Abstract
CRELD1 is a pivotal factor for heart development, the function of which is unknown in adult life. We here provide evidence that CRELD1 is an important gatekeeper of immune system homeostasis. Exploiting expression variance in large human cohorts contrasting individuals with the lowest and highest CRELD1 expression levels revealed strong phenotypic, functional and transcriptional differences, including reduced CD4+ T cell numbers. These findings were validated in T cell–specific Creld1-deficient mice. Loss of Creld1 was associated with simultaneous overactivation and increased apoptosis, resulting in a net loss of T cells with age. Creld1 was transcriptionally and functionally linked to Wnt signaling. Collectively, gene expression variance in large human cohorts combined with murine genetic models, transcriptomics and functional testing defines CRELD1 as an important modulator of immune homeostasis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Data are available from the corresponding author upon reasonable request. Data from previous human cohorts are available from the following sources: Human Functional Genomics Project, GSE134080; CEDAR cohort, http://cedar-web.giga.ulg.ac.be/; ImmVar project, GSE56035; curated PBMC dataset collection from Warnat-Herresthal et al. 202035, GSE122517. New processed and raw RNA-seq data from the Creld1 conditional mouse model are available through the GEO database (GSE133863, GSE153210; SuperSeries GSE153238). Source data are provided with this paper.
Code availability
Code for reproducing bioinformatic analyses of this study are available from the corresponding author upon reasonable request.
References
Mass, E., Wachten, D., Aschenbrenner, A. C., Voelzmann, A. & Hoch, M. Murine Creld1 controls cardiac development through activation of calcineurin/NFATc1 signaling. Dev. Cell 28, 711–726 (2014).
Robinson, S. W. et al. Missense mutations in CRELD1 are associated with cardiac atrioventricular septal defects. Am. J. Hum. Genet. 72, 1047–1052 (2003).
Aschenbrenner, A. C. et al. A cross-species approach to identify transcriptional regulators exemplified for Dnajc22 and Hnf4a. Sci. Rep. 7, 4056 (2017).
Netea, M. G. et al. Understanding human immune function using the resources from the Human Functional Genomics Project. Nat. Med. 22, 831–833 (2016).
Momozawa, Y. et al. IBD risk loci are enriched in multigenic regulatory modules encompassing putative causative genes. Nat. Commun. 9, 2427 (2018).
Raj, T. et al. Polarization of the effects of autoimmune and neurodegenerative risk alleles in leukocytes. Science 344, 519–523 (2014).
Brodin, P. et al. Variation in the human immune system is largely driven by non-heritable influences. Cell 160, 37–47 (2015).
Piasecka, B. et al. Distinctive roles of age, sex, and genetics in shaping transcriptional variation of human immune responses to microbial challenges. Proc. Natl Acad. Sci. USA 115, E488–E497 (2018).
Li, Y. et al. A functional genomics approach to understand variation in cytokine production in humans. Cell 167, 1099–1110.e14 (2016).
Li, J., Liu, Y., Kim, T., Min, R. & Zhang, Z. Gene expression variability within and between human populations and implications toward disease susceptibility. PLoS Comput. Biol. 6, e1000910 (2010).
De Jager, P. L. et al. ImmVar project: insights and design considerations for future studies of “healthy” immune variation. Semin. Immunol. 27, 51–57 (2015).
Liston, A., Carr, E. J. & Linterman, M. A. Shaping variation in the human immune system. Trends Immunol. 37, 637–646 (2016).
Schirmer, M. et al. Linking the human gut microbiome to inflammatory cytokine production capacity. Cell 167, 1125–1136.e8 (2016).
ter Horst, R. et al. Host and environmental factors influencing individual human cytokine responses. Cell 167, 1111–1124.e13 (2016).
Alpert, A. et al. A clinically meaningful metric of immune age derived from high-dimensional longitudinal monitoring. Nat. Med. 25, 487–495 (2019).
FANTOM Consortium and the RIKEN PMI and CLST (DGT). A promoter-level mammalian expression atlas. Nature 507, 462–470 (2014).
Barrett, T. et al. NCBI GEO: archive for functional genomics data sets—update. Nucleic Acids Res. 41, D991–D995 (2013).
Nikolich-Žugich, J. The twilight of immunity: emerging concepts in aging of the immune system. Nat. Immunol. 19, 10–19 (2018).
Lu, W. et al. CD4:CD8 ratio as a frontier marker for clinical outcome, immune dysfunction and viral reservoir size in virologically suppressed HIV-positive patients. J. Int. AIDS Soc. 18, 20052 (2015).
Leng, S. X. & Margolick, J. B. Aging, sex, inflammation, frailty, and CMV and HIV infections. Cell. Immunol. 348, 104024 (2020).
Voehringer, D. et al. Viral infections induce abundant numbers of senescent CD8 T cells. J. Immunol. 167, 4838–4843 (2001).
Henson, S. M. et al. KLRG1 signaling induces defective Akt (ser473) phosphorylation and proliferative dysfunction of highly differentiated CD8+ T cells. Blood 113, 6619–6628 (2009).
Shi, L. et al. KLRG1 impairs CD4+ T cell responses via p16ink4a and p27kip1 pathways: role in hepatitis B vaccine failure in individuals with hepatitis C virus infection. J. Immunol. 192, 649–657 (2014).
Hamad, A. R. & Schneck, J. P. Antigen-induced T cell death is regulated by CD4 expression. Int. Rev. Immunol. 20, 535–546 (2001).
Chae, W.-J. & Bothwell, A. L. M. Canonical and non-canonical wnt signaling in immune cells. Trends Immunol. 39, 830–847 (2018).
Driessens, G. et al. β-Catenin inhibits T cell activation by selective interference with linker for activation of T cells–phospholipase C-γ1 phosphorylation. J. Immunol. 186, 784–790 (2011).
Yu, Q. et al. T cell factor 1 initiates the T helper type 2 fate by inducing the transcription factor GATA-3 and repressing interferon-γ. Nat. Immunol. 10, 992–999 (2009).
Ma, J., Wang, R., Fang, X., Ding, Y. & Sun, Z. Critical role of TCF-1 in repression of the IL-17 gene. PLoS ONE 6, e24768 (2011).
Lorenz, H., Hailey, D. W., Wunder, C. & Lippincott-Schwartz, J. The fluorescence protease protection (FPP) assay to determine protein localization and membrane topology. Nat. Protoc. 1, 276–279 (2006).
Wright, S. C. et al. FZD5 is a Gαq-coupled receptor that exhibits the functional hallmarks of prototypical GPCRs. Sci. Signal. 11, eaar5536 (2018).
McQuate, A., Latorre-Esteves, E. & Barria, A. A Wnt/calcium signaling cascade regulates neuronal excitability and trafficking of NMDARs. Cell Rep. 21, 60–69 (2017).
Grandy, D. et al. Discovery and characterization of a small molecule inhibitor of the PDZ domain of Dishevelled. J. Biol. Chem. 284, 16256–16263 (2009).
Fuerer, C. & Nusse, R. Lentiviral vectors to probe and manipulate the Wnt signaling pathway. PLoS ONE 5, e9370 (2010).
Veeman, M. T., Slusarski, D. C., Kaykas, A., Louie, S. H. & Moon, R. T. Zebrafish prickle, a modulator of noncanonical Wnt/Fz signaling, regulates gastrulation movements. Curr. Biol. 13, 680–685 (2003).
Warnat-Herresthal, S. et al. Scalable prediction of acute myeloid leukemia using high-dimensional machine learning and blood transcriptomics. iScience 23, 100780 (2020).
Barnes, P. J. Senescence in COPD and its comorbidities. Annu. Rev. Physiol. 79, 517–539 (2017).
MacNee, W. Is chronic obstructive pulmonary disease an accelerated aging disease? Ann. Am. Thorac. Soc. 13, S429–S437 (2016).
Li, Y. et al. Deficient activity of the nuclease MRE11A induces T cell aging and promotes arthritogenic effector functions in patients with rheumatoid arthritis. Immunity 45, 903–916 (2016).
Weyand, C. M., Yang, Z. & Goronzy, J. J. T-cell aging in rheumatoid arthritis. Curr. Opin. Rheumatol. 26, 93–100 (2014).
Storey, J. D. et al. Gene-expression variation within and among human populations. Am. J. Hum. Genet. 80, 502–509 (2007).
Redig, J. K. et al. Allelic interaction between CRELD1 and VEGFA in the pathogenesis of cardiac atrioventricular septal defects. AIMS Genet. 1, 1–19 (2014).
Sprent, J. & Surh, C. D. Normal T cell homeostasis: the conversion of naive cells into memory-phenotype cells. Nat. Immunol. 12, 478–484 (2011).
Staal, F. J. T. & Clevers, H. C. WNT signalling and haematopoiesis: a WNT–WNT situation. Nat. Rev. Immunol. 5, 21–30 (2005).
Zhou, B. O. et al. Bone marrow adipocytes promote the regeneration of stem cells and haematopoiesis by secreting SCF. Nat. Cell Biol. 19, 891–903 (2017).
Sugimura, R. et al. Noncanonical Wnt signaling maintains hematopoietic stem cells in the niche. Cell 150, 351–365 (2012).
Scholz, B. et al. Endothelial RSPO3 controls vascular stability and pruning through non-canonical WNT/Ca2+/NFAT signaling. Dev. Cell 36, 79–93 (2016).
Thrasivoulou, C., Millar, M. & Ahmed, A. Activation of intracellular calcium by multiple Wnt ligands and translocation of β-catenin into the nucleus: a convergent model of Wnt/Ca2+ and Wnt/β-catenin pathways. J. Biol. Chem. 288, 35651–35659 (2013).
Yu, S. et al. The TCF-1 and LEF-1 transcription factors have cooperative and opposing roles in T cell development and malignancy. Immunity 37, 813–826 (2012).
Gullicksrud, J. A. et al. Differential requirements for Tcf1 long isoforms in CD8+ and CD4+ T cell responses to acute viral infection. J. Immunol. 199, 911–919 (2017).
Notani, D. et al. Global regulator SATB1 recruits β-catenin and regulates TH2 differentiation in Wnt-dependent manner. PLoS Biol. 8, e1000296 (2010).
Aguirre-Gamboa, R. et al. Differential effects of environmental and genetic factors on T and B cell immune traits. Cell Rep. 17, 2474–2487 (2016).
Rodríguez, C. I. et al. High-efficiency deleter mice show that FLPe is an alternative to Cre-loxP. Nat. Genet. 25, 139–140 (2000).
Sawada, S., Scarborough, J. D., Killeen, N. & Littman, D. R. A lineage-specific transcriptional silencer regulates CD4 gene expression during T lymphocyte development. Cell 77, 917–929 (1994).
Wilkins, B. J. et al. Calcineurin/NFAT coupling participates in pathological, but not physiological, cardiac hypertrophy. Circ. Res. 94, 110–118 (2004).
A, S. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Korotkevich, G., Sukhov, V. & Sergushichev, A. Fast gene set enrichment analysis. Preprint at bioRxiv https://doi.org/10.1101/060012 (2019).
Acknowledgements
We thank K. Händler, M. Kraut, H. Theis and M. Thielisch for technical assistance and K. Baßler, F. Gondorf, S. Burgdorf and M. Embgenbroich for technical advice. Furthermore, we acknowledge the assistance of the Flow Cytometry Core Facility at the Institute of Experimental Immunology, University of Bonn. We thank M. Hoch for his continuous support of our work; P. Rosenstiel, F. Berberich-Siebelt, M. Väth, D. Cantrell, D. Littmann and L. Fülle for helpful discussion of our work; and M. Davis, S. Turner, L. Klein, J. Bluestone and J. Riley for critically reading our manuscript. A.C.A. was supported by an intramural grant from the Department of Genomics & Immunoregulation at the LIMES Institute. The work was supported by the German Research Foundation (DFG) to J.L.S. under Germany’s Excellence Strategy (EXC2151-390873048) as well as under SCHU 950/8-1; GRK 2168, TP11; SFB704, the BMBF-funded excellence project Diet–Body–Brain (DietBB) and the EU project SYSCID under grant number 733100. M.G.N. was supported by a Spinoza grant of the Netherlands Organization for Scientific Research and an ERC Advanced Grant (833247). E.M. was supported by the DFG under Germany’s Excellence Strategy EXC2151-390873048, the Daimler and Benz Foundation and the Fritz Thyssen Foundation. S.R. and M.G. are supported by the University of Liège (ARC and Fonds Spéciaux) and Fonds de la Recherche Scientifique de Belgique (F.R.S.-FNRS) under grant number EOS 30770923. C.S.F. was supported by the DFG (SFB738/TP3).
Author information
Authors and Affiliations
Contributions
Conceptualization was by A.C.A., L.B., E.M. and J.L.S. The methodology was devised by A.C.A., M.B., L.B., E.M., J.S.-S., J.L.S. and T.U. L.B., P.G., R.t.H., Y.L., J.S.-S., T.U. and S.W.-H. performed formal analysis. A.C.A., L.B., C.S.F., A.H., P.K., M.K., E.M. and L.S. carried out the investigations. Resources were provided by M.B., C.S.F., M.G., M.J., L.A.B.J., M.G.N., S.R. and J.L.S. The draft manuscript was written by A.C.A. and J.L.S. All authors reviewed and edited the manuscript. Visualization was by L.B. and T.U. The project was supervised by A.C.A., L.B. and M.B. Funding acquisition was by A.C.A., J.L.S., M.G.N., E.M., S.R. and M.G.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Editor recognition statement L. A. Dempsey was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
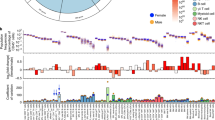
Extended Data Fig. 1 Reduced CRELD1 expression in healthy individuals correlates with reduced T cell numbers and increased immunosenescence-associated signature in blood.
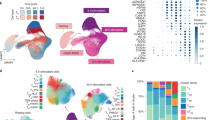
a, Expression of selected genes across 712 human samples of the FANTOM5 dataset ordered by anatomic region. b, Expression fold changes (FCs) in the top 100 human datasets (based on subgroup effect) listed on NCBI GEO profiles independently ranked by FC for each of the analyzed genes. Within the analyzed datasets, 1371 comparisons are interrogated. c, Distribution of expression of selected human and murine genes across the FANTOM5 specimens. d, Expression FCs in the top 100 murine datasets (based on subgroup effect) listed on NCBI GEO profiles independently ranked by FC for each of the analyzed genes. Within the analyzed datasets, 1380 comparisons are interrogated. e, Detailed schematic of the analysis performed on the 500FG dataset. f–l, Box plots of cellular counts in CRELD1hi and CRELD1lo groups in (f) CD4+ T cells (p = 0.0093), g, CD4+ effector memory (CCR7–/CD45RO+ EM, p = 0.024) T cells, h, CD8+ T cells, (i) CD8+ EM T cells (p = 0.014, j, IgM+ B cells (p = 0.047), k, IgD+IgM+ B cells (p = 0.039), and (l) IgD+IgM+CD27– B cells (p = 0.04) (n = 10, unpaired two-sided t-test, *p < 0,05 **p < 0.01 ***p < 0,001). m, Age of the donors in the CRELD1hi and CRELD1lo groups (n = 10). n, Correlation between IMM-AGE down and IMM-AGE up signatures with dots representing all donors in the study, colored by CRELD1 expression (p-value shown for Pearson’s correlation test of CRELD1 expression and signature enrichment, error bar shows 95th confidence interval). o, Ratio between the number of CD4+ and CD8+ T cells for the CRELD1hi and CRELD1lo groups (n = 10, unpaired two-sided t-test, *p = 0.036). p, Percentage of CMV-positive donors in the CRELD1hi and CRELD1lo groups. q–r, Box plots of cellular counts in CMV– CRELD1hi and CRELD1lo groups in (q) CD4+ T cells, (r) CD8+ T cells (n = 10, unpaired two-sided t-test, **p = 0.0043). (Box plots in figure represent median value as well as 25th-75th percentiles; whiskers show the 5th-95th percentiles.).
Extended Data Fig. 2 Creld1 deficiency leads to reduced T cell numbers in middle-aged mice.
a, Schematic representation of Creld1 WT and flox locus; activation of Cre recombinase leads to the deletion of exons 3 to 6 (CD4-Cre+; Creld1cKO). b, Creld1 expression levels in control (grey) or Creld1-deficient (red) CD4+ T cells (n = 3, unpaired two-tailed t-test, ***p = 6.1×10-6). c, Flow cytometry analysis of CD4+ T cells stained for Creld1. d, Body weight of 3-months-old animals (n = 6). e, Weight of spleen and thymus normalized to body weight of 3-months-old animals (spleen n = 6; thymus, control n = 5, Creld1cKO n = 7). f, Number of cells for double negative (DN), double positive (DP) and CD4/CD8 single positive T cells in the thymus of 3-months-old control or Creld1cKO animals (n = 6). g–i, Total cell number for the main T cell populations of control or Creld1cKO 3-months-old animals (n = 6) in (g) spleen, (h) peripheral lymph nodes (pLN, inguinal and brachial) and (i) mesenteric lymph nodes. j, Total cell number for main T cell populations in colon of 1-year-old control or Creld1cKO animals (control n = 4, Creld1cKO n = 5, unpaired two-sided t-test, p > 0.05). k, Body weight of 1-year-old animals (n = 9). l, Normalized spleen and thymus weight of 1-year-old animals (n = 9 and 4, respectively). m, Percentage of CD8+ T cells from mesenteric lymph nodes of middle-age animals producing IFN-γ or TNF (n = 5).(Boxplots in figure represent median value as well as 25th-75th percentiles; whiskers show the 5th-95th percentiles.).
Extended Data Fig. 3 RNA-sequencing reveals loss of Wnt signaling in Creld1-deficient cells.
a, Percentage of dead cells after one day of ex vivo culture with PMA/ionomycin stimulation (n = 6, unpaired two-sided t-test, **p = 0.001). b, Schematic representation of the analysis performed on the RNA-seq data. c, Number of genes used for the generation of the co-expression network from each experimental condition. d, GSEA plot of the IMM-AGE down signature in the 8 h-control condition for Creld1-deficient vs. control cells. e, Mean scaled heat map of the gene set GO_T_CELL_HOMEOSTASIS in 0 h (blue) and 8 h-control (green) samples of Creld1-deficient (red) vs. control (grey) cells. f, GSEA plot of the DEBIASI_APOPTOSIS_BY_REOVIRUS_INFECTION_UP signature in the 8 h-control condition for Creld1-deficient vs. control cells. g, Mean scaled heat map of the leading edge of the GSEA of the DEBIASI_APOPTOSIS_BY_REOVIRUS_INFECTION_UP for 0 h (blue) and 8 h-control (green) samples of Creld1-deficient (red) vs. control (grey) cells. h, Mean scaled heat map of the leading edge of the GSEA of the GO_POSITIVE_REGULATION_OF_ALPHA_BETA_T_CELL_ACTIVATION signature for 8 h stimulated (orange) samples of Creld1-deficient (red) vs. control (grey) cells. i, CoCena2 co-expression network colored according to the identified modules (Fig. 3i) (large network, left panel) or the group fold change (GFC) of each comparison (right panel). j, Heat map for the GFC of each module of the CoCena2 transcription factor network. Module number and number of included genes are indicated in the colored boxes on the right of the heat map with selected gene sets denoted next to it for each of the CoCena2 modules color-coded for the database used for the enrichment. k, GSEA for selected Wnt signaling signatures in naïve and 8h-control samples (p represents statistical summary of GSEA). l, Schematic representation of the sampling performed for RNA-seq analysis of 1-year-old Creld1cKO animals. m, Mean scaled heat map of selected genes in naïve cKO T cells from 3-months-old (young 0 h and 8 h ctrl) and 1-year-old (1yo 0 h). n–o, Enrichment scores for the murine (young animal) Creld1cKO-derived ‘cKO UP’ and ‘cKO DOWN’ signatures plotted against the ranked gene list of the naïve Creld1cKO vs. control CD4+ T cells from 1-year-old animals (n) as well as NES and p-values for the ‘cKO UP’ and ‘cKO DOWN’ gene signatures (p represents statistical summary of GSEA) (o). (ES: enrichment score, GFC: group fold change, NES: normalized enrichment score).
Extended Data Fig. 4 Creld1 deficiency modulates cell activation and is associated with reduced cell survival.
a, Division index calculated on CFSE-labelled conventional CD4+ T cells stimulated for 3 days with anti-CD3/anti-CD28-conjugated beads (n = 3, unpaired two-sided t-test *p = 0.031). b, Un-normalized CFSE dilution profile of control (grey) or Creld1-deficient (red) CD4+ T cells after 3d-stimulation with anti-CD3/anti-CD28-conjugated beads. c, Representative flow cytometry plot for CD25, CD44, and CD69 staining in control or Creld1cKO CD4+ naïve T cells after 24 h of anti-CD3/anti-CD28 stimulation. d, Exemplary plots for Bcl-2, Fas (CD95) and cleaved Caspase3 staining after 8 h ex vivo culture without stimulation. e, Exemplary histogram of flow cytometry data for Bcl-2 expression and box plot for percentage of Bcl-2– cells in freshly isolated CD4+ T cells from spleen (n = 5, unpaired two-tailed t-test, *p = 0.015). f, Percentage and mean fluorescence intensity (MFI) of Fas (CD95) in freshly isolated CD4+ T cells from spleen (n = 4). g–i, Quantification of β-Catenin protein levels in (g) blood (n = 3) and (h) spleen (n = 3) CD4+ T cells measured by flow cytometry. i, β-Catenin protein levels in control and Creld1-deficient CD4+ T cells after 8 h ex vivo stimulation + /- TCR stimulation (n = 3). j, Representative flow cytometry plot for intracellular IFN-γ, IL-4 and IL-17 in in vitro-differentiated control and Creld1-deficient CD4+ T cells. k–l, In vitro CD4+ T cell differentiation towards (k) TH9 (control n = 5, Creld1cKO n = 4, One-Way ANOVA with Bonferroni post-hoc) and (l) iTreg (control n = 5, Creld1cKO n = 4, One-Way ANOVA with Bonferroni post-hoc) subsets. (Box plots in figure represent median value as well as 25th-75th percentiles; whiskers show the 5th-95th percentiles.).
Extended Data Fig. 5 The transmembrane protein Creld1 is linked to Wnt signaling.
Anti-Creld1 staining in fixed/non-permeabilized RFP-Creld1 expressing cells (scale bar 25μm, representative result of n = 4 independent experiments). b, Schematic overview of the fluorescence protease protection (FPP) assay. EYFP, CD3δ-CFP, and YFP-CD3δ expressing cells were used as control for the procedure (scale bar 25μm, representative result of n = 4 independent experiments). c, Schematic representation of the RFP-Creld1 and Creld1-RFP fusion proteins used for the FPP assay. d, Exemplary experiment for the kinetics of the FPP assay. e, Anti-RFP membrane staining in fixed, non-permeabilized Creld1-RFP expressing cells (scale bar 25 µm, representative result of n = 2 independent experiments). f. Creld1 membrane staining in cells expressing different Creld1 mutants (scale bar 25 μm, representative result of n = 3 independent experiments). g, Exemplary confocal image of RFP-Creld1, NFATc1-GFP co-expressing cells after treatment with Wnt signaling inhibitors (scale bar 25 μm). h, Schematic overview of the Wnt pathway and the used inhibitors (i–k) Western blot quantification of β-Catenin protein levels in dependency of Creld1. Values were normalized to β-Actin. i, β-Catenin fold change of either RFP-Creld1 or RFP-Creld1ΔC2 vs. the RFP control condition in cells transfected with RFP-Creld1 (grey) or RFP-Creld1ΔC2 (green) respectively (RFP-Creld1 n = 7, RFP-Creld1ΔC2 n = 9, unpaired two-sided t-test **p = 0.0056). j, Pearson correlation between β-Catenin fold change (see h) and RFP expression (used as control of transfection efficiency for RFP-Creld1ΔC2, error bar shows 95th confidence interval). k, Quantification of β-Catenin sensitive luciferase assay: TOP/FOP ratios upon co-expression with the different RFP-Creld1 mutants normalized to RFP control (RFP_Creld1 n = 9 p = 0.00022, RFP-Creld1ΔN n = 3 p = 0.0028, RFP-Creld1ΔC2 n = 10 p = 0.039, unpaired two-sided t-test *p < 0.05, **p < 0.01, ***p < 0.001). l, Flow cytometry quantification of cleaved Caspase3 after 8 h ex vivo incubation with IL-7, IL-15, or IL-7+IL-15 (n = 3, unpaired two-sided t-test, **p = 0.0045) in control (grey) or Creld1-deficient (red) CD4+ T cells. (Box plots in figure represent median value as well as 25th-75th percentiles, whiskers show the 5th-95th percentiles.).
Extended Data Fig. 6 Human variation in CRELD1 expression reflects transcriptional changes in Creld1 cKO.
a, Schematic representation of the selection of genes used for the cKO DOWN and cKO UP signatures. b, Schematic overview of the analysis performed on the CEDAR dataset. Gene set variation analysis performed on all healthy donors in the CEDAR cohort for both (c) cKO UP and (d) cKO DOWN signatures vs. CRELD1 expression (error bar shows 95th confidence interval, p-value shown for Pearson’s correlation test). e, Schematic overview of the analysis performed on the ImmVar dataset. f, g, Gene set enrichment analysis (GSEA) of the ImmVar CD4+ T cell samples in the CRELD1lo vs. CRELD1hi comparison for the (f) ‘cKO UP’ and ‘cKO DOWN’ gene signatures as well as (g) selected apoptosis and WNT signaling signatures (p-value derived from statistical summary of GSEA). h, GSEA of the conventional monocyte samples of the ImmVar dataset in the CRELD1lo vs. CRELD1hi comparison for the ‘cKO UP’ and ‘cKO DOWN’ gene signatures (p-value derived from statistical summary of GSEA). i, Schematic overview of the analysis performed on our curated PBMC dataset collection.
Extended Data Fig. 7 Modulation of immune effector molecules by CRELD1.
a, Schematic overview of the analysis performed on the 500FG dataset including transcriptome and cytokine production. b–d, Cytokine secretion from PBMC cultures of CRELD1hi and CRELD1lo donors after 24 h of stimulation (b) with Pam3Cys for TNF (p = 0.0002) and with B. burgdorferi for (c) IL-1β (p = 0.0087) or (d) IL-6 (p = 0.037, CRELD1hi n = 9, CRELD1lo n = 10, unpaired two-sided t-test, *p < 0.05, **p < 0.01, ***p < 0.001). e, CRELD1 expression in PBMC from healthy individuals (n = 638) and juvenile idiopathic arthritis (JIA) patients (n = 161, unpaired two-sided t-test, ***p = 3.1×10-5). f, GSEA for the ‘cKO down’, WNT signaling, apoptosis and IMM-AGE down signatures in the comparison JIA vs. healthy (p-value derived from statistical summary of GSEA). (Box plots in figure represent median value as well as 25th-75th percentiles;, whiskers show the 5th-95th percentiles.).
Supplementary information
Supplementary Information
Exemplary gating strategy.
Supplementary Tables
Supplementary Tables 1–9.
Source data
Source Data Fig. 5
Exemplary and unprocessed immunoblots.
Rights and permissions
About this article
Cite this article
Bonaguro, L., Köhne, M., Schmidleithner, L. et al. CRELD1 modulates homeostasis of the immune system in mice and humans. Nat Immunol 21, 1517–1527 (2020). https://doi.org/10.1038/s41590-020-00811-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-020-00811-2
This article is cited by
-
Identification of biomarkers for glycaemic deterioration in type 2 diabetes
Nature Communications (2023)
-
A guide to systems-level immunomics
Nature Immunology (2022)
-
CRELD1 controls T cell homeostasis
Nature Immunology (2020)