Abstract
Understanding the mechanisms that modulate helper T lymphocyte functions is crucial to decipher normal and pathogenic immune responses in humans. To identify molecular determinants influencing the pathogenicity of T cells, we separated ex vivo-isolated primary human memory T lymphocytes on the basis of their ability to produce high levels of inflammatory cytokines. We found that the inflammatory, cytokine-producing phenotype of memory T lymphocytes was defined by a specific core gene signature and was mechanistically regulated by the constitutive activation of the NF-κB pathway and by the expression of the transcriptional repressor BHLHE40. BHLHE40 attenuated the expression of anti-inflammatory factors, including miR-146a, a negative regulator of NF-κB activation and ZC3H12D, an RNase of the Regnase-1 family able to degrade inflammatory transcripts. Our data reveal a molecular network regulating the proinflammatory phenotype of human memory T lymphocytes, with the potential to contribute to disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
All Nanostring, RNA-seq, ChIP-seq and ATAC-seq datasets are available in GEO (accession number GSE122946). Source data are provided for Figs. 4e and 5b. Other supporting raw data are available from the corresponding author upon request.
References
Newell, E. W., Sigal, N., Bendall, S. C., Nolan, G. P. & Davis, M. M. Cytometry by time-of-flight shows combinatorial cytokine expression and virus-specific cell niches within a continuum of CD8+ T cell phenotypes. Immunity 36, 142–152 (2012).
Helmstetter, C. et al. Individual T helper cells have a quantitative cytokine memory. Immunity 42, 108–122 (2015).
Cao, Y. et al. Functional inflammatory profiles distinguish myelin-reactive T cells from patients with multiple sclerosis. Sci. Transl. Med. 7, 287ra274 (2015).
Galli, E. et al. GM-CSF and CXCR4 define a T helper cell signature in multiple sclerosis. Nat. Med. 25, 1290–1300 (2019).
Hartmann, F. J. et al. Multiple sclerosis-associated IL2RA polymorphism controls GM-CSF production in human TH cells. Nat. Commun. 5, 5056 (2014).
Hirota, K. et al. Autoimmune Th17 cells induced synovial stromal and innate lymphoid cell secretion of the cytokine GM-CSF to initiate and augment autoimmune arthritis. Immunity 48, 1220–1232 (2018).
Behrens, F. et al. MOR103, a human monoclonal antibody to granulocyte–macrophage colony-stimulating factor, in the treatment of patients with moderate rheumatoid arthritis: results of a phase Ib/IIa randomised, double-blind, placebo-controlled, dose-escalation trial. Ann. Rheum. Dis. 74, 1058–1064 (2015).
Burmester, G. R. et al. Efficacy and safety of mavrilimumab in subjects with rheumatoid arthritis. Ann. Rheum. Dis. 72, 1445–1452 (2013).
de Vries, E. G. et al. Flare-up of rheumatoid arthritis during GM-CSF treatment after chemotherapy. Lancet 338, 517–518 (1991).
Codarri, L. et al. RORγT drives production of the cytokine GM-CSF in helper T cells, which is essential for the effector phase of autoimmune neuroinflammation. Nat. Immunol. 12, 560–567 (2011).
Sonderegger, I. et al. GM-CSF mediates autoimmunity by enhancing IL-6-dependent Th17 cell development and survival. J. Exp. Med. 205, 2281–2294 (2008).
McQualter, J. L. et al. Granulocyte–macrophage colony-stimulating factor: a new putative therapeutic target in multiple sclerosis. J. Exp. Med. 194, 873–882 (2001).
Campbell, I. K. et al. Protection from collagen-induced arthritis in granulocyte–macrophage colony-stimulating factor-deficient mice. J. Immunol. 161, 3639–3644 (1998).
Wawro, M., Kochan, J., Krzanik, S., Jura, J. & Kasza, A. Intact NYN/PIN-like domain is crucial for the degradation of inflammation-related transcripts by ZC3H12D. J. Cell Biochem. 118, 487–498 (2017).
Taganov, K. D., Boldin, M. P., Chang, K. J. & Baltimore, D. NF-κB-dependent induction of microRNA miR-146, an inhibitor targeted to signaling proteins of innate immune responses. Proc. Natl Acad. Sci. USA 103, 12481–12486 (2006).
McGeachy, M. J. et al. The interleukin 23 receptor is essential for the terminal differentiation of interleukin 17-producing effector T helper cells in vivo. Nat. Immunol. 10, 314–324 (2009).
Sakaguchi, S., Miyara, M., Costantino, C. M. & Hafler, D. A. FOXP3+ regulatory T cells in the human immune system. Nat. Rev. Immunol. 10, 490–500 (2010).
Ascherio, A., Munger, K. L. & Simon, K. C. Vitamin D and multiple sclerosis. Lancet Neurol. 9, 599–612 (2010).
Kongsbak, M., Levring, T. B., Geisler, C. & von Essen, M. R. The vitamin D receptor and T cell function. Front. Immunol. 4, 148 (2013).
Annunziato, F. et al. Phenotypic and functional features of human Th17 cells. J. Exp. Med. 204, 1849–1861 (2007).
Lin, C. C. et al. IL-1-induced Bhlhe40 identifies pathogenic T helper cells in a model of autoimmune neuroinflammation. J. Exp. Med. 213, 251–271 (2016).
Sun, H., Lu, B., Li, R. Q., Flavell, R. A. & Taneja, R. Defective T cell activation and autoimmune disorder in Stra13-deficient mice. Nat. Immunol. 2, 1040–1047 (2001).
Martinez-Llordella, M. et al. CD28-inducible transcription factor DEC1 is required for efficient autoreactive CD4+ T cell response. J. Exp. Med. 210, 1603–1619 (2013).
Lin, C. C. et al. Bhlhe40 controls cytokine production by T cells and is essential for pathogenicity in autoimmune neuroinflammation. Nat. Commun. 5, 3551 (2014).
Huynh, J. P. et al. Bhlhe40 is an essential repressor of IL-10 during Mycobacterium tuberculosis infection. J. Exp. Med. 215, 1823–1838 (2018).
Yu, F. et al. The transcription factor Bhlhe40 is a switch of inflammatory versus anti-inflammatory Th1 cell fate determination. J. Exp. Med. 215, 1813–1821 (2018).
Miyazaki, K. et al. The role of the basic helix-loop-helix transcription factor Dec1 in the regulatory T cells. J. Immunol. 185, 7330–7339 (2010).
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Geiger, R., Duhen, T., Lanzavecchia, A. & Sallusto, F. Human naive and memory CD4+ T cell repertoires specific for naturally processed antigens analyzed using libraries of amplified T cells. J. Exp. Med. 206, 1525–1534 (2009).
Mele, F. et al. ERK phosphorylation and miR-181a expression modulate activation of human memory TH17 cells. Nat. Commun. 6, 6431 (2015).
St-Pierre, B., Flock, G., Zacksenhaus, E. & Egan, S. E. Stra13 homodimers repress transcription through class B E-box elements. J. Biol. Chem. 277, 46544–46551 (2002).
Sun, H. & Taneja, R. Stra13 expression is associated with growth arrest and represses transcription through histone deacetylase (HDAC)-dependent and HDAC-independent mechanisms. Proc. Natl Acad. Sci. USA 97, 4058–4063 (2000).
Ivanova, A. V. et al. STRA13 interacts with STAT3 and modulates transcription of STAT3-dependent targets. J. Mol. Biol. 340, 641–653 (2004).
Li, Y. et al. DEC1 negatively regulates the expression of DEC2 through binding to the E-box in the proximal promoter. J. Biol. Chem. 278, 16899–16907 (2003).
Housley, W. J. et al. Genetic variants associated with autoimmunity drive NF-κB signaling and responses to inflammatory stimuli. Sci. Transl. Med. 7, 291ra293 (2015).
Monticelli, S., Aijo, T. & Trifari, S. Approaches to detect microRNA expression in T cell subsets and T cell differentiation. Methods Mol. Biol. 1514, 153–172 (2017).
Matsushita, K. et al. Zc3h12a is an RNase essential for controlling immune responses by regulating mRNA decay. Nature 458, 1185–1190 (2009).
Liang, J. et al. MCP-induced protein 1 deubiquitinates TRAF proteins and negatively regulates JNK and NF-κB signaling. J. Exp. Med. 207, 2959–2973 (2010).
Minagawa, K. et al. Post-transcriptional modulation of cytokine production in T cells for the regulation of excessive inflammation by TFL. J. Immunol. 192, 1512–1524 (2014).
Ferber, I. A. et al. Mice with a disrupted IFN-γ gene are susceptible to the induction of experimental autoimmune encephalomyelitis (EAE). J. Immunol. 156, 5–7 (1996).
Bettelli, E. et al. Loss of T-bet, but not STAT1, prevents the development of experimental autoimmune encephalomyelitis. J. Exp. Med. 200, 79–87 (2004).
Oestreich, K. J. & Weinmann, A. S. Master regulators or lineage-specifying? Changing views on CD4+ T cell transcription factors. Nat. Rev. Immunol. 12, 799–804 (2012).
Kanda, M. et al. Transcriptional regulator Bhlhe40 works as a cofactor of T-bet in the regulation of IFN-γ production in iNKT cells. Proc. Natl Acad. Sci. USA 113, e3394–e3402 (2016).
Zhang, L. et al. Lineage tracking reveals dynamic relationships of T cells in colorectal cancer. Nature 564, 268–272 (2018).
Hosokawa, H. et al. Transcription factor PU.1 represses and activates gene expression in early T cells by redirecting partner transcription factor binding. Immunity 48, 1119–1134 (2018).
Oestreich, K. J. & Weinmann, A. S. T-bet employs diverse regulatory mechanisms to repress transcription. Trends Immunol. 33, 78–83 (2012).
Curina, A. et al. High constitutive activity of a broad panel of housekeeping and tissue-specific cis-regulatory elements depends on a subset of ETS proteins. Genes Dev. 31, 399–412 (2017).
Balestrieri, C. et al. Co-optation of tandem DNA repeats for the maintenance of mesenchymal identity. Cell 173, 1150–1164 (2018).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Ross-Innes, C. S. et al. Differential oestrogen receptor binding is associated with clinical outcome in breast cancer. Nature 481, 389–393 (2012).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
McLean, C. Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010).
Diaferia, G. R. et al. Dissection of transcriptional and cis-regulatory control of differentiation in human pancreatic cancer. EMBO J. 35, 595–617 (2016).
Zambelli, F., Pesole, G. & Pavesi, G. Pscan: finding over-represented transcription factor binding site motifs in sequences from co-regulated or co-expressed genes. Nucleic Acids Res. 37, W247–W252 (2009).
Zambelli, F., Pesole, G. & Pavesi, G. PscanChIP: finding over-represented transcription factor-binding site motifs and their correlations in sequences from ChIP-Seq experiments. Nucleic Acids Res. 41, W535–W543 (2013).
Mahony, S. & Benos, P. V. STAMP: a web tool for exploring DNA-binding motif similarities. Nucleic Acids Res. 35, W253–W258 (2007).
Letunic, I. & Bork, P. Interactive tree of life (iTOL) v3: an online tool for the display and annotation of phylogenetic and other trees. Nucleic Acids Res. 44, W242–W245 (2016).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Delaleu, N., Nguyen, C. Q., Tekle, K. M., Jonsson, R. & Peck, A. B. Transcriptional landscapes of emerging autoimmunity: transient aberrations in the targeted tissue’s extracellular milieu precede immune responses in Sjogren’s syndrome. Arthritis Res. Ther. 15, R174 (2013).
Subramanian, A. et al. Gene-set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Su, G., Morris, J. H., Demchak, B. & Bader, G. D. Biological network exploration with Cytoscape 3. Curr. Protoc. Bioinformatics 47, 11–24 (2014).
Merico, D., Isserlin, R., Stueker, O., Emili, A. & Bader, G. D. Enrichment map: a network-based method for gene-set enrichment visualization and interpretation. PloS One 5, e13984 (2010).
Lindner, S. E. et al. The miR-15 family reinforces the transition from proliferation to differentiation in pre-B cells. EMBO Rep. 18, 1604–1617 (2017).
Acknowledgements
The authors thank D. Jarossay, S. Notarbartolo and F. Mele for invaluable technical support and input; M. Perez for help with Fig. 1; L. Vincenzetti for pilot CRISPR-Cas9 experiments and E. Džafo for help with experiments involving Regnase-4. This work was supported by the Swiss National Science Foundation (grant number 156875 and 175569), the NCCR ‘RNA & Disease’, the Swiss MS Society, the Ceresio Foundation, the Vontobel Stiftung and the Kurt und Senta Herrmann Stiftung (all to S. Monticelli). S. Montagner was supported by a Marie Heim-Vögtlin postdoctoral fellowship (number 164489). This work was also partially supported by the Italian Ministry of Health with Ricerca Corrente and 5×1000 funds (to G.N.).
Author information
Authors and Affiliations
Contributions
S.E., N.B., C.L., S. Montagner and M.C. designed and performed experiments and analyzed data; S.P., C.B. and G.N. performed and analyzed all the sequencing experiments; N.D. analyzed data; S. Monticelli oversaw the project and analyzed data. S. Monticelli and G.N. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
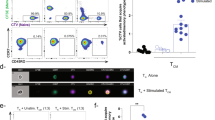
Extended Data Fig. 1 Sorting scheme and principal component analysis.
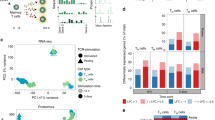
a, Sorting scheme for GM-CSF secretion assay. TEM cells were separated by sorting and further divided by secretion assay and sorting. b, Principal component analysis of RNA-seq data. Cells from nine individual donors were separated by GM-CSF secretion assay and pooled in three pools of three donors each prior to RNA-seq. This analysis showed robust results, with differences due to sample-to-sample variability accounting for only 15.45% (PC#2) of the variance, while the majority of the observed differences were due to the phenotype of the cells (65.03%, GM-CSFpos vs. neg, PC#1).
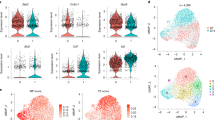
Extended Data Fig. 2 Expression of specific markers in T cell subsets.
a, Cytokine expression in different T cell subsets. Each indicated cytokine (IL-22, IL-17A and IFN-γ) is expressed with or without concomitant co-expression of GM-CSF. Also, the IL-22+ cells are not simply a subset of the larger GM-CSF+ pool of cells. Mean ± SD. Each dot represents one donor (n = 5). b, Expression of IL23R. Expression of the IL23R gene in GM-CSF+ and GM-CSF− cells, as determined by RNA-seq (n = 9). FDR = False discovery rate calculated for RNA-seq samples. c, FOXP3 expression. Comparison of FOXP3 expression in Treg cells (CD4+CD25highCD127low), GM-CSF+ and GM-CSF− cells, as assessed by intracellular staining. Representative of n = 2 experiments.
Extended Data Fig. 3 Differences in gene expression patterns between GM-CSF+ and GM-CSF− TEM cells.
Systematic delineation of coordinated changes in gene expression within specific gene sets (that is, pathways) was achieved by applying the data analysis and data visualization pipeline described in the Methods. 119 gene sets yielded significant enrichment in GM-CSF+ vs. GM-CSF− cells (false-discovery-rate (FDR)-q < 0.01, nom. p-value of < 0.005 and TAGS > 50) whereas none were found depleted. Defining the network’s organization is the degree of overlap in leading edge-genes (genes contributing to each gene set’s significant enrichment) two gene sets (nodes) linked by a connection (edge) share (threshold connectivity 0.05). Node-size corresponds to FDR-q value threshold passed by the gene set (small < 0.01, medium < 0.001, large < 0.0001). Node color denotes MCL-cluster membership that subsequently would be summarized within 5 biological themes that is color-coded network areas. Their FDR-q value distribution is displayed in Fig. 1f.
Extended Data Fig. 4 RNA-seq, Nanostring, ATAC-seq and ChIP-seq analyses.
a, Comparison of RNA-seq and Nanostring data. Concordant results of gene expression profiling obtained by RNA-seq of TEM cells and Nanostring profiling of TCM cells. b, Expression of BHLHE40 in different T cell subsets. Expression of BHLHE40 was determined by qRT-PCR in different T cell subsets freshly sorted from peripheral blood. Each dot represents one donor (n = 6). Mean ± SD; paired t-test, two-tailed. c, ATAC-seq and ChIP-seq analyses. Representative snapshots of ChIP-seq and ATAC-seq tracks for selected genomic loci.
Extended Data Fig. 5 Optimization of CRISP-Cas9-mediated deletion and screening.
a, As a proof-of-principle and test of efficiency, Jurkat T cells were transfected using the Neon transfection system with ribonucleoparticles of Cas9 and one gRNA against the TCRα chain. After three days the cells were stained for surface TCR expression, showing a loss of expression in 62% of the cells (green). Grey: mock transfected control cells. Representative of at least n = 3 independent experiments. b, Schematic representation of the BHLHE40 locus with indicated the location of gRNAs and primers used for screening of the clones. c, Optimization of screening procedure by mismatch cleavage assay. After transfection with two gRNAs against BHLHE40, Jurkat cells were single cell-cloned by seeding in a 384-well plate in 20% FBS. Genomic DNA was extracted from 10 clones and from the entire population (no cloning) and negative controls. The first ‘long’ PCR showed already the presence of indels in most clones (left panel) and even at the level of whole population. This was further confirmed by denaturing, reannealing and T7 endonuclease digestion (middle panel). The presence of a homozygous deletion was further confirmed using a ‘short’ PCR that cannot provide an amplification product if the region between the two gRNAs is deleted. The highlighted Clone 7 (black box) is one of several clones in which the deletion was most likely identical on both alleles, leading to a shorter PCR product that cannot be digested by the T7 endonuclease because of the absence of mismatches. Representative of at least n = 2 independent transfections and cloning procedures. d, The genomic DNA from Clone 7 was sequenced in the BHLHE40 locus region, revealing a deletion of 196 nucleotides involving part of exon 1 and exon 2, as well as the intron. e, Schematic representation of the ZC3H12D locus with indicated the location of gRNAs and primers used for screening of the clones in Fig. 8f.
Extended Data Fig. 6 Analysis of primary human T cell clones transfected with Cas9 ribonucleoparticles directed against the BHLHE40 gene.
Primary human memory T lymphocytes were transfected with ribonucleoparticles of Cas9 and gRNAs against exon 1/2 and exon 5 of BHLHE40. After single cell cloning and expansion, individual clones were tested for the presence of the deletion by PCR. Median; Mann-Whitney test. Each dot represents one clone (n = 77 for KO and n = 90 for control clones).
Extended Data Fig. 7 MiRNA analysis.
a, Transduction with a miR-146a sponge of primary human T lymphocytes leads to increased NF-kB p65 phosphorylation. Primary memory T lymphocytes were transduced with a lentivirus expressing a miR-146a sponge and GFP as an independent reporter of the efficiency of transduction (schematic representation on top); a few days after transduction cells were stimulated with PMA and ionomycin for 5 min and levels of p65 phosphorylation were determined by intracellular staining. Cells were gated as GFPneg (non-expressing the sponge) and GFPpos (expressing the sponge) and compared also to untransduced cells. Representative of n = 2 experiments. b, Same as (c) except that the results (expressed as mean fluorescence intensity, MFI) of two independent experiments in which cells were stimulated for the indicated times are shown. c, Effect of BHLHE40 expression on miR-181a in Jurkat T cells. Jurkat T cells were transduced with a BHLHE40-expressing lentivirus. After selection and expansion of the transduced cells, expression of miR-181a was measured by qRT-PCR. Each dot represents one independent experiment (n = 4). Mean ± SD; paired t-test, two-tailed. d, Phylogenetic analysis of the top 50 bHLH matrices recovered from the BHLHE40 ChIP-seq data.
Extended Data Fig. 8 Schematic diagram describing the regulatory module identified in this study.
Using GM-CSF secretion as a proxy for an inflammatory phenotype of primary human memory TH lymphocytes, we identified a regulatory module that influences the activity of these pro-inflammatory and potentially pathogenic cells. In GM-CSF− cells, mechanisms are in place to actively repress the expression of inflammatory cytokines, including high levels of miR-146a expression (a negative regulator of NF-κB activation), and relatively higher expression of ZC3H12D, an RNase enzyme involved in the negative regulation of cytokine expression. Conversely, in GM-CSF+ cells, the expression of the transcriptional repressor BHLHE40 leads to the direct downregulation of ZC3H12D expression, and indirectly affects also miR-146a. This in turn allows full-blown NF-κB activation and expression of inflammatory cytokines.
Supplementary information
Source data
Source Data Fig. 4
Uncropped images of western blots in Fig. 4. After transfer, the membrane was cut using the marker as a reference and the two parts of the same membrane were blotted with the indicated antibody. The boxes represent the parts of the gel shown in the figure.
Source Data Fig. 5
Uncropped images of western blots in Fig. 4. After transfer, the membrane was cut using the marker as a reference and the two parts of the same membrane were blotted with the indicated antibody. For each experiment, the same lysate was loaded twice on the same gel, the membrane was cut and part of the membrane was blotted with an anti-phosphorylated-p65 antibody, while the other part was blotted with a total anti-p65 antibody. The boxes represent the parts of the gels shown in the figure.
Rights and permissions
About this article
Cite this article
Emming, S., Bianchi, N., Polletti, S. et al. A molecular network regulating the proinflammatory phenotype of human memory T lymphocytes. Nat Immunol 21, 388–399 (2020). https://doi.org/10.1038/s41590-020-0622-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-020-0622-8
This article is cited by
-
Cholinergic control of Th17 cell pathogenicity in experimental autoimmune encephalomyelitis
Cell Death & Differentiation (2023)
-
NF-κB subunits direct kinetically distinct transcriptional cascades in antigen receptor-activated B cells
Nature Immunology (2023)
-
Steady-state memory-phenotype conventional CD4+ T cells exacerbate autoimmune neuroinflammation in a bystander manner via the Bhlhe40/GM-CSF axis
Experimental & Molecular Medicine (2023)
-
MHBSt167 induced autophagy promote cell proliferation and EMT by activating the immune response in L02 cells
Virology Journal (2022)
-
Identification of a ZC3H12D-regulated competing endogenous RNA network for prognosis of lung adenocarcinoma at single-cell level
BMC Cancer (2022)