Abstract
Clinical manifestations of COVID-19 caused by the new coronavirus SARS-CoV-2 are associated with age1,2. Adults develop respiratory symptoms, which can progress to acute respiratory distress syndrome (ARDS) in the most severe form, while children are largely spared from respiratory illness but can develop a life-threatening multisystem inflammatory syndrome (MIS-C)3,4,5. Here, we show distinct antibody responses in children and adults after SARS-CoV-2 infection. Adult COVID-19 cohorts had anti-spike (S) IgG, IgM and IgA antibodies, as well as anti-nucleocapsid (N) IgG antibody, while children with and without MIS-C had reduced breadth of anti-SARS-CoV-2-specific antibodies, predominantly generating IgG antibodies specific for the S protein but not the N protein. Moreover, children with and without MIS-C had reduced neutralizing activity as compared to both adult COVID-19 cohorts, indicating a reduced protective serological response. These results suggest a distinct infection course and immune response in children independent of whether they develop MIS-C, with implications for developing age-targeted strategies for testing and protecting the population.
Similar content being viewed by others
Main
The clinical manifestations of SARS-CoV-2 infection in children are distinct from adults. Children with COVID-19 rarely exhibit severe respiratory symptoms and often remain asymptomatic2, whereas adults experience respiratory symptoms of varying severity; older adults and those with comorbidities such as hypertension and diabetes have substantially higher risks of developing COVID-19-associated ARDS with high mortality2,6. In children, a rare but severe clinical manifestation of SARS-CoV-2 infection designated MIS-C, exhibits similarities to Kawasaki disease in certain inflammatory features and cardiovascular involvement while generally lacking severe respiratory symptoms3,4,5. The nature of the immune response to SARS-CoV-2 in children with different clinical manifestations ranging from asymptomatic to MIS-C relative to the more common respiratory manifestations of COVID-19 in adults is unclear.
The generation of virus-specific antibodies that neutralize or block infectivity is the most consistent correlate of protective immunity for multiple infections and vaccines7,8. Antibodies specific for the major SARS-CoV-2 antigens, including the S protein which binds the cellular receptor for viral entry and the N protein necessary for viral replication, have been detected in actively infected patients and in patients with mild disease who recovered9,10,11,12. Anti-S antibodies, in particular, can exhibit potent neutralizing activity and are currently being pursued as a therapeutic option for infusion into patients during severe disease and for targeted generation in vaccines13,14,15. Defining the nature of the antibody response to SARS-CoV-2 infection as a function of age and clinical syndrome can provide essential insights for improved screening and targeted protection for the global population that continues to suffer from this relentless pandemic.
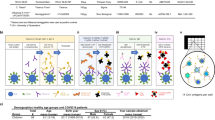
In this study, we investigated the specificity and functionality of the antibody response and its protective capacity in adult and pediatric patients seen at NewYork-Presbyterian/Columbia University Irving Medical Center (NYP/CUIMC) hospital and the Morgan Stanley Children’s Hospital of New York (MSCHONY) during the height of the pandemic in New York City from March to June 20203,13,16,17. We present 4 patient cohorts comprising a total of 79 individuals, including adults recruited as convalescent plasma donors (CPDs) who recovered from mild COVID-19 respiratory disease without requiring hospitalization (CPD, n = 19), adults hospitalized with severe COVID-19 ARDS (COVID-ARDS, n = 13) and 2 pediatric cohorts including children hospitalized with MIS-C (MIS-C, n = 16) and children who were infected with SARS-CoV-2 but did not develop MIS-C (pediatric non-MIS-C, n = 31) (clinical characteristics are shown in Table 1). The adult cohorts represented a broad age range (19–84 years) while members of the pediatric cohorts were younger (3–18 years) (Table 1). Individuals were diagnosed as infected with SARS-CoV-2 based on a history of symptoms, PCR+ test for virus and/or serology (Table 1). While comorbidities were rare among pediatric individuals, they were frequently present in adults with COVID-ARDS (Supplementary Table 1). Samples from COVID-ARDS and MIS-C patients were obtained within 24–36 h of being admitted or intubated for respiratory failure, largely before the initiation of therapeutic interventions (Supplementary Table 1). Samples from pediatric non-MIS-C individuals were obtained during phlebotomy for various clinical reasons, including routine screening for hospital admission and medical procedures (Supplementary Table 2), with 48% having experienced no COVID-like symptoms and designated as asymptomatic. Both MIS-C and COVID-ARDS individuals exhibited markers of systemic inflammation including highly elevated concentrations of interleukin 6 (IL-6) and C-reactive protein (CRP), while ferritin and lactate dehydrogenase (LDH) were significantly increased in COVID-ARDS compared to MIS-C individuals (Table 1). Only 2 pediatric individuals developed respiratory failure and ARDS (Table 1; 1 with MIS-C and 1 non-MIS-C), indicating distinct inflammatory responses and clinical manifestations between children and adults in response to infection.
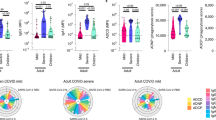
We quantitated SARS-CoV-2 specific antibodies for each cohort in terms of specificity and antibody class, including IgM generated initially in a primary response and IgG and IgA classes prominent in serum and secretions, respectively. Anti-S antibodies were present as IgG (Fig. 1a), IgM (Fig. 1b) and IgA (Fig. 1c) classes in adult COVID-ARDS and CPD donors, with significantly higher concentration in COVID-ARDS patients for all classes (Fig. 1a–c). By contrast, anti-S antibody titers and isotype predominance in both pediatric cohorts (MIS-C and non-MIS-C) were similar to each other and to the adult CPD subjects—showing predominant anti-S IgG (Fig. 1a), low titers of anti-S IgM (Fig. 1b) (similar to negative control pre-pandemic plasma), and variable titers of anti-S IgA antibodies (Fig. 1c). We further assessed the specificity of anti-S IgG for SARS-CoV-2 S protein compared to other coronavirus strains using a cell-based ELISA (Methods). Plasma IgG from subject samples but not pre-pandemic control samples bound SARS-CoV-2 S protein and the common circulating D614G S protein variant18, but did not significantly bind S protein from SARS-CoV-1 or Middle East respiratory syndrome coronaviruses (Extended Data Fig. 1), establishing the specificity of the anti-S IgG response for SARS-CoV-2 in all cohorts. However, the abundance of IgG antibodies specific for the SARS-CoV-2 nucleocapsid (N) protein, which complexes with viral RNA and is involved in viral replication19 was significantly lower in both pediatric cohorts compared to the two adult cohorts (Fig. 1d). The low amounts of anti-N IgG were similar in children with and without MIS-C, and the higher anti-N IgG titers in adults were similar in the CPD and COVID-ARDS cohort, suggesting that generation of anti-N antibody is age- but not symptom-dependent.
a–d, Levels of antibodies to SARS-CoV-2 S and N proteins were measured using serial dilutions of patient plasma in an indirect ELISA assay to detect anti-S IgG (a), anti-S IgM (b), anti-S IgA (c) and anti-N IgG (d). The absorbance sum across 6 serial 1:4 plasma dilutions from adult CPDs (open black squares, n = 19), adult patients with COVID-19-induced ARDS (COVID-ARDS, closed red squares, n = 13), pediatric patients with a history of SARS-CoV-2 infection but not MIS-C (non-MIS-C, open blue circles, n = 31), patients with MIS-C (closed green circles, n = 16) and control plasma from pre-pandemic donors (negative control, gray triangles, n = 10) is shown. The black bar indicates the median + interquartile range (IQR). P values were calculated by one-way ANOVA with Šidák’s multiple comparisons test. Anti-S IgG (a): CPD versus COVID-ARDS: P = 1.32 × 10−4; CPD versus pediatric non-MIS-C: P = 0.59; COVID-ARDS versus MIS-C: P = 8.53 × 10−6; pediatric non-MIS-C versus MIS-C: P = 0.24. Anti-S IgM (b): CPD versus COVID-ARDS: P = 6.93 × 10−5; CPD versus pediatric non-MIS-C: P = 0.33; COVID-ARDS versus MIS-C: P = 2.54 × 10−6; pediatric non-MIS-C versus MIS-C: P = 0.99. Anti-S IgA (c): CPD versus COVID-ARDS: P = 3.82 × 10−7; CPD versus pediatric non-MIS-C: P = 0.08, COVID-ARDS versus MIS-C: P = 9.06 × 10−7; pediatric non-MIS-C versus MIS-C: P = 0.11. Anti-N IgG (d): CPD versus COVID-ARDS: P = 0.93; CPD versus pediatric non-MIS-C: P = 3.31 × 10−5; COVID-ARDS versus MIS-C: P = 3.88 × 10−5; pediatric non-MIS-C versus MIS-C: P = 0.99. Significance is indicated as ***P < 0.001 or P > 0.05 (not significant (NS)). e,f, For anti-S IgG (e) and anti-N IgG (f), the antibody levels of individuals are also plotted against patient age in the adult (left) and pediatric cohorts (right) with the best fit lines and P values calculated using simple linear regression. Anti-S IgG versus age (pediatric non-MIS-C: R2 = 0.23, slope = −0.077, y-intercept = 2.70). Anti-N IgG versus age (CPD: R2 = 0.34, slope = 0.023, y-intercept = 0.12).
The potential effects of age and time post-symptom onset (that is, disease course) on the differential antibody abundance for each cohort were examined. While there was no significant correlation between anti-S IgG and age among adults and the pediatric MIS-C cohort, a modest but significant negative correlation between age and anti-S IgG titers was observed in the pediatric non-MIS-C cohort (Fig. 1e, right). Moreover, there was a significant correlation of anti-N IgG titers with age in the CPD group with younger adults having lower anti-N titers than older adults, while both pediatric groups had low anti-N titers across all ages (Fig. 1f). Analysis of antibody abundance as a function of time post-symptom onset revealed a significant correlation between anti-S IgG titers and increased time post-symptom for both pediatric groups and adult COVID-ARDS group, which is suggestive of an evolving response over time (Fig. 2a). No correlation with symptom onset and anti-S IgM was observed (Fig. 2b). These results show that the anti-SARS-CoV-2 antibody response generated in children is predominantly anti-S IgG antibodies independent of clinical syndrome. By contrast, adults generate broader antibody responses to infection in terms of isotypes and specificities and exhibit increased magnitude and breadth of the anti-S antibody response with more severe disease.
a,b, Levels of anti-S IgG (a) and anti-S IgM (b) were plotted against the time post-symptom onset for those individuals who were symptomatic either with COVID-19 or MIS-C. The adult groups (CPD, open black squares, n = 19 and COVID-ARDS, closed red squares, n = 13) are plotted on the left and the pediatric groups (MIS-C, closed green circles, n = 16 and non-MIS-C, open blue circles, n = 16) are plotted on the right with the best fit line and P value reported to 4 decimal places, calculated using simple linear regression. Anti-S IgG versus time post-symptom onset (COVID-ARDS: R2 = 0.39, slope = 0.11, y-intercept = 1.59; MIS-C: R2 = 0.25, slope = 0.055, y-intercept = 1.87; pediatric non-MIS-C: R2 = 0.30, slope = 0.021, y-intercept = 1.29).
The functional capacity of antibodies to provide protection correlates with their neutralizing activity in blocking virus infection. We developed a cell-based pseudovirus assay based on a system reported previously20,21 where multicycle infection of red fluorescent protein (RFP)-expressing vesicular stomatitis virus (VSV) pseudotyped with SARS-CoV-2 S protein is measured in the presence of serially diluted plasma samples (Methods). We validated this assay by comparing the neutralizing activity of plasma samples tested in the pseudovirus assay to the activity measured in the live virus microneutralization assay based on inhibition of cytopathic effect22 and found a direct correlation in neutralizing activity calculated from the pseudovirus and live virus assay over a wide range of neutralizing activity (Fig. 3a).
a, Plasma neutralizing activity in the pseudovirus assay was correlated with the end point titers in a live virus microneutralization assay based on the inhibition of the cytopathic effect (n = 13; Methods). b, The neutralizing activity for SARS-CoV-2-specific antibodies was determined using the pseudovirus assay (Methods). Neutralizing activity is shown from adult CPDs (open black squares, n = 19), adult patients with COVID-19-induced ARDS (COVID-ARDS, closed red squares, n = 13), pediatric patients with a history of SARS-CoV-2 infection but not MIS-C (non-MIS-C, open blue circles, n = 31), patients with MIS-C (closed green circles, n = 16) and control plasma from pre-pandemic donors (negative control, gray triangles, n = 10). The black bar indicates the median + IQR. The P values were calculated by one-way ANOVA with Šidák’s multiple comparisons test (CPD versus COVID-ARDS: P = 0.019; CPD versus pediatric non-MIS-C: P = 0.0031; COVID-ARDS versus MIS-C: P = 3.35 × 10−6; pediatric non-MIS-C versus MIS-C: P = 1.0). Significance is indicated as *P < 0.05, **P < 0.01, ***P < 0.001 or P > 0.05 (NS). c, The percentage inhibition values of S protein–mediated pseudoviral replication plotted against the plasma dilution factors for all individuals in each group are shown. d,e, Neutralizing activity was plotted against anti-S IgG levels (d) and patient age (e) in the adult (left) and pediatric (right) cohorts. The best fit lines and P values (reported to four decimal places) were calculated using simple linear regression. Neutralizing activity versus anti-S IgG (CPD: R2 = 0.29, slope = 0.36, y-intercept = 0.88; MIS-C: R2 = 0.51, slope = 0.38, y-intercept = 0.43; pediatric non-MIS-C: R2 = 0.21, slope = 0.22, y-intercept = 0.86). Neutralizing activity versus age (pediatric non-MIS-C: R2 = 0.20, slope = −0.034, y-intercept = 1.64).
Neutralizing activity as measured by the pseudovirus assay showed differences between the four cohorts associated with age group and/or clinical severity. The pediatric MIS-C and non-MIS-C groups both exhibited significantly lower neutralizing activity than the adult CPD and COVID-ARDS groups, while plasma from COVID-ARDS patients showed the highest neutralizing potency of the four groups across the dilution series (Fig. 3b,c). No differences were observed in neutralizing activity in the MIS-C compared to the pediatric non-MIS-C group (Fig. 3b,c). Only a small fraction of antibodies raised against viral antigens will have neutralizing activity against the virus, which correlates with protective capacity23. By linear regression, there was significant correlation between the abundance of anti-S IgG and neutralizing activity in the CPD, MIS-C and pediatric non-MIS-C groups, albeit with a significantly lower elevation and y-intercept for the MIS-C group relative to the COVID-ARDS and CPD groups (Fig. 3d). Together, these results establish a significant quantitative difference in neutralizing activity of anti-SARS-CoV-2 antibodies between pediatric and adult groups.
We examined the potential effects of age and disease course on neutralizing activity in the different groups. There was no correlation between neutralizing activity and patient age in either adult group (Fig. 3d). However, there was a significant decline of neutralizing activity with patient age in the pediatric non-MIS-C group (Fig. 3e, right) similar to the decrease in anti-S IgG abundance with age observed during the teenage years (Fig. 1e, right). Neutralizing activity within each group did not correlate with time post-symptom onset except in the severely ill COVID-ARDS group (Fig. 4a). Moreover, MIS-C patients also maintained the same titers of anti-S IgG and neutralizing activity 2–4 weeks after hospital discharge based on paired analysis of the follow-up compared to the retested primary sample in 10 out of 16 (62.5%) patients (Fig. 4b). Together, these results indicate that lower magnitude of functional antibody responses in pediatric SARS-CoV-2 infection compared to adults is age-associated and not related to infection course.
a, The levels of neutralization activity were plotted against the time post-symptom onset for those individuals who were symptomatic either for COVID-19 or MIS-C. The adult groups (CPD, open black squares, n = 19 and COVID-ARDS, closed red squares, n = 13) are plotted on the left and the pediatric groups (MIS-C, closed green circles, n = 16 and non-MIS-C, open blue circles, n = 16) are plotted on the right with the best fit line and P value calculated using simple linear regression (COVID-ARDS: R2 = 0.36, slope = 0.040, y-intercept = 1.34). b, The anti-S IgG levels (left) and neutralizing activity (right) of MIS-C individuals (n = 10) during the acute phase of illness and at a follow-up visit 2–4 weeks after hospital discharge. The P values were calculated using a two-way paired t-test.
To better define how SARS-CoV-2 antibody responses are related to age and clinical syndrome, we performed multivariable linear regression analysis to control for the effects of demographic and clinical covariates. Consistent with the grouped analysis (Figs. 1a–d and 3b), analysis of all pediatric and adult data showed that the pediatric age group was a significant predictor of lower SARS-CoV-2 neutralizing activity, anti-S IgM and anti-N IgG, and that these relationships were independent of time post-symptom onset, clinical syndrome or sex (Supplementary Table 3). In addition, ARDS was a significant independent predictor of higher SARS-CoV-2 neutralizing activity, anti-S IgG and anti-S IgM (Supplementary Table 3). Within the subgroup of pediatric individuals, age was a significant independent predictor of SARS-CoV-2 neutralizing activity (Supplementary Table 3), which is consistent with the pairwise analysis (Fig. 3e, right). These results show that the observed relationships of age and clinical syndrome with SARS-CoV-2 antibody responses are independent of potentially confounding factors, including being male.
Together, our results show quantitative and qualitative differences in the anti-SARS-CoV-2-specific antibody response across the spectrum of infection in children compared to adults. Children exhibited a SARS-CoV-2-specific antibody response that was largely limited to IgG anti-S antibodies with the lowest overall level of neutralizing activity compared to adult COVID-19 cohorts. In addition, children with different disease severities (that is, with or without MIS-C) exhibited similar antibody profiles, while in the adult cohorts, those with the most severe disease (ARDS) had higher abundance, breadth and neutralizing activity of anti-SARS-CoV-2 antibodies compared to adults who recovered from mild disease. While there was an association with increased amounts of anti-S IgG and time post-symptom onset, age remained the major factor distinguishing antibody profiles. Additionally, the durable responses seen in follow-up samples from MIS-C individuals provide evidence for relative stability of antibody abundance over a period of weeks. These findings suggest distinct primary SARS-CoV-2 infection courses and immune responses in children and adults.
Optimal protection to viral respiratory tract infections is mediated by virus-specific immunological memory developed during previous exposures24. The majority of primary exposures, especially to viral respiratory pathogens that are ubiquitous in the population, occur during infancy and childhood and virus-specific memory is established by adult life25,26. Consequently, it is largely unknown how primary immune responses to viral pathogens may differ between children and adults. The sudden and widespread emergence of SARS-CoV-2 as a new pathogen enables the study of primary immune responses across all ages. The reduced respiratory symptoms and low incidence of ARDS in the pediatric population2 suggest a distinct infection course, possibly due to lower expression of the viral receptor (angiotensin-converting enzyme 2 (ACE2)) in pediatric airway epithelial cells27 or a more robust innate immune response in children28,29,30. A milder infection course in the pediatric groups is further consistent with the lower abundance of anti-N-specific antibodies identified in this study since release of N proteins requires lysis of virally infected cells. The age association of anti-N antibodies in the adult CPD group is consistent with the age-associated risk for more severe and prolonged disease from SARS-CoV-2 infection. While current platforms to determine previous infection with SARS-CoV-2 rely heavily on the detection of anti-N IgG, our results suggest that these testing platforms may have decreased sensitivity for assessing previous infections among the pediatric population.
The reduced functional antibody response in children compared to adults could also be due to efficacious immune-mediated viral clearance resulting in fewer respiratory symptoms and severe illness. The presence of SARS-CoV-2-specific T cells in the peripheral blood of recovered and COVID-ARDS adults has been demonstrated in multiple cohorts31,32,33, although the protective capacity of these T cells is unclear. The pediatric T cell response to SARS-CoV-2 requires investigation but may exceed the adult responses due to an increased number of naïve T cells available to respond to new pathogens34 or more recently acquired T cell memory to related coronavirus strains35 due to children experiencing more respiratory illnesses. The IgG predominance in the majority of children examined in this study is consistent with preexisting immunological memory. Interestingly, less severe manifestations of COVID-19 have been associated with a more coordinated adaptive immune responses in adults36, suggesting that the quality and quantity of the immune response is important for protection from severe disease. These are important future areas of investigation for understanding the immune response to SARS-CoV-2 infection in children.
The similar antibody profiles in children with and without MIS-C suggest that the adaptive immune response per se is not associated with MIS-C pathogenesis. However, reduced neutralizing activity may predispose children to develop low-level, persistent infection in other sites resulting in MIS-C. Children can present with gastrointestinal symptoms rather than respiratory illness and demonstrate prolonged fecal shedding of the virus37. Alternately, the presence of non-neutralizing anti-S antibodies could lead to antibody-dependent enhancement of infection, which is known to occur in viral infections including SARS-CoV-1 (ref. 38). Additionally, autoreactive antibodies recently identified in children with MIS-C may promote aberrant immune responses leading to systemic inflammation29,30. Further studies delineating the differences in adult and pediatric immune responses to SARS-CoV-2 are warranted to define how protection or pathology is mediated in response to this pathogen. In summary, our results suggest a distinct infection course and immune response in children independent of whether they develop MIS-C, with implications for developing age-targeted strategies for testing and protecting the population.
Methods
Study cohorts
We recruited a total of 79 individuals from MSCHONY and NYP/CUIMC who represented distinct clinical manifestations of SARS-CoV-2 infection and different age groups divided into four cohorts: (1) individuals (n = 19) donating blood as part of our institution’s convalescent plasma trial (CPDs) after a history of recent illness consistent with COVID-19 but not requiring hospitalization and subsequently identified as positive for anti-SARS-CoV-2 antibodies; (2) patients with severe COVID-19 and ARDS (n = 13) who tested positive for SARS-CoV-2 by PCR from nasopharyngeal swabs; (3) pediatric patients with MIS-C (n = 16) and confirmed SARS-CoV-2 antibody-positive serology; and (4) pediatric patients without MIS-C (n = 31) receiving medical attention at NYP/CUIMC and confirmed to have active or previous SARS-CoV-2 infection by PCR from nasopharyngeal swabs or antibody-positive serology. ARDS was defined by clinical consensus criteria, including infiltrates on chest radiograph and a PaO2/FiO2 ratio of less than 300 or equivalent pediatric criteria39,40. MIS-C was defined using the Centers for Disease Control and Prevention definition: <21 years of age; fever >38 °C for >24 h; laboratory evidence of inflammation; hospital admission; multisystem involvement; no alternative plausible diagnosis; and positive SARS-Cov-2 serology41. Sequential organ failure assessment (SOFA) scores were calculated on all hospitalized individuals using previously validated adult and pediatric score tools to provide additional clinical insight into patient disease severity42,43,44. This study was approved by the institutional review board at CUIMC. Written consent was obtained from CPD individuals. Due to the limitations placed on direct contact with infected individuals and a need to conserve personal protective equipment, verbal informed consent was obtained from surrogates of critically ill COVID-ARDS patients and verbal parental consent was obtained for MIS-C individuals. Biospecimens and data from non-MIS-C pediatric patients were obtained from the Columbia University Biobank.
Sample processing
Blood samples were obtained at the time of outpatient donation for CPD individuals, at the time of admission for MIS-C individuals, during clinical care for pediatric non-MIS-C individuals and after diagnosis of ARDS for COVID-ARDS patients. Plasma was isolated from whole blood via centrifugation. Aliquots were frozen at −80 ˚C before analysis.
Purification of SARS-CoV-2 viral proteins
The ectodomain of the SARS-CoV-2 spike trimer45 was cloned into mammalian expression vector pCAGGS (Addgene) with a fold-on tag followed by 6×His tag and Strep-tag II at the C terminal. This expression vector was transiently transfected into HEK293F cells and the spike trimer secreted in the supernatant was purified 3–5 d post-transfection by metal affinity chromatography using an Ni-NTA (QIAGEN) column. SARS-CoV-2 N protein was cloned into pET28a(+) vector (Merck Millipore) with an AAALE linker and 6×His tag at the C terminal. The nanoparticle construct was then used to transform into Escherichia coli strain BL21 (DE3) pLysS cells and the target protein was produced and purified from the bacterial lysate by metal affinity chromatography using an Ni-NTA column, followed by size-exclusion chromatography on a Superdex 200 10/300 GL column.
ELISA to detect virus-specific antibodies
The SARS-CoV-2 spike trimer and N were coated on 96-well ELISA plates at 4 °C overnight and unbound proteins were then removed by washing with PBS, followed by blocking with PBS/3% nonfat dry milk. Plasma samples were serially diluted in PBS with Tween 20 (PBST; 0.1% Tween-20 in PBS) + 10% FCS starting with 1:100 and 5 successive fourfold dilutions into each well of the coated plate, which was incubated at 37 °C for 1 h, followed by washing 6 times with PBST. Peroxidase AffiniPure goat anti-human IgG (H+L) antibody (1:3,000 dilution), anti-human IgM antibody (1:10,000 dilution) (Jackson ImmunoResearch) or anti-human IgA antibody (1:5,000 dilution) (Thermo Fisher Scientific) was subsequently added to each well and incubated for 1 h at 37 °C, washed and tetramethylbenzidine substrate (Sigma-Aldrich) was added and the reaction was stopped using 1 M of sulfuric acid. Absorbance was measured at 450 nm and expressed as an optical density or OD450 value. Identical serial dilutions were performed for all samples with no missing titrations. For the cell-based ELISA, HEK293T cells were transfected with full-length, codon-optimized S protein from SARS-CoV-2, SARS-CoV-2 D614G variant, MERS, SARS-CoV-1 or empty pCAGGS control vector (Epoch Life Science). Transfected cells were seeded onto 96-well plates (40,000 cells per well) and cultured at 37 °C, 5% CO2 overnight, then heat-inactivated plasma samples were added for 30 min on ice, followed by washing with cold PBS and fixation in 4% paraformaldehyde. The fixed cells were stained with protein G Alexa Fluor 488 (ThermoFisher) and 4′,6-diamidino-2-phenylindole (DAPI), and fluorescence images were acquired using the IN Cell Analyzer 2500HS high content analysis imaging system (Cytiva).
Pseudovirus neutralization assay
We adapted a pseudovirus-based neutralization strategy we previously developed to measure the inhibition of infection by high biocontainment enveloped viruses in a large number of samples under low-level biocontainment20,21. For this assay, SARS-CoV-2 S protein was pseudotyped onto recombinant VSV that expresses RFP but does not express the VSV attachment protein, G (VSV-ΔG-RFP). Initially, VSV-ΔG-RFP pseudotyped with VSV G was used to infect 293T (human kidney epithelial) cells that were cotransfected with full-length codon-optimized SARS-CoV-2 S protein (Epoch Life Science), the viral entry receptor ACE2 (Epoch Life Science) and green fluorescent protein (GFP). Infected HEK293T cells were then mixed at a 2:1 ratio with Vero (African green monkey kidney) cells, which have high endogenous expression of ACE2 (ref. 46). Cells were then combined with diluted serum or plasma in 96-well plates. During the assay, infected S protein–expressing HEK293T cells generated VSV-ΔG-RFP viruses bearing S protein, which subsequently infected and drove RFP expression in Vero cells and underwent multiple cycles of entry and budding in HEK293T cells due to the coexpression of S protein with ACE2. The GFP and RFP signals were measured 24–48 h after plating (Infinite M1000 PRO Microplate Reader; Tecan), resulting in robust amplification of the S protein pseudovirus-driven RFP signal between 24 and 48 h. Inhibition of RFP signal amplification indicated S protein neutralizing activity in patient plasma (Extended Data Fig. 1). Identical, fivefold serial dilutions were performed for all samples and there were no missing titration data points for any of the samples.
SARS-CoV-2 viral stock production
SARS-CoV-2 (2019-nCoV/USA_WA1/2020) was kindly provided to B.H. by the World Reference Center for Emerging Viruses and Arboviruses. To generate virus stocks, Vero E6 cells (kindly provided by F. Cosset, International Center for Research in Infectious Diseases, Institut National de la Santé et de la Recherche Médicale) were inoculated with virus at a multiplicity of infection of 0.01. The virus-containing medium was collected at 72 h post infection, clarified by low-speed centrifugation, aliquoted and stored at −80 °C. Virus stock was quantified by limiting the dilution plaque assay on Vero E6 cells as described elsewhere47,48.
Live virus neutralization assay
Twofold dilutions of plasma in 50 μl of DMEM were incubated with 200 plaque-forming units of SARS-CoV-2 in 50 μl of DMEM for 30 min at 4 °C. Then, 100 μl of DMEM and 4% FCS containing 4 × 104 Vero E6 cells were added on top of the former mix to have final dilution of sera from 1:50 to 1:6,400 (4 wells per dilution). Cells were then incubated for 3 d at 37 °C and 5% CO2. The cytopathic effect was revealed by crystal violet staining and scored by an observer blinded to study design and sample identity. Neutralization end point titers were expressed as the value of the last serum dilution that completely inhibited the virus-induced cytopathic effect.
Quantitation of neutralization titers in the pseudovirus assay
For the quantitation of neutralization titers in the pseudovirus assay, the RFP signal driven by the pseudovirus normalized to the GFP signal derived from the SARS-CoV-2 S protein and ACE2 transfected cells was measured at 24 and 48 h; the ratio of normalized RFP at 48 h (RFP48) to normalized RFP at 24 h (RFP24) was calculated. This ratio provides a readout of multicycle infection of the S protein/ACE2-transfected cell monolayer by S protein–bearing pseudoviruses. Neutralizing activity for each sample was calculated by taking the sum of the reciprocal of the RFP48/RFP24 ratio at all 6 plasma dilutions for each sample, as described by Hartman et al.49, and also by percentage inhibition of multicycle replication at each dilution calculated based on the RFP48/RFP24 ratio of the sample, control wells of maximal multicycle replication without inhibition (MAX) and control wells with 100% inhibition of multicycle replication using a lipidated SARS-CoV-2-derived peptide (MIN)50,51. The equation for percentage inhibition of multicycle replication is 100 × (1 − (sample − MIN)/(MAX – MIN)).
Statistical analysis
All statistical analysis was performed using Prism v.8.4.3 (GraphPad Software). Comparisons of clinical data between groups was performed using the Mann–Whitney U-test, one-way analysis of variance (ANOVA) and Dunn’s multiple comparisons test. Comparisons of antibody levels and neutralization activity were performed using a one-way ANOVA and Tukey’s multiple comparisons test. Pairwise correlation analysis was performed using simple linear regression. Multiple linear regression analysis was performed on the combined adult and pediatric data and the adult and pediatric cohorts individually. For all analyses, the outcome variables included the abundance of anti-S IgG, anti-S IgM, anti-N IgG and SARS-CoV-2 neutralizing activity. For the combined adult and pediatric datasets, the independent variable was pediatric age group (‘pediatric’) and covariates included sex, clinical syndrome and time post-symptom onset (days). For the adult and pediatric subgroup analyses, the independent variables were clinical syndrome and age (years) and the covariates included sex and time post-symptom onset (days). For each variable, P values were calculated using the t statistic with two-sided hypothesis testing.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The raw data analyzed for this study are provided as source data. Additional supporting data are available from the authors upon request.
References
Dong, Y. et al. Epidemiology of COVID-19 among children in China. Pediatrics 145, e20200702 (2020).
Wu, Z. & McGoogan, J. M. Characteristics of and important lessons from the coronavirus disease 2019 (COVID-19) outbreak in China: summary of a report of 72 314 cases from the Chinese Center for Disease Control and Prevention. JAMA 323, 1239–1242 (2020).
Cheung, E. W. et al. Multisystem inflammatory syndrome related to COVID-19 in previously healthy children and adolescents in New York city. JAMA 324, 294–296 (2020).
Feldstein, L. R. et al. Multisystem inflammatory syndrome in U.S. children and adolescents. N. Engl. J. Med. 383, 334–346 (2020).
Whittaker, E. et al. Clinical characteristics of 58 children with a pediatric inflammatory multisystem syndrome temporally associated with SARS-CoV-2. JAMA 324, 259–269 (2020).
Singh, A. K. et al. Prevalence of co-morbidities and their association with mortality in patients with COVID-19: a systematic review and meta-analysis. Diabetes Obes. Metab. https://doi.org/10.1111/dom.14124 (2020).
Lanzavecchia, A., Frühwirth, A., Perez, L. & Corti, D. Antibody-guided vaccine design: identification of protective epitopes. Curr. Opin. Immunol. 41, 62–67 (2016).
Corti, D. & Lanzavecchia, A. Broadly neutralizing antiviral antibodies. Annu. Rev. Immunol. 31, 705–742 (2013).
Wang, X. et al. Neutralizing antibodies responses to SARS-CoV-2 in COVID-19 inpatients and convalescent patients. Clin. Infect. Dis. https://doi.org/10.1093/cid/ciaa721 (2020).
Ni, L. et al. Detection of SARS-CoV-2-specific humoral and cellular immunity in COVID-19 convalescent individuals. Immunity 52, 971–977.e3 (2020).
Long, Q.-X. et al. Antibody responses to SARS-CoV-2 in patients with COVID-19. Nat. Med. 26, 845–848 (2020).
Amanat, F. et al. A serological assay to detect SARS-CoV-2 seroconversion in humans. Nat. Med. 26, 1033–1036 (2020).
Bloch, E. M. et al. Deployment of convalescent plasma for the prevention and treatment of COVID-19. J. Clin. Invest. 130, 2757–2765 (2020).
Amanat, F. & Krammer, F. SARS-CoV-2 vaccines: status report. Immunity 52, 583–589 (2020).
Huang, A. T. et al. A systematic review of antibody mediated immunity to coronaviruses: kinetics, correlates of protection, and association with severity. Nat. Commun. 11, 4704 (2020).
Cummings, M. J. et al. Epidemiology, clinical course, and outcomes of critically ill adults with COVID-19 in New York City: a prospective cohort study. Lancet 395, 1763–1770 (2020).
Zachariah, P. et al. Epidemiology, clinical features, and disease severity in patients with coronavirus disease 2019 (COVID-19) in a children’s hospital in New York City, New York. JAMA Pediatr. 174, e202430 (2020).
Korber, B. et al. Tracking changes in SARS-CoV-2 spike: evidence that D614G increases infectivity of the COVID-19 virus. Cell 182, 812–827.e19 (2020).
Cong, Y. et al. Nucleocapsid protein recruitment to replication-transcription complexes plays a crucial role in coronaviral life cycle. J. Virol. 94, e01925-19 (2020).
Talekar, A. et al. Rapid screening for entry inhibitors of highly pathogenic viruses under low-level biocontainment. PLoS ONE 7, e30538 (2012).
Porotto, M. et al. Simulating henipavirus multicycle replication in a screening assay leads to identification of a promising candidate for therapy. J. Virol. 83, 5148–5155 (2009).
Chan, K.-H. et al. Cross-reactive antibodies in convalescent SARS patients’ sera against the emerging novel human coronavirus EMC (2012) by both immunofluorescent and neutralizing antibody tests. J. Infect. 67, 130–140 (2013).
Gao, Q. et al. Development of an inactivated vaccine candidate for SARS-CoV-2. Science 369, 77–81 (2020).
Kohlmeier, J. E. & Woodland, D. L. Immunity to respiratory viruses. Annu. Rev. Immunol. 27, 61–82 (2009).
He, X.-S. et al. Analysis of the frequencies and of the memory T cell phenotypes of human CD8+ T cells specific for influenza A viruses. J. Infect. Dis. 187, 1075–1084 (2003).
PrabhuDas, M. et al. Challenges in infant immunity: implications for responses to infection and vaccines. Nat. Immunol. 12, 189–194 (2011).
Bunyavanich, S., Do, A. & Vicencio, A. Nasal gene expression of angiotensin-converting enzyme 2 in children and adults. JAMA 323, 2427–2429 (2020).
Pierce, C. A. et al. Immune responses to SARS-CoV-2 infection in hospitalized pediatric and adult patients. Sci. Transl. Med. 12, eabd5487 (2020).
Gruber, C. N. et al. Mapping systemic inflammation and antibody responses in multisystem inflammatory syndrome in children (MIS-C). Cell https://doi.org/10.1016/j.cell.2020.09.034 (2020).
Consiglio, C. R. et al. The immunology of multisystem inflammatory syndrome in children with COVID-19. Cell https://doi.org/10.1016/j.cell.2020.09.016 (2020).
Weiskopf, D. et al. Phenotype and kinetics of SARS-CoV-2-specific T cells in COVID-19 patients with acute respiratory distress syndrome. Sci. Immunol. 5, eabd2071 (2020).
Grifoni, A. et al. Targets of T cell responses to SARS-CoV-2 coronavirus in humans with COVID-19 disease and unexposed individuals. Cell 181, 1489–1501.e15 (2020).
Sekine, T. et al. Robust T cell immunity in convalescent individuals with asymptomatic or mild COVID-19. Cell 183, 158–168.e14 (2020).
Kumar, B. V., Connors, T. J. & Farber, D. L. Human T cell development, localization, and function throughout life. Immunity 48, 202–213 (2018).
Mateus, J. et al. Selective and cross-reactive SARS-CoV-2 T cell epitopes in unexposed humans. Science 370, 89–94 (2020).
Rydyznski Moderbacher, C. et al. Antigen-specific adaptive immunity to SARS-CoV-2 in acute COVID-19 and associations with age and disease severity. Cell https://doi.org/10.1016/j.cell.2020.09.038 (2020).
Xu, Y. et al. Characteristics of pediatric SARS-CoV-2 infection and potential evidence for persistent fecal viral shedding. Nat. Med. 26, 502–505 (2020).
Iwasaki, A. & Yang, Y. The potential danger of suboptimal antibody responses in COVID-19. Nat. Rev. Immunol. 20, 339–341 (2020).
Ranieri, V. M. et al. Acute respiratory distress syndrome: the Berlin Definition. JAMA 307, 2526–2533 (2012).
Khemani, R. G., Smith, L. S., Zimmerman, J. J., Erickson, S. & Pediatric Acute Lung Injury Consensus Conference Group. Pediatric acute respiratory distress syndrome: definition, incidence, and epidemiology: proceedings from the Pediatric Acute Lung Injury Consensus Conference. Pediatr. Crit. Care Med. 16, S23–S40 (2015).
Multisystem Inflammatory Syndrome in Children (MIS-C) Associated with Coronavirus Disease 2019 (COVID-19) (CDC Health Alert Network, 2020); https://emergency.cdc.gov/han/2020/han00432.asp
Singer, M. et al. The Third International Consensus Definitions for Sepsis and Septic Shock (Sepsis-3). JAMA 315, 801–810 (2016).
Matics, T. J. & Sanchez-Pinto, L. N. Adaptation and validation of a pediatric sequential organ failure assessment score and evaluation of the Sepsis-3 definitions in critically ill children. JAMA Pediatr. 171, e172352 (2017).
Vasilevskis, E. E. et al. Validity of a modified sequential organ failure assessment score using the Richmond Agitation-Sedation Scale. Crit. Care Med. 44, 138–146 (2016).
Wrapp, D. et al. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science 367, 1260–1263 (2020).
Hoffmann, M. et al. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell 181, 271–280.e8 (2020).
Guillaume, V. et al. Nipah virus: vaccination and passive protection studies in a hamster model. J. Virol. 78, 834–840 (2004).
Mathieu, C. et al. Nipah virus uses leukocytes for efficient dissemination within a host. J. Virol. 85, 7863–7871 (2011).
Hartman, H., Wang, Y., Schroeder, H. W. Jr. & Cui, X. Absorbance summation: a novel approach for analyzing high-throughput ELISA data in the absence of a standard. PLoS ONE 13, e0198528 (2018).
Pessi, A. et al. A general strategy to endow natural fusion-protein-derived peptides with potent antiviral activity. PLoS ONE 7, e36833 (2012).
Xia, S. et al. Fusion mechanism of 2019-nCoV and fusion inhibitors targeting HR1 domain in spike protein. Cell Mol. Immunol. 17, 765–767 (2020).
Acknowledgements
We thank F. Cosset for the donation of the Vero E6 cells. We thank the Medical ICU nurse champions, C. Garcellano, T. Drukdak, H. Avila Raymundo, L. Wagner and R. Lee, who all led the efforts to obtain patient samples for the adult ARDS patients; E. Hernandez and L. Gomez for their roles as clinical coordinators; and the nurses and clinical staff in the Pediatric Intensive Care Unit of MSCHONY. We acknowledge the dedication, commitment and sacrifice of the other nurses, providers and personnel who helped care for these patients during the COVID-19 crisis. We acknowledge the suffering and loss of our COVID-19 patients and that of their families and our community. We also acknowledge the staff of the Columbia University Biobank for their efforts in collecting and preserving COVID-19 patient samples for research studies. This work was supported by National Institutes of Health (NIH) grant nos. AI128949, AI100119 and AI106697 awarded to D.L.F.; NIH grant nos. AI121349, NS091263, NS105699 and AI146980 awarded to M.P.; Fondation de France and ANR Flash-Covid-19, ANR-20-COVI-000 awarded to B.H. and M.P.; and grant no. AI114736 awarded to A.M. S.W. is supported by NIH grant no. K08DK122130. T.J.C. is supported by NIH grant no. K23 AI141686. The funders/sponsors had no role in the design and conduct of the study; the collection, management, analysis and interpretation of the data; the preparation, review or approval of the manuscript; and the decision to submit the manuscript for publication.
Author information
Authors and Affiliations
Contributions
All authors meet the authorship criteria and approved publication of the manuscript. S.P.W., T.J.C., M.P., D.L.F., A.M. and E.H. conceived and designed the study and assays and wrote and/or edited the paper. T.J.C., M.R.B., D.L.F., E.M., J.M. and E.I. coordinated sample acquisition and recruited and consented the MIS-C and COVID-ARDS patients. T.J.C., J.D-P., S.P.W., E.M., J.M. and M.R.B. performed the compilation and analysis of clinical data from the MIS-C and COVID-ARDS patients. E.H. and Z.C.B. recruited and consented CPDs. S.P.W. and Z.C.B. performed the compilation and analysis of data from the convalescent plasma study. F.L.C., S.A.F, M.C. and S.P.W. performed the collection, isolation and storage of samples from CPDs. P.A.S., S.B.W., P.D., J.G., E.I., R.M. and M.M.L.P. performed the collection, isolation and storage of samples from the COVID-ARDS patients. E.H., S.W., K.E.H., F.D.Z. and W.L. established and performed the ELISA assays for the quantification of anti-SARS-CoV-2 S and N protein IgG, IgM and IgA from human serum and plasma. M.P., A.M., F.T.B., D.S. and Y.Z. established and performed the SARS-CoV-2 S protein pseudovirus neutralization assays. C.M., B.H. and D.D. established and performed the SARS-CoV-2 live virus neutralization assay. S.P.W., D.L.F. and T.J.C. performed the compilation and analysis of data from the ELISA and the pseudovirus neutralization and live virus neutralization assays.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer reviewer information Peer reviewer reports are available. Nature Immunology thanks Adrienne Randolph, Stephen Kent and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. L. A. Dempsey was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Specificity of subjects’ antibodies for SARS-CoV-2 S protein.
The specificity of antibodies for SARS-CoV-2 S protein compared to SARS-CoV-1 and MERS S protein was assayed in a cell-based IgG binding assay. HEK293T cells were transfected with S protein and its common variants from the indicated coronaviruses. The transfected cells were then incubated with human plasma from the indicated study groups and bound human IgG was detected using fluorescently tagged protein G (Methods). Shown are the percentage of the S protein transfected cells that are positive for human IgG in each patient group: CPD, black squares, n = 19; COVID-ARDS, red squares, n = 13; pediatric non-MIS-C, n = 28; MIS-C, green circles, n = 16; and control plasma from pre-pandemic donors (grey triangles; Neg, n = 6). Black bar indicates the median+interquartile range. P values were calculated by one-way ANOVA with Dunnett’s multiple comparisons test (CPD, SARS-CoV-2 vs. SARS-CoV-1: P = 0, SARS-CoV-2 vs. MERS: P = 0; COVID-ARDS, SARS-CoV-2 vs. SARS-CoV-1: P = 0, SARS-CoV-2 vs. MERS: P = 0; Ped non-MIS-C, SARS-CoV-2 vs. SARS-CoV-1: P = 0, SARS-CoV-2 vs. MERS: P = 0; MIS-C, SARS-CoV-2 vs. SARS-CoV-1: P = 0, SARS-CoV-2 vs. MERS: P = 0; Negative control, SARS-CoV-2 vs. SARS-CoV-1: P = 0.32, SARS-CoV-2 vs. MERS: P = 0.52).
Supplementary information
Source data
Source Data Fig. 1
Statistical source data
Source Data Fig. 2
Statistical source data
Source Data Fig. 3
Statistical source data
Source Data Fig. 4
Statistical source data
Source Data Extended Data Fig. 1
Statistical source data
Rights and permissions
About this article
Cite this article
Weisberg, S.P., Connors, T.J., Zhu, Y. et al. Distinct antibody responses to SARS-CoV-2 in children and adults across the COVID-19 clinical spectrum. Nat Immunol 22, 25–31 (2021). https://doi.org/10.1038/s41590-020-00826-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-020-00826-9
This article is cited by
-
The mechanisms of milder clinical symptoms of COVID-19 in children compared to adults
Italian Journal of Pediatrics (2024)
-
Severe pediatric COVID-19: a review from the clinical and immunopathophysiological perspectives
World Journal of Pediatrics (2024)
-
SARS-CoV-2 seroprevalence among children in Greece during Omicron variant period
European Journal of Pediatrics (2024)
-
Cellular and molecular features of COVID-19 associated ARDS: therapeutic relevance
Journal of Inflammation (2023)
-
The protective effect of COVID-19 vaccines on developing multisystem inflammatory syndrome in children (MIS-C): a systematic literature review and meta-analysis
Pediatric Rheumatology (2023)