Abstract
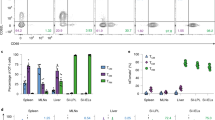
Tissue-resident memory T (TRM) cells, functionally distinct from circulating memory T cells, have a critical role in protective immunity in tissues, are more efficacious when elicited after vaccination and yield more effective antitumor immunity, yet the signals that direct development of TRM cells are incompletely understood. Here we show that type 1 regulatory T (Treg) cells, which express the transcription factor T-bet, promote the generation of CD8+ TRM cells. The absence of T-bet-expressing type 1 Treg cells reduces the presence of TRM cells in multiple tissues and increases pathogen burden upon infectious challenge. Using infection models, we show that type 1 Treg cells are specifically recruited to local inflammatory sites via the chemokine receptor CXCR3. Close proximity with effector CD8+ T cells and Treg cell expression of integrin-β8 endows the bioavailability of transforming growth factor-β in the microenvironment, thereby promoting the generation of CD8+ TRM cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Masopust, D., Vezys, V., Marzo, A. L. & Lefrancois, L. Preferential localization of effector memory cells in nonlymphoid tissue. Science 291, 2413–2417 (2001).
Reinhardt, R. L., Khoruts, A., Merica, R., Zell, T. & Jenkins, M. K. Visualizing the generation of memory CD4+ T cells in the whole body. Nature 410, 101–105 (2001).
Mueller, S. N. & Mackay, L. K. Tissue-resident memory T cells: local specialists in immune defence. Nat. Rev. Immunol. 16, 79–89 (2016).
Konjar, S., Ferreira, C., Blankenhaus, B. & Veldhoen, M. Intestinal barrier interactions with specialized CD8+ T Cells. Front. Immunol. 8, 1281 (2017).
Ariotti, S. et al. Tissue-resident memory CD8+ T cells continuously patrol skin epithelia to quickly recognize local antigen. PNAS 109, 19739–19744 (2012).
Konjar, S. et al. Mitochondria maintain controlled activation state of epithelial-resident T lymphocytes. Sci. Immunol. 3, eaan2543 (2018).
Joshi, N. S. et al. Inflammation directs memory precursor and short-lived effector CD8(+) T cell fates via the graded expression of T-bet transcription factor. Immunity 27, 281–295 (2007).
Intlekofer, A. M. et al. Requirement for T-bet in the aberrant differentiation of unhelped memory CD8+ T cells. J. Exp. Med. 204, 2015–2021 (2007).
Joshi, N. S. et al. Increased numbers of preexisting memory CD8+ T cells and decreased T-bet expression can restrain terminal differentiation of secondary effector and memory CD8+ T cells. J. Immunol. 187, 4068–4076 (2011).
Pipkin, M. E. et al. Interleukin-2 and inflammation induce distinct transcriptional programs that promote the differentiation of effector cytolytic T cells. Immunity 32, 79–90 (2010).
Mackay, L. K. et al. The developmental pathway for CD103(+)CD8+ tissue-resident memory T cells of skin. Nat. Immunol. 14, 1294–1301 (2013).
Wakim, L. M. et al. The molecular signature of tissue-resident memory CD8+ T cells isolated from the brain. J. Immunol. 189, 3462–3471 (2012).
Zhang, N. & Bevan, M. J. Transforming growth factor-β signaling controls the formation and maintenance of gut-resident memory T cells by regulating migration and retention. Immunity 39, 687–696 (2013).
Laidlaw, B. J. et al. CD4+ T cell help guides formation of CD103+ lung-resident memory CD8+ T cells during influenza viral infection. Immunity 41, 633–645 (2014).
Mackay, L. K. et al. T-box transcription factors combine with the cytokines TGF-β and IL-15 to control tissue-resident memory T cell fate. Immunity 43, 1101–1111 (2015).
Laidlaw, B. J., Craft, J. E. & Kaech, S. M. The multifaceted role of CD4(+) T cells in CD8(+) T cell memory. Nat. Rev. Immunol. 16, 102–111 (2016).
Feau, S., Arens, R., Togher, S. & Schoenberger, S. P. Autocrine IL-2 is required for secondary population expansion of CD8(+) memory T cells. Nat. Immunol. 12, 908–913 (2011).
Yu, F., Sharma, S., Edwards, J., Feigenbaum, L. & Zhu, J. Dynamic expression of transcription factors T-bet and GATA-3 by regulatory T cells maintains immunotolerance. Nat. Immunol. 16, 197–206 (2015).
de Goer de Herve, M. G., Jaafoura, S., Vallee, M. & Taoufik, Y. FoxP3(+) regulatory CD4+ T cells control the generation of functional CD8+ memory. Nat. Commun. 3, 986 (2012).
Pace, L. et al. Regulatory T cells increase the avidity of primary CD8+ T cell responses and promote memory. Science 338, 532–536 (2012).
Laidlaw, B. J. et al. Production of IL-10 by CD4(+) regulatory T cells during the resolution of infection promotes the maturation of memory CD8(+) T cells. Nat. Immunol. 16, 871–879 (2015).
Kalia, V., Penny, L. A., Yuzefpolskiy, Y., Baumann, F. M. & Sarkar, S. Quiescence of memory CD8(+) T cells is mediated by regulatory T cells through inhibitory receptor CTLA-4. Immunity 42, 1116–1129 (2015).
Koch, M. A. et al. The transcription factor T-bet controls regulatory T cell homeostasis and function during type 1 inflammation. Nat. Immunol. 10, 595–602 (2009).
Lord, G. M. et al. T-bet is required for optimal proinflammatory CD4+ T cell trafficking. Blood 106, 3432–3439 (2005).
Levine, A. G. et al. Stability and function of regulatory T cells expressing the transcription factor T-bet. Nature 546, 421–425 (2017).
Tan, T. G., Mathis, D. & Benoist, C. Singular role for T-BET+CXCR3+ regulatory T cells in protection from autoimmune diabetes. PNAS 113, 14103–14108 (2016).
Janssen, E. M. et al. CD4+ T cell help controls CD8+ T cell memory via TRAIL-mediated activation-induced cell death. Nature 434, 88–93 (2005).
Oh, S. et al. IL-15 as a mediator of CD4+ help for CD8+ T cell longevity and avoidance of TRAIL-mediated apoptosis. PNAS 105, 5201–5206 (2008).
Madakamutil, L. T. et al. CD8αα-mediated survival and differentiation of CD8+ memory T cell precursors. Science 304, 590–593 (2004).
Herndler-Brandstetter, D. et al. KLRG1(+) Effector CD8(+) T cells lose KLRG1, differentiate into all memory T cell lineages, and convey enhanced protective immunity. Immunity 48, 716–729 (2018).
Rose, M. E., Wakelin, D. & Hesketh, P. Gamma-interferon controls Eimeria vermiformis primary infection in BALB/c mice. Infect. Immun. 57, 1599–1603 (1989).
Belkaid, Y., Piccirillo, C. A., Mendez, S., Shevach, E. M. & Sacks, D. L. CD4+CD25+ regulatory T cells control Leishmania major persistence and immunity. Nature 420, 502–507 (2002).
Fernandez-Ruiz, D. et al. Liver-resident memory CD8(+) T cells form a front-line defense against malaria liver-stage infection. Immunity 45, 889–902 (2016).
Ramsburg, E., Tigelaar, R., Craft, J. & Hayday, A. Age-dependent requirement for γδ T cells in the primary but not secondary protective immune response against an intestinal parasite. J. Exp. Med. 198, 1403–1414 (2003).
Miragaia, R. J. et al. Single-cell transcriptomics of regulatory T cells reveals trajectories of tissue adaptation. Immunity 50, 493–504 (2019).
Zemmour, D. et al. Single-cell gene expression reveals a landscape of regulatory T cell phenotypes shaped by the TCR. Nat. Immunol. 19, 291–301 (2018).
Xiong, Y., Ahmad, S., Iwami, D., Brinkman, C. C. & Bromberg, J. S. T-bet regulates natural regulatory T cell afferent lymphatic migration and suppressive function. J. Immunol. 196, 2526–2540 (2016).
Bergsbaken, T. & Bevan, M. J. Proinflammatory microenvironments within the intestine regulate the differentiation of tissue-resident CD8(+) T cells responding to infection. Nat. Immunol. 16, 406–414 (2015).
Bergsbaken, T., Bevan, M. J. & Fink, P. J. Local inflammatory cues regulate differentiation and persistence of CD8(+) tissue-resident memory T cells. Cell Rep. 19, 114–124 (2017).
Khan, T. N., Mooster, J. L., Kilgore, A. M., Osborn, J. F. & Nolz, J. C. Local antigen in nonlymphoid tissue promotes resident memory CD8+ T cell formation during viral infection. J. Exp. Med. 213, 951–966 (2016).
Wakim, L. M., Woodward-Davis, A. & Bevan, M. J. Memory T cells persisting within the brain after local infection show functional adaptations to their tissue of residence. PNAS 107, 17872–17879 (2010).
Iijima, N. & Iwasaki, A. T cell memory. A local macrophage chemokine network sustains protective tissue-resident memory CD4+ T cells. Science 346, 93–98 (2014).
Natsuaki, Y. et al. Perivascular leukocyte clusters are essential for efficient activation of effector T cells in the skin. Nat. Immunol. 15, 1064–1069 (2014).
Worthington, J. J. et al. Integrin αvβ8-mediated TGF-β activation by effector regulatory T cells is essential for suppression of T cell-mediated inflammation. Immunity 42, 903–915 (2015).
Masopust, D. et al. Dynamic T cell migration program provides resident memory within intestinal epithelium. J. Exp. Med. 207, 553–564 (2010).
Butcher, E. C. & Picker, L. J. Lymphocyte homing and homeostasis. Science 272, 60–66 (1996).
Gebhardt, T. et al. Memory T cells in nonlymphoid tissue that provide enhanced local immunity during infection with herpes simplex virus. Nat. Immunol. 10, 524–530 (2009).
Mackay, L. K. et al. Long-lived epithelial immunity by tissue-resident memory T (TRM) cells in the absence of persisting local antigen presentation. PNAS 109, 7037–7042 (2012).
Graham, J. B., Da Costa, A. & Lund, J. M. Regulatory T cells shape the resident memory T cell response to virus infection in the tissues. J. Immunol. 192, 683–690 (2014).
Hickman, H. D. et al. CXCR3 chemokine receptor enables local CD8(+) T cell migration for the destruction of virus-infected cells. Immunity 42, 524–537 (2015).
Paust, H. J. et al. CXCR3+ regulatory T cells control TH1 responses in crescentic GN. J. Am. Soc. Nephrol. 27, 1933–1942 (2016).
Harrison, O. J. et al. Commensal-specific T cell plasticity promotes rapid tissue adaptation to injury. Science 363, eaat6280 (2019).
Casey, K. A. et al. Antigen-independent differentiation and maintenance of effector-like resident memory T cells in tissues. J. Immunol. 188, 4866–4875 (2012).
Mohammed, J. et al. Stromal cells control the epithelial residence of DCs and memory T cells by regulated activation of TGF-β. Nat. Immunol. 17, 414–421 (2016).
Intlekofer, A. M. et al. Anomalous type 17 response to viral infection by CD8+ T cells lacking T-bet and eomesodermin. Science 321, 408–411 (2008).
Rubtsov, Y. P. et al. Regulatory T cell-derived interleukin-10 limits inflammation at environmental interfaces. Immunity 28, 546–558 (2008).
Luche, H., Weber, O., Nageswara Rao, T., Blum, C. & Fehling, H. J. Faithful activation of an extra-bright red fluorescent protein in ‘knock-in’ Cre-reporter mice ideally suited for lineage tracing studies. Eur. J. Immunol. 37, 43–53 (2007).
Hancock, W. W. et al. Requirement of the chemokine receptor CXCR3 for acute allograft rejection. J. Exp. Med. 192, 1515–1520 (2000).
Kuhn, R., Lohler, J., Rennick, D., Rajewsky, K. & Muller, W. Interleukin-10-deficient mice develop chronic enterocolitis. Cell 75, 263–274 (1993).
Nieuwenhuis, E. E. et al. Disruption of T helper 2-immune responses in Epstein–Barr virus-induced gene 3-deficient mice. PNAS 99, 16951–16956 (2002).
Travis, M. A. et al. Loss of integrin α(v)β8 on dendritic cells causes autoimmunity and colitis in mice. Nature 449, 361–365 (2007).
Azhar, M. et al. Generation of mice with a conditional allele for transforming growth factor-β 1 gene. Genesis 47, 423–431 (2009).
Li, Y. et al. Exogenous stimuli maintain intraepithelial lymphocytes via aryl hydrocarbon receptor activation. Cell 147, 629–640 (2011).
Figueiredo-Campo, P., Ferreira, C., Blankenhaus, B. & Veldhoen, M. Eimeria vermiformis infection model of murine small intestine. Bio. Protoc. 8, e3122 (2018).
Cossarizza, A. et al. Guidelines for the use of flow cytometry and cell sorting in immunological studies (second edition). Eur. J. Immunol. 49, 1457–1973 (2019).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Acknowledgements
We acknowledge the excellent contributions from the iMM flow cytometry, rodent, histology and microscopy facilities. The project leading to these results has received funding from the European Union H2020 ERA project (no. 667824, EXCELLtoINNOV), Fundo iMM-Laço and ‘la caixa’ Foundation (ID 100010434) under agreement LCF/PR/HR19/52160005 for work in the Veldhoen laboratory, with additional funding from the Fundação para a Ciência e a Tecnologia to P.F.-C. (SFRH/BD/131605/2017), to L.B. (PD/BD/138847/2018) and to A.B. (SFRH/BD/138900/2018). In addition, the work was funded by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) SFB1366 (project no. 394046768-SFB 1366; C02) and by SPP 1937 (CE 140/2-1) to A.C and SFB1292 (TP13) and TR156 (TPB02) to H.C.P. A.L. was supported by Ligue Nationale Contre le Cancer. Other grants supporting this study: Foncer Contre le Cancer (JCM) and EL-2016 LNCC Labelisation Ligue Nationale Contre Cancer. TL-tetramer was obtained through the National Institutes of Health Tetramer Core Facility.
Author information
Authors and Affiliations
Contributions
C.F., L.B., S.K., B.B., P.F.-C. and M.B. performed the experiments and provided assistance. A.B. performed bioinformatics analysis. A.S. and A.C. provided CXCR3-deficient animals. N.K. and H.C.P. provided EBI3-deficient animals and performed analysis. A.L. and J.C.M. provided Foxp3Cre Itgb8f/f and Foxp3Cre Tgfb1f/f animals and performed analysis. C.F. and M.V. conceived and directed the experiments and wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Zoltan Fehervari was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–8.
Rights and permissions
About this article
Cite this article
Ferreira, C., Barros, L., Baptista, M. et al. Type 1 Treg cells promote the generation of CD8+ tissue-resident memory T cells. Nat Immunol 21, 766–776 (2020). https://doi.org/10.1038/s41590-020-0674-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-020-0674-9
This article is cited by
-
Tissue-resident memory T cells: decoding intra-organ diversity with a gut perspective
Inflammation and Regeneration (2024)
-
Structural insights into the activation and inhibition of CXC chemokine receptor 3
Nature Structural & Molecular Biology (2024)
-
Transient regulatory-T-cell interruption promotes skin-resident memory T cells mediated tumor protection
Scientific Reports (2023)
-
IFNγ-induction of TH1-like regulatory T cells controls antiviral responses
Nature Immunology (2023)
-
Construction and validation of a prognostic model for colon adenocarcinoma based on bile acid metabolism-related genes
Scientific Reports (2023)