Abstract
Efficient generation of germinal center (GC) responses requires directed movement of B cells between distinct microenvironments underpinned by specialized B cell–interacting reticular cells (BRCs). How BRCs are reprogrammed to cater to the developing GC remains unclear, and studying this process is largely hindered by incomplete resolution of the cellular composition of the B cell follicle. Here we used genetic targeting of Cxcl13-expressing cells to define the molecular identity of the BRC landscape. Single-cell transcriptomic analysis revealed that BRC subset specification was predetermined in the primary B cell follicle. Further topological remodeling of light and dark zone follicular dendritic cells required CXCL12-dependent crosstalk with B cells and dictated GC output by retaining B cells in the follicle and steering their interaction with follicular helper T cells. Together, our results reveal that poised BRC-defined microenvironments establish a feed-forward system that determines the efficacy of the GC reaction.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All scRNA-seq and BCR-seq datasets are available in ArrayExpress (accession numbers E-MTAB-8445 (Cxcl13-Cre+ cell scRNA-seq), E-MTAB-8454 (B cell scRNA-seq and BCR-seq; VSV immunization) and E-MTAB-8904 (BCR-seq; NP–KLH immunization). The data that support the findings of this study are available from the corresponding author.
References
Schwickert, T. A. et al. A dynamic T cell–limited checkpoint regulates affinity-dependent B cell entry into the germinal center. J. Exp. Med. 208, 1243–1252 (2011).
Sander, S. et al. PI3 kinase and FOXO1 transcription factor activity differentially control B cells in the germinal center light and dark zones. Immunity 43, 1075–1086 (2015).
Dominguez-Sola, D. et al. The FOXO1 transcription factor instructs the germinal center dark zone program. Immunity 43, 1064–1074 (2015).
Shinnakasu, R. et al. Regulated selection of germinal-center cells into the memory B cell compartment. Nat. Immunol. 17, 861–869 (2016).
Gitlin, A. D., Shulman, Z. & Nussenzweig, M. C. Clonal selection in the germinal centre by regulated proliferation and hypermutation. Nature 509, 637–640 (2014).
Fletcher, A. L. et al. Lymph node fibroblastic reticular cells directly present peripheral tissue antigen under steady-state and inflammatory conditions. J. Exp. Med. 207, 689–697 (2010).
Chai, Q. et al. Maturation of lymph node fibroblastic reticular cells from myofibroblastic precursors is critical for antiviral immunity. Immunity 38, 1013–1024 (2013).
Gil-Cruz, C. et al. Fibroblastic reticular cells regulate intestinal inflammation via IL-15-mediated control of group 1 ILCs. Nat. Immunol. 17, 1388–1396 (2016).
Perez-Shibayama, C. et al. Fibroblastic reticular cells initiate immune responses in visceral adipose tissues and secure peritoneal immunity. Sci. Immunol. 3, eaar4539 (2018).
Cremasco, V. et al. B cell homeostasis and follicle confines are governed by fibroblastic reticular cells. Nat. Immunol. 15, 973–981 (2014).
Dubey, L. K. et al. Lymphotoxin-dependent B cell–FRC crosstalk promotes de novo follicle formation and antibody production following intestinal helminth infection. Cell Rep. 15, 1527–1541 (2016).
Zhang, Y. et al. Plasma cell output from germinal centers is regulated by signals from Tfh and stromal cells. J. Exp. Med. 215, 1227–1243 (2018).
Suzuki, K., Grigorova, I., Phan, T. G., Kelly, L. M. & Cyster, J. G. Visualizing B cell capture of cognate antigen from follicular dendritic cells. J. Exp. Med. 206, 1485–1493 (2009).
Barrington, R. A., Pozdnyakova, O., Zafari, M. R., Benjamin, C. D. & Carroll, M. C. B lymphocyte memory: role of stromal cell complement and FcγRIIB receptors. J. Exp. Med. 196, 1189–1199 (2002).
Heesters, B. A. et al. Endocytosis and recycling of immune complexes by follicular dendritic cells enhances B cell antigen binding and activation. Immunity 38, 1164–1175 (2013).
Allen, C. D. et al. Germinal center dark and light zone organization is mediated by CXCR4 and CXCR5. Nat. Immunol. 5, 943–952 (2004).
Pereira, J. P., Kelly, L. M. & Cyster, J. G. Finding the right niche: B-cell migration in the early phases of T-dependent antibody responses. Int. Immunol. 22, 413–419 (2010).
Bannard, O. et al. Germinal center centroblasts transition to a centrocyte phenotype according to a timed program and depend on the dark zone for effective selection. Immunity 39, 912–924 (2013).
Mionnet, C. et al. Identification of a new stromal cell type involved in the regulation of inflamed B cell follicles. PLoS Biol. 11, e1001672 (2013).
Rodda, L. B. et al. Single-cell RNA sequencing of lymph node stromal cells reveals niche-associated heterogeneity. Immunity 48, 1014–1028 (2018).
Onder, L. et al. Lymphatic endothelial cells control initiation of lymph node organogenesis. Immunity 47, 80–92 (2017).
Ansel, K. M. et al. A chemokine-driven positive feedback loop organizes lymphoid follicles. Nature 406, 309–314 (2000).
Mackay, F. & Browning, J. L. Turning off follicular dendritic cells. Nature 395, 26–27 (1998).
Wang, X. et al. Follicular dendritic cells help establish follicle identity and promote B cell retention in germinal centers. J. Exp. Med. 208, 2497–2510 (2011).
Rodda, L. B., Bannard, O., Ludewig, B., Nagasawa, T. & Cyster, J. G. Phenotypic and morphological properties of germinal center dark zone Cxcl12-expressing reticular cells. J. Immunol. 195, 4781–4791 (2015).
Fink, K. et al. B cell activation state-governed formation of germinal centers following viral infection. J. Immunol. 179, 5877–5885 (2007).
Kalinke, U. et al. The role of somatic mutation in the generation of the protective humoral immune response against vesicular stomatitis virus. Immunity 5, 639–652 (1996).
Calado, D. P. et al. The cell-cycle regulator c-Myc is essential for the formation and maintenance of germinal centers. Nat. Immunol. 13, 1092–1100 (2012).
Ersching, J. et al. Germinal center selection and affinity maturation require dynamic regulation of mTORC1 kinase. Immunity 46, 1045–1058 (2017).
Spillane, K. M. & Tolar, P. B cell antigen extraction is regulated by physical properties of antigen-presenting cells. J. Cell Biol. 216, 217–230 (2017).
Moran, I. et al. Memory B cells are reactivated in subcapsular proliferative foci of lymph nodes. Nat. Commun. 9, 3372 (2018).
Roco, J. A. et al. Class-switch recombination occurs infrequently in germinal centers. Immunity 51, 337–350 (2019).
Nie, Y. et al. The role of CXCR4 in maintaining peripheral B cell compartments and humoral immunity. J. Exp. Med. 200, 1145–1156 (2004).
Gitlin, A. D. et al. T cell help controls the speed of the cell cycle in germinal center B cells. Science 349, 643–646 (2015).
Pratama, A. & Vinuesa, C. G. Control of TFH cell numbers: why and how? Immunol. Cell Biol. 92, 40–48 (2014).
Ara, T. et al. Long-term hematopoietic stem cells require stromal cell-derived factor-1 for colonizing bone marrow during ontogeny. Immunity 19, 257–267 (2003).
Greenbaum, A. et al. CXCL12 in early mesenchymal progenitors is required for haematopoietic stem-cell maintenance. Nature 495, 227–230 (2013).
Wimmer, N. et al. Lymphotoxin β receptor activation on macrophages induces cross-tolerance to TLR4 and TLR9 ligands. J. Immunol. 188, 3426–3433 (2012).
Ludewig, B. et al. Induction of optimal anti-viral neutralizing B cell responses by dendritic cells requires transport and release of virus particles in secondary lymphoid organs. Eur. J. Immunol. 30, 185–196 (2000).
Cunningham, A. F. et al. Salmonella induces a switched antibody response without germinal centers that impedes the extracellular spread of infection. J. Immunol. 178, 6200–6207 (2007).
Cervantes-Barragan, L. et al. TLR2 and TLR4 signaling shapes specific antibody responses to Salmonella typhi antigens. Eur. J. Immunol. 39, 126–135 (2009).
Cossarizza, A. et al. Guidelines for the use of flow cytometry and cell sorting in immunological studies (second edition). Eur. J. Immunol. 49, 1457–1973 (2019).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Zheng, G. X. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
McCarthy, D. J., Campbell, K. R., Lun, A. T. & Wills, Q. F. Scater: pre-processing, quality control, normalization and visualization of single-cell RNA-seq data in R. Bioinformatics 33, 1179–1186 (2017).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Kolde, R. Pheatmap: pretty heatmaps. R package version 61 (2012).
Allen, C. D., Okada, T., Tang, H. L. & Cyster, J. G. Imaging of germinal center selection events during affinity maturation. Science 315, 528–531 (2007).
Lun, A. T. L., McCarthy, D. J. & Marioni, J. C. A step-by-step workflow for low-level analysis of single-cell RNA-seq data with Bioconductor [version 2]. F1000Res 5, 2122 (2016).
Reich, M. et al. The GenePattern Notebook Environment. Cell Syst. 5, 149–151 (2017).
Street, K. et al. Slingshot: cell lineage and pseudotime inference for single-cell transcriptomics. BMC Genomics 19, 477 (2018).
Gupta, N. T. et al. Change-O: a toolkit for analyzing large-scale B cell immunoglobulin repertoire sequencing data. Bioinformatics 31, 3356–3358 (2015).
Acknowledgements
This study received financial support from the Swiss National Science Foundation (grants 177208, 166500 and 159188 to B.L. and grant 180011 to N.B.P.) and the Novartis Foundation for Biomedical Research. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
B.L. designed the study, discussed data and wrote the paper. N.B.P. designed the study, performed experiments, analyzed data and wrote the paper. U.M., C.G.-C., C.P.-S., M.N., H.-W.C. and L.O. performed experiments and analyzed and discussed data. M.L. performed bioinformatics analyses and discussed data. M.A.L. provided reagents and discussed data. C.N.-A. and T.N. provided reagents.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information L. A. Dempsey was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The Cxcl13-Cre/TdTom transgene faithfully demarcates non-endothelial, non-hematopoietic, CXCL13-expressing cells.
a, Representative immunofluorescence images of EYFP, TdTom, and CXCL13 expression in naive Cxcl13-Cre/TdTom EYFP mice. Scale bars, 100 μm and 10 μm. b, Representative images of TdTom expression in the medullary cords of naive and day 12 VSV-immunized Cxcl13-Cre/TdTom EYFP mice. Scale bars, 500 μm and 100 μm. c, Quantification of the percentage of EYFP+ cells expressing CD31 or CD45 in naive Cxcl13-Cre/TdTom EYFP mice. Mean and SEM are depicted. d, Representative images of EYFP, TdTom and podoplanin (PDPN) expression in naive Cxcl13-Cre/TdTom EYFP mice. Arrows point to the cell body. Scale bars, 50 μm and 20 μm. e, Flow cytometric gating strategy of non-hematopoietic, non-endothelial reticular cells from Cxcl13-Cre/TdTom EYFP mice. (a, b, d) Images are representative of at least five mice. (c, e) N = 8 naive mice, 4 independent experiments; P values as per one-way ANOVA with Tukey’s multiple comparisons test.
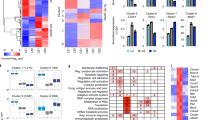
Extended Data Fig. 2 Single cell transcriptomic analysis of B cell-interacting reticular cells.
a, Schematic representation of the experimental setup. b, Heatmap of the averaged gene expression of top cluster-specific genes across all BRC subsets. Hierarchical grouping based on cluster-specific marker genes is depicted in the left panel. c, Feature plots depicting the expression of the indicated genes. d, Representative overview of RANKL staining in naïve Cxcl13-Cre/TdTom mouse lymph nodes. e, The indicated interfollicular region from (d) stained for B220, RANKL and MAdCAM1. Images are representative of at least three mice.
Extended Data Fig. 3 Molecular and topological identity of light and dark zone FDC at a single cell level.
a, b, Strategy for demarcating distinct regions of the B cell follicle for topological BRC analysis using positioning, TdTom fluorescence intensity, and IgD and CD21/35 immunostaining. Each dot in (b) represents one cell body, color-coded according to the occupied region in the follicle. Scale bars, 100 μm. c, BRC enumeration per region of B cell follicle. d, Quantification of the mean fluorescence intensity (MFI) of TdTom in each region of the B cell follicle (according to color-code). e, f, Merged images and single channels corresponding to Fig. 3c,d. Scale bars, 100 μm. Images are representative of at least two mice per condition. g, Confocal microscopy analysis of IgD, TdTom, CD21/35 and MYH11 staining in secondary B cell follicles of Cxcl13-Cre/TdTom EYFP mice. Images are representative of at least 5 immunized Cxcl13-Cre/TdTom mice. (c, d). N = 7 naïve Cxcl13-Cre/TdTom mice, 3 independent experiments, N = 5 VSV-immunized Cxcl13-Cre/TdTom mice, 2 independent experiments. Mean and SEM are depicted. P values as per one-way ANOVA with Tukey’s multiple comparisons test.
Extended Data Fig. 4 LTβR signaling in Cxcl13-Cre+ cells governs BRC maturation.
a, b, Representative immunofluorescence images of EYFP, TdTom and B220 staining in lymph nodes (a) and regions underpinning B cell aggregates (b) in naive Cxcl13-Cre/TdTom Ltbrfl/fl mice (Ltbrfl/fl). c, Enumeration of TdTom+ EYFP+ cells per lymph node (LN) in naive Cxcl13-Cre/TdTom control or Ltbrfl/fl mice. d, Quantification of the mean fluorescence intensity (MFI) of TdTom on EYFP+ cells from naive control or Ltbrfl/fl mice. e, Representative flow cytometry plots of MAdCAM1 and CD21/35 on TdTom+ EYFP+ cells from naive Ltbrfl/fl mice. Mean percentages and SEM are indicated. N = 14 mice, 4 independent experiments. f, g, Representative immunofluorescence images of EYFP, TdTom and B220 staining in lymph nodes (f) and regions underpinning B cell aggregates (g) in day 12 VSV-immunized Ltbrfl/fl mice. h, Enumeration of TdTom+ EYFP+ cells per lymph node in immunized control or Ltbrfl/fl mice. i, Quantification of the MFI of TdTom on EYFP+ cells from lymph nodes of immunized control or Ltbrfl/fl mice. j, Representative flow cytometry plots of MAdCAM1 and CD21/35 on TdTom+ EYFP+ cells from immunized Ltbrfl/fl mice. Mean percentages and SEM are indicated. N = 13 Ltbrfl/fl mice, 5 independent experiments. (a, b, f, g) Images are representative of 6 Ltbrfl/fl per condition. (c, d) N = 11 control mice and 13 Ltbrfl/fl mice; 4 independent experiments. (h, i) N = 16 control mice and 11 Ltbrfl/fl mice, 4 independent experiments. (c, d, h, i) Mean and SEM are indicated. P values as per two-tailed Mann Whitney test.
Extended Data Fig. 5 CXCL12 expression in in Cxcl13-Cre+ cells governs BRC topological remodeling.
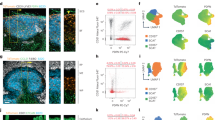
a, b, Percentage (a) and number (b) per lymph node (LN) of the indicated hematopoietic cell populations in naive Cxcl13-Cre/TdTom control, Cxcl12fl/fl and Ltbrfl/fl mice. N = 9 control mice, 7 Cxcl12fl/fl, and 8 Ltbrfl/fl mice; 2 independent experiments. c–e, Scaled gene expression of naive versus immunized LZ FDC (c), DZ FDC (d) or TBRC (e). Red dots indicate differentially expressed genes with an adjusted p-value < 0.01 and an effect size (logFC) > 0.4. f, Left hand panels depict representative images of CD21/35 staining in immunized control and Cxcl12fl/fl mice. The yellow dotted line demarcates the GC perimeter according to IgD staining (inset). Right hand panels depict the volumetric reconstruction of CD21/35 fluorescence and the division of the GC into quadrants. g, Quantification of the volume of CD21/35 per GC quadrant in control and Cxcl12fl/fl mice. N = 5 control and 4 Cxcl12fl/fl mice. Mean and SEM are indicated. h, GFP, CD21/35, PDLIM3 and B220 staining in lymph nodes from naïve Cxcl12-GFP mice. Data is representative of 2 Cxcl12-GFP mice. i, Representative images of PDLIM3, B220 and EYFP in naïve Cxcl13-Cre/TdTom control or Cxcl12fl/fl mice. N = 3 mice per condition. j, Left hand panels show representative images of Ki67, TdTom and CD21/35 staining in immunized control and Cxcl12fl/fl mice. The yellow dotted line demarcates the GC perimeter. Right hand panels depict Ki67 staining and the demarcation of GC quadrants. k, Enumeration of Ki67+ cells per GC quadrant in control and Cxcl12fl/fl mice. N = 5 control and 5 Cxcl12fl/fl mice. (a, b, g, k) Mean and SEM are indicated. P values as per two-way Anova with Bonferroni multiple comparisons test.
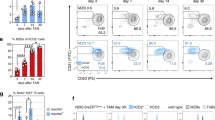
Extended Data Fig. 6 CXCL12 production by Cxcl13-Cre+ cells is required for efficient humoral immunity.
a, Enumeration of CD19+ cells per lymph node (LN) in naive or VSV-immunized control mice, or immunized Cxcl13-Cre/TdTom Cxcl12fl/fl (Cxcl12fl/fl) and Cxcl13-Cre/TdTom Ltbrfl/fl (Ltbrfl/fl) mice. N = 5 naive mice; 27 littermate control mice, 5 independent experiments; 20 Cxcl12fl/fl, 5 independent experiments; 12 Ltbrfl/fl mice, 4 independent experiments. b, c, Representative flow cytometry plots and quantification of CD86− CXCR4+ DZ and CD86+ CXCR4− LZ germinal center CD19+ cells in immunized control, Cxcl12fl/fl mice or Ltbrfl/fl mice. N = 20 immunized littermate controls, 5 independent experiments; 13 Cxcl12fl/fl and 9 Ltbrfl/fl mice, 3 independent experiments. d, e, Expression of DAPI in GL7+ B cells and quantification of GC B cells in the indicated cell cycle stages. N = 15 littermate control mice, 3 independent experiments; 4 Cxcl12fl/fl mice, 2 experiments; 13 Ltbrfl/fl mice, 2 experiments. f, Representative flow cytometry plots of 7AAD and Annexin V incorporation by GL7+ B cells in immunized littermate control, Cxcl12fl/fl mice or Ltbrfl/fl mice. g, Quantification of the mean fluorescence intensity (MFI) of Annexin V on GL7+ B cells from littermate control, Cxcl12fl/fl mice or Ltbrfl/fl mice. (f, g) N = 10 littermate control mice, 10 Cxcl12fl/fl mice and 9 Ltbrfl/fl mice per group; 2 independent experiments. h, GL7, CD138, B220 and CD21/35 staining of lymph nodes from Ltbrfl/fl mice on day 8 post VSV-immunization. Data is representative of 3 Ltbrfl/fl mice. (a, c, e, f, g) Mean and SEM are shown. P values as per one-way Anova with Tukey’s multiple comparisons test.
Extended Data Fig. 7 Cxcl12-proficient Cxcl13-Cre+ cells govern high-affinity humoral immunity to complex protein antigens and haptens.
a, An example of reconstructed clonal lineage trees based on B cell receptor sequencing for the quantification of maximum branch lengths. b, Quantification of the frequency of GL7+ CD19+ cells in draining lymph nodes of littermate control or Cxcl13-Cre/TdTom Cxcl12fl/fl (Cxcl12fl/fl) mice 12 day following NP-KLH immunization. N = 10 control mice and 11 Cxcl12fl/fl mice; 2 independent experiments. c, d, Quantification of total (c) or high-affinity (d) NP-specific serum IgG titers from control or Cxcl12fl/fl mice 12 days following NP-KLH immunization. N = 8 control mice and 11 Cxcl12fl/fl mice; 2 independent experiments. e, Quantification of the proportion of GC B cells harboring the high-affinity W33L mutation in control and Cxcl12fl/fl mice. (b–d) Mean and SEM are shown. P values as per two-tailed Mann-Whitney test. (e) Data is representative of two independent mice per condition; the number of clonotypes is indicated. P values as per two-tailed Wilcoxon test.
Extended Data Fig. 8 ScRNA-Seq of germinal center B cells from VSV immunized mice.
a, Sorting strategy for scRNA-seq of GC B cells. b, Top marker genes for each subset, and normalized expression scores (NES) of top GSEA pathways reflected by each subset’s marker genes. c, Median cell cycle scores. The middle line demarcates the median; box limits demarcate the upper and lower quartiles; whiskers depict the 1.5x interquartile range and points indicate outliers. (b, c) scRNA-seq data is representative of 4140 cells from littermate control mice and 4677 cells from Cxcl12fl/fl mice; 2 biological replicates over 2 independent experiments.
Extended Data Fig. 9 CXCL12 regulates the gene expression profile of B cells and positioning of TFH within the GC.
a, Pseudotime ordering of GC B cells. Violin plots indicate the density distribution of cells. b, Average gene expression of GC marker genes along inferred pseudotime for littermate control (black line) and Cxcl13-Cre/TdTom Cxcl12fl/fl (red line) mice. c, The relative abundance of cells expressing the indicated genes in control and Cxcl12fl/fl mice. d, Representative flow cytometry plots of the indicated markers in GL7+ B cells from littermate control or Cxcl12fl/fl mice, GL7− non-GC B cells and FMO controls. e, Quantification of the mean fluorescence intensity (MFI) of the indicated markers in CD86+ CXCR4− LZ and CD86− CXCR4+ DZ B cells from control (black boxes) or Cxcl12fl/fl (white boxes) mice. AKT: N = 5 mice per condition; BIM: N = 6 control and 7 Cxcl12fl/fl mice; Bcl6: N = 7 control and 9 Cxcl12fl/fl mice; all over 2 independent experiments. f, Representative flow cytometry plots and mean percentage and SEM of PD-1+ Bcl6+ CD4+ cells in immunized control and Cxcl12fl/fl mice. g, Enumeration of PD-1+ Bcl6+ CD4+ cells per lymph node (LN) in immunized control or Cxcl12fl/fl mice. N = 15 control and 13 Cxcl12fl/fl mice; 3 independent experiments. h, TdTom, PD-1 and CD4 staining in immunized Cxcl13-Cre/TdTom mice. The dotted line demarcates the GC perimeter, which is divided into four quadrants for manual PD-1+ cell enumeration. Scale bar, 100 μm. Images are representative of 2 mice. (a–c) scRNA-seq data is representative of 4140 cells from littermate control mice and 4677 cells from Cxcl12fl/fl mice; 2 biological replicates over 2 independent experiments. (c) Statistical significance calculated using the Pearson’s Chi-squared test with Bonferroni multiple comparisons test. (e, g) Mean and SEM are shown. (e) P values as per one-way ANOVA with Tukey’s multiple comparisons test. (g) P values as per two-tailed Mann-Whitney.
Supplementary information
Rights and permissions
About this article
Cite this article
Pikor, N.B., Mörbe, U., Lütge, M. et al. Remodeling of light and dark zone follicular dendritic cells governs germinal center responses. Nat Immunol 21, 649–659 (2020). https://doi.org/10.1038/s41590-020-0672-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-020-0672-y
This article is cited by
-
Bone morphogenic protein-4 availability in the cardiac microenvironment controls inflammation and fibrosis in autoimmune myocarditis
Nature Cardiovascular Research (2024)
-
PI16+ reticular cells in human palatine tonsils govern T cell activity in distinct subepithelial niches
Nature Immunology (2023)
-
Long-term retention of antigens in germinal centers is controlled by the spatial organization of the follicular dendritic cell network
Nature Immunology (2023)
-
Splenic stromal niches in homeostasis and immunity
Nature Reviews Immunology (2023)
-
Conserved stromal–immune cell circuits secure B cell homeostasis and function
Nature Immunology (2023)