14, 317–324 (2018).
Moon, E. K. et al. Expression of a functional CCR2 receptor enhances tumor localization and tumor eradication by retargeted human T cells expressing a mesothelin-specific chimeric antibody receptor. Clin. Cancer Res. 17, 4719–4730 (2011).
Chmielewski, M., Kopecky, C., Hombach, A. A. & Abken, H. IL-12 release by engineered T cells expressing chimeric antigen receptors can effectively muster an antigen-independent macrophage response on tumor cells that have shut down tumor antigen expression. Cancer Res. 71, 5697–5706 (2011).
Avanzi, M. P. et al. Engineered tumor-targeted T cells mediate enhanced anti-tumor efficacy both directly and through activation of the endogenous immune system. Cell Rep. 23, 2130–2141 (2018).
Chmielewski, M. & Abken, H. CAR T cells releasing IL-18 convert to T-Bethigh FoxO1low effectors that exhibit augmented activity against advanced solid tumors. Cell Rep. 21, 3205–3219 (2017).
Kuhn, N. F. et al. CD40 ligand-modified chimeric antigen receptor T cells enhance antitumor function by eliciting an endogenous antitumor response. Cancer Cell 35, 473–488.e6 (2019).
Adachi, K. et al. IL-7 and CCL19 expression in CAR-T cells improves immune cell infiltration and CAR-T cell survival in the tumor. Nat. Biotechnol. 36, 346–351 (2018).
de Mingo Pulido, A. et al. TIM-3 regulates CD103+ dendritic cell function and response to chemotherapy in breast cancer. Cancer Cell 33, 60–74.e6 (2018).
Beavis, P. A. et al. Dual PD-1 and CTLA-4 checkpoint blockade promotes antitumor immune responses through CD4+Foxp3− cell-mediated modulation of CD103+ dendritic cells. Cancer Immunol. Res. 6, 1069–1081 (2018).
Spranger, S., Dai, D., Horton, B. & Gajewski, T. F. Tumor-residing Batf3 dendritic cells are required for effector T cell trafficking and adoptive T cell therapy. Cancer Cell 31, 711–723.e4 (2017).
Durai, V. et al. Altered compensatory cytokine signaling underlies the discrepancy between Flt3 −/− and Flt3l −/− mice. J. Exp. Med. 215, 1417–1435 (2018).
Brown, C. E. et al. Regression of glioblastoma after chimeric antigen receptor T-cell therapy. N. Engl. J. Med. 375, 2561–2569 (2016).
Adusumilli, P. S. et al. Regional delivery of mesothelin-targeted CAR T cell therapy generates potent and long-lasting CD4-dependent tumor immunity. Sci. Transl. Med. 6, 261ra151 (2014).
Tchou, J. et al. Safety and efficacy of intratumoral injections of chimeric antigen receptor (CAR) T cells in metastatic breast cancer. Cancer Immunol. Res. 5, 1152–1161 (2017).
Piechocki, M. P., Ho, Y. S., Pilon, S. & Wei, W. Z. Human ErbB-2 (Her-2) transgenic mice: a model system for testing Her-2 based vaccines. J. Immunol. 171, 5787–5794 (2003).
Darcy, P. K. et al. Redirected perforin-dependent lysis of colon carcinoma by ex vivo genetically engineered CTL. J. Immunol. 164, 3705–3712 (2000).
Haynes, N. M. et al. Redirecting mouse CTL against colon carcinoma: superior signaling efficacy of single-chain variable domain chimeras containing TCR-ζ vs Fc ε RI-γ. J. Immunol. 166, 182–187 (2001).
Haynes, N. et al. Single-chain antigen recognition receptors that costimulate potent rejection of established experimental tumors. Blood 100, 3155–3163 (2002).
Duong, C. P. et al. Engineering T cell function using chimeric antigen receptors identified using a DNA library approach. PLoS ONE 8, e63037 (2013).
Naik, S. H. et al. Development of plasmacytoid and conventional dendritic cell subtypes from single precursor cells derived in vitro and in vivo. Nat. Immunol. 8, 1217–1226 (2007).
Mayer, C. T. et al. Selective and efficient generation of functional Batf3-dependent CD103+ dendritic cells from mouse bone marrow. Blood 124, 3081–3091 (2014).
Helft, J. et al. GM-CSF mouse bone marrow cultures comprise a heterogeneous population of CD11c+MHCII+ macrophages and dendritic cells. Immunity 42, 1197–1211 (2015).
Chen, E. Y. et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinformatics 14, 128 (2013).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Betel, I. The occurrence of cytochromes in nuclei isolated from rat thymus. Biochim. Biophys. Acta 143, 62–69 (1967).
Zotos, D. et al. IL-21 regulates germinal center B cell differentiation and proliferation through a B cell–intrinsic mechanism. J. Exp. Med. 207, 365–378 (2010).
Acknowledgements
We acknowledge assistance from the staff of the Animal Facility, the Centre for Advanced Histology and Microscopy, the Flow Cytometry Facility, the Molecular Genomics Core and the Pathology departments at the Peter MacCallum Cancer Centre. We also acknowledge the contribution of consumer advocates K. Gill, M. Rear and G. Sissing for this study. This work was funded by the National Health and Medical Research Council (NHMRC; grant nos. 1062580, 1143976 and 1150425), the Cancer Council Victoria (grant no. 1156382), the National Breast Cancer Foundation (grant no. IIRS-19-016 19-22), the CLEARbridge Foundation and the Tour de Cure Foundation. P.A.B. is supported by a National Breast Cancer Foundation Fellowship (ID no. ECF-17-005). P.K.D., A.M.L. and K.K. are supported by NHMRC Research Fellowships (grant nos. APP1041828, APP1080321 and APP1102792, respectively).
Author information
Authors and Affiliations
Contributions
J.L., S.M., P.A.B. and P.K.D. designed the experiments, developed the methodology, analyzed and interpreted data and wrote the manuscript. J.L., S.M., P.A.B., P.K.D., I.G.H., K.S., M.A.H., L.G., A.X.Y.C., K.L.T., E.V.P. and J.D.C. performed experiments and acquired data. E.M.C., A.M.L., B.J.S., J.A.T., K.K., M.E., S.J.V. and J.W. provided technical support and advice on data analysis and interpretation. P.A.B. and P.K.D. supervised the study and were responsible for coordination and strategy.
Corresponding authors
Ethics declarations
Competing interests
P.A.B. and P.K.D. are inventors on a patent filed for the following study (filing no. PCT/AU2019/050660).
Additional information
Peer review information Zoltan Fehervari was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Tumor-derived Flt3L mediates tumor regression. Related to Fig. 1.
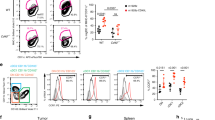
a, Flt3L levels from 24JK-Her2 and MC38 cell lines transduced to express Cherry control (Cherry) or Flt3L Cherry (Cherry FL). Data represent technical replicates from one of two independent experiments with similar results. b, Tumor and serum Flt3L levels from hHer2 mice 6 days post inoculation with 24JK-Her2 Cherry or Cherry FL cells (3 mice/group), from one of two independent experiments. c, Tumor measurement (2-way ANOVA) and survival (tumor area > 100mm2) of C57BL/6 mice subcutaneously injected with 5 × 104 MC38 expressing Flt3L cherry (Cherry FL, n = 7) or cherry control (Cherry, n = 8) (two-tailed Mantel-Cox, two pooled independent experiments). d, Tumor myeloid cell gating strategy. Lineage markers include Thy1.2, CD19, B220/CD45R, NK1.1 and Ly6G. e, Frequency of tumor infiltrating CD4+ T cells, Treg, CD11b+ cDC2, macrophages and neutrophils 9 days post tumor inoculation with 24JK-Her2 Cherry FL or control. Bars represent mean ± SEM, b, e Unpaired Student’s t-test, symbols represent individual mice.
Extended Data Fig. 2 Characterization of in vitro differentiated BM DCs. Related to Fig. 2.
a, In vitro differentiated BM DC gating strategy. b, Contour plots of BM cells differentiated with (FL-DCs) or without (GM-DCs) exogenous Flt3L (20 ng/mL). c, IFN-γ and TNF levels in supernatants of 4 × 104 naive OT-I T cells co-cultured for 5 days with OVA257-264 peptide (0.1 μg/mL) pulsed GM- or FL-DCs at a 2:1 ratio. Where indicated, DCs were activated with poly (I:C) (25 μg/mL) for 24 hours prior to co-culture. Bars represent mean ± SEM of technical triplicates from one of three independent experiments with similar results. d, Schematic illustration of OT-I T cell transduction and preconditioning with cytokines IL-7 and IL-15 (IL-7/15).
Extended Data Fig. 3 Preconditioning regimens for lymphodepletion and cDC1 expansion following OT-I FL therapy.
a, Analysis of splenic immune cell subsets 2 days after total body irradiation (TBI), representative of one of two independent experiments with similar results. b, Fold change of immune cell subsets normalized to control non-irradiated mice. c, Comparison of splenic immune cell subsets after preconditioning with titrated doses of TBI or cyclophosphamide (CP). d, Fold change of immune cell subsets after 0.5 Gy TBI or 50 mg/kg CP normalized to control mice. e, hHer2 mice were irradiated at indicated doses and subcutaneously inoculated with 5 × 105 Flt3L expressing 24JK-Her2 cells (n = 6 mice/group), from of one of two independent experiments (2-way ANOVA). f–g, Absolute numbers of BM pre-cDC, pre-cDC1 and pre-cDC2 per femur, and splenic cDC1 and cDC2 populations 7 days post adoptive transfer, after preconditioning with 0.5 Gy TBI or 50 mg/kg CP. Bars represent mean ± SEM, a, f, g 1-way ANOVA, symbols represent individual mice.
Extended Data Fig. 4 Flt3L-secreting T cells promote DC proliferation and tumor infiltration of endogenous CD8+ T cells. Related to Fig. 3.
a, BM pre-cDC gating strategy. Lineage markers include Thy1.2, CD19, and NK1.1. b, Frequency of pre-cDC1 and pre-cDC2 as a percentage of total BM pre-cDC (NT n = 4 mice, OT-I and FL n = 6 mice/group). c, Absolute numbers of macrophages and neutrophils per mg of tumor, and d, Frequency of tumor cDC1, cDC2 and cDC1 precursors as a percentage of total tumor DCs post adoptive transfer (NT n = 4, OT-1 n = 6, FL n = 5). e, Adoptive transfer of E0771-OVA tumor-bearing mice with 2 × 107 antigen-specific OT-I T cells or antigen non-specific pmel-1 T cells transduced to secrete Flt3L (OT-I FL and pmel-1 FL, respectively). (Left) Expression of surface NGFR marker in OT-I FL and pmel-1 FL cells. (Mid) Flt3L from tumor supernatants (day 7). (Right) Absolute numbers of tumor cDC1 and cDC1 precursors on day 7 post adoptive transfer with OT-I FL or pmel-1 FL. f, Representative contour plots of tumor cDC1 and cDC2 respectively secreting IL-12 and TNF. g, Absolute numbers of migratory and resident dLN DCs (day 5 NT n = 4 mice, OT-I n = 6, FL n = 5; day 7 NT and FL n = 8/group, OT-I n = 11) and h, splenic cDC1, cDC2, macrophages and neutrophils (NT n = 3 – 4/group, OT-I n = 6/group, FL = 5 – 6/group). Bars represent mean ± SEM, b–e, g, h 1-way ANOVA, a–g Data representative of one of two independent experiments with similar results, d,e Symbols represent individual mice.
Extended Data Fig. 5 Treg and CD62L+TCF−1+ CD8+ T cells in tumors.
a, Absolute numbers of Treg per mg tumor on day 7 post adoptive transfer (1-way ANOVA), from one of two independent experiments. b, Frequency of CD62L+TCF-1+ OT-I or endogenous CD8+ T cells in the tumor on day 7 post adoptive transfer from two pooled independent experiments (unpaired Student’s t-test). Symbols represent individual mice.
Extended Data Fig. 6 Characterization of the effects of adjuvants in vitro and in vivo.
a, Activated BM DCs were pulsed with OVA257-264 before co-culture with naive OT-I T cells (1:2). Frequencies of CD69+ and 4-1BB+ CD8+ OT-I T cells day 5 post co-culture, data represent technical triplicates from one of three independent experiments with similar results. b, Survival (tumor area > 100mm2) from one of two independent experiments (two-tailed Mantel-Cox) of mice subcutaneously inoculated with 5 × 105 MC38 Cherry FL cells or control. Ten days later, mice were injected with poly (I:C:) (200 μg) and/or anti-4-1BB (100 μg). Subsequent poly (I:C) were administered on days 4 and 8 post initial treatment. c, E0771-OVA tumor bearing C57BL/6 WT mice were treated accordingly with OT-I FL, poly (I:C), anti-4-1BB (top; NT and FL + anti-4-1BB n = 5 mice/group, rest n = 4/group; 2-way ANOVA) or adjuvants with no T cells (bottom; n = 5 mice/group). Arrows (top) indicate adjuvants (days 5 and 8). Data points from NT, OT-I FL and OT-I FL + poly (I:C) + anti-4-1BB were included in Fig. 4c pooled data. d, Significantly upregulated genes based on GO term ‘Response to type I Interferon’ (GO: 0034340) (day 6). e, Representative CD40+CD86+ activated DCs (day 7). f–g, Absolute numbers of PMA and ionomycin stimulated, IFN-γ-producing CD8+ T cells on day 7 post adoptive transfer from 2 – 4 pooled independent experiments, symbols represent individual mice, 1-way ANOVA. h, hHer2 mice were subcutaneously injected with 1 × 106 MC38-Her2, preconditioned and treated with 1 × 107 CAR T or CAR T FL cells on days 5 and 6 post tumor inoculation (NT and FL n = 5 mice/group, CAR T n = 6, CAR T + adjuvants n = 12, FL + adjuvants n = 13, 2-way ANOVA, Sidak’s multiple comparison). Arrow indicate adjuvants on day 5 post adoptive transfer. a, c, f–h Bars represent mean ± SEM.
Extended Data Fig. 7 In vitro and in vivo characterization of CD4+ CAR T cells.
a, Frequency of mCherry+ CD4+ and CD8+ CAR T cells from bulk transduced CAR T cell population prior to adoptive transfer, from two pooled independent experiments with n = 2 technical replicates per experiment. b, Frequency of CD4+ and CD8+ mCherry+ CAR T cells in the dLN of E0771-OVA-Her2 tumor-bearing mice on 7 days post adoptive transfer (CAR n = 8 mice, FL n = 9, CAR T + adjuvants n = 12, FL + adjuvants n = 12). Bars represent mean ± SEM.
Extended Data Fig. 8 Safety assessment of Flt3L-secreting T cells and adjuvant therapy.
hHer2 transgenic mice were inoculated with 4 × 105 E0771-OVA-Her2 tumor cells and treated with transduced CAR T cells as per Fig. 6f. Mice were monitored for changes in weight. Alternatively, sera and organs were assessed at 48 hours post adjuvant therapy for toxicity studies. a, Liver function and/or damage was assessed by serum alanine transaminase (ALT), aspartate aminotransferase (AST), total albumin, bilirubin, and gamma glutamyl transferase (GGT) levels. b, Kidney function and/or damage was assessed by serum urea and creatinine levels. c, Sera were assessed for levels of IFN-γ, IL-1β, IL-6, GM-CSF, G-CSF, MCP-1 and TNF cytokines. d, Percentage weight change relative to day 0 measurements. Dotted line (90%) indicates the maximum ethical limit allowed for weight reduction. Arrows indicate adjuvant administration (NT n = 5 mice, CAR T and FL + Isotype n = 6 mice/group, CAR T and FL + adjuvants n = 8/group). e, Representative hematoxylin and eosin histology staining of cerebellum, liver, kidney, lung and spleen of control non-treated (NT), CAR T or CAR T FL adjuvant treated mice (3 mice/group). f, Sera were assessed for antinuclear antibodies as an indication of autoimmunity (n = 5 mice/group). Sera from NZB/W (with proteinuria; n = 3) and C57BL/6 mice (n = 3) immunized with an irrelevant antigen (NP-KLH) were respectively used as positive and negative controls. Bars represent mean ± SEM, a–c Symbols represent individual mice, 1-way ANOVA.
Extended Data Fig. 9 Assessment of BM hematopoietic stem cells and progenitors.
a, Gating strategy for BM populations. b, Absolute numbers of hematopoietic stem cells (HSC), common lymphoid progenitors (CLP), common dendritic progenitors (CDP) and myeloid and dendritic progenitors (MDP) in the BM from E0771-OVA tumor-bearing mice 7 or 14 days post adoptive transfer with Flt3L-secreting OT-I T cells. Data shown as mean ± SEM of n = 5 mice/group, 1-way ANOVA.
Supplementary information
Supplementary Information
Supplementary Figs. 1 and 2.
Supplementary Table
Information on antibodies, reagents, software and service platforms.
Source data
Source Data Fig. 1
Statistical Source Data for Fig. 1.
Source Data Fig. 2
Statistical Source Data for Fig. 2.
Source Data Fig. 3
Statistical Source Data for Fig. 3.
Source Data Fig. 4
Statistical Source Data for Fig. 4.
Source Data Fig. 5
Statistical Source Data for Fig. 5.
Source Data Fig. 6
Statistical Source Data for Fig. 6.
Source Data Fig. 7
Statistical Source Data for Fig. 7.
Source Data Extended Data Fig. 1
Statistical Source Data for Extended Data Fig. 1.
Source Data Extended Data Fig. 2
Statistical Source Data for Extended Data Fig. 2.
Source Data Extended Data Fig. 3
Statistical Source Data for Extended Data Fig. 3.
Source Data Extended Data Fig. 4
Statistical Source Data for Extended Data Fig. 4.
Source Data Extended Data Fig. 5
Statistical Source Data for Extended Data Fig. 5.
Source Data Extended Data Fig. 6
Statistical Source Data for Extended Data Fig. 6.
Source Data Extended Data Fig. 7
Statistical Source Data for Extended Data Fig. 7.
Source Data Extended Data Fig. 8
Statistical Source Data for Extended Data Fig. 8.
Source Data Extended Data Fig. 9
Statistical Source Data for Extended Data Fig. 9.
Rights and permissions
About this article
Cite this article
Lai, J., Mardiana, S., House, I.G. et al. Adoptive cellular therapy with T cells expressing the dendritic cell growth factor Flt3L drives epitope spreading and antitumor immunity. Nat Immunol 21, 914–926 (2020). https://doi.org/10.1038/s41590-020-0676-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-020-0676-7
This article is cited by
-
FLT3L-induced virtual memory CD8 T cells engage the immune system against tumors
Journal of Biomedical Science (2024)
-
CAR T cell therapy for patients with solid tumours: key lessons to learn and unlearn
Nature Reviews Clinical Oncology (2024)
-
Dendritic cell-targeted therapy expands CD8 T cell responses to bona-fide neoantigens in lung tumors
Nature Communications (2024)
-
From complexity to clarity: unravelling tumor heterogeneity through the lens of tumor microenvironment for innovative cancer therapy
Histochemistry and Cell Biology (2024)
-
An oncolytic virus delivering tumor-irrelevant bystander T cell epitopes induces anti-tumor immunity and potentiates cancer immunotherapy
Nature Cancer (2024)