Abstract
Ptpn6 is a cytoplasmic phosphatase that functions to prevent autoimmune and interleukin-1 (IL-1) receptor-dependent, caspase-1-independent inflammatory disease. Conditional deletion of Ptpn6 in neutrophils (Ptpn6∆PMN) is sufficient to initiate IL-1 receptor-dependent cutaneous inflammatory disease, but the source of IL-1 and the mechanisms behind IL-1 release remain unclear. Here, we investigate the mechanisms controlling IL-1α/β release from neutrophils by inhibiting caspase-8-dependent apoptosis and Ripk1–Ripk3–Mlkl-regulated necroptosis. Loss of Ripk1 accelerated disease onset, whereas combined deletion of caspase-8 and either Ripk3 or Mlkl strongly protected Ptpn6∆PMN mice. Ptpn6∆PMN neutrophils displayed increased p38 mitogen-activated protein kinase-dependent Ripk1-independent IL-1 and tumor necrosis factor production, and were prone to cell death. Together, these data emphasize dual functions for Ptpn6 in the negative regulation of p38 mitogen-activated protein kinase activation to control tumor necrosis factor and IL-1α/β expression, and in maintaining Ripk1 function to prevent caspase-8- and Ripk3–Mlkl-dependent cell death and concomitant IL-1α/β release.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The data supporting the findings of this study are available within the manuscript and its Supplementary Information files, and are available from the corresponding author upon request. Source data for Figs. 2 and 4–7 are provided with the paper.
Code availability
The custom-scripted macro used for automated image analysis of live-cell imaging data is available from https://doi.org/10.26180/5db913c24b884.
References
Uihlein, L. C., Brandling-Bennett, H. A., Lio, P. A. & Liang, M. G. Sweet syndrome in children. Pediatr. Dermatol. 29, 38–44 (2012).
Prat, L., Bouaziz, J. D., Wallach, D., Vignon-Pennamen, M. D. & Bagot, M. Neutrophilic dermatoses as systemic diseases. Clin. Dermatol. 32, 376–388 (2014).
Going, J. J., Going, S. M., Myskow, M. W. & Beveridge, G. W. Sweet’s syndrome: histological and immunohistochemical study of 15 cases. J. Clin. Pathol. 40, 175–179 (1987).
Bourke, J. F., Jones, J. L., Fletcher, A. & Graham-Brown, R. A. An immunohistochemical study of the dermal infiltrate and epidermal staining for interleukin 1 in 12 cases of Sweet’s syndrome. Br. J. Dermatol. 134, 705–709 (1996).
Giasuddin, A. S., El-Orfi, A. H., Ziu, M. M. & El-Barnawi, N. Y. Sweet’s syndrome: is the pathogenesis mediated by helper T cell type 1 cytokines? J. Am. Acad. Dermatol. 39, 940–943 (1998).
Kluger, N., Gil-Bistes, D., Guillot, B. & Bessis, D. Efficacy of anti-interleukin-1 receptor antagonist anakinra (Kineret®) in a case of refractory Sweet’s syndrome. Dermatology 222, 123–127 (2011).
Beynon, C. et al. Successful treatment of autoimmune disease-associated pyoderma gangrenosum with the IL-1 receptor antagonist anakinra: a case series of 3 patients. J. Clin. Rheumatol. 23, 181–183 (2017).
Leuenberger, M. et al. PASS syndrome: an IL-1-driven autoinflammatory disease. Dermatology 232, 254–258 (2016).
Acquitter, M., Plantin, P., Kupfer, I., Auvinet, H. & Marhadour, T. Anakinra improves pyoderma gangrenosum in psoriatic arthritis: a case report. Ann. Intern. Med. 163, 70–71 (2015).
Nesterovitch, A. B. et al. Alteration in the gene encoding protein tyrosine phosphatase nonreceptor type 6 (PTPN6/SHP1) may contribute to neutrophilic dermatoses. Am. J. Pathol. 178, 1434–1441 (2011).
Croker, B. A. et al. Inflammation and autoimmunity caused by a SHP1 mutation depend on IL-1, MyD88, and a microbial trigger. Proc. Natl Acad. Sci. USA 105, 15028–15033 (2008).
Croker, B. A. et al. Neutrophils require SHP1 to regulate IL-1β production and prevent inflammatory skin disease. J. Immunol. 186, 1131–1139 (2011).
Abram, C. L., Roberge, G. L., Pao, L. I., Neel, B. G. & Lowell, C. A. Distinct roles for neutrophils and dendritic cells in inflammation and autoimmunity in motheaten mice. Immunity 38, 489–501 (2013).
Shultz, L. D. & Green, M. C. Motheaten, an immunodeficient mutant of the mouse. II. Depressed immune competence and elevated serum immunoglobulins. J. Immunol. 116, 936–943 (1976).
Watson, H. A. et al. Purity of transferred CD8+ T cells is crucial for safety and efficacy of combinatorial tumor immunotherapy in the absence of SHP-1. Immunol. Cell Biol. 94, 802–808 (2016).
Gurung, P. et al. Tyrosine kinase SYK licenses MyD88 adaptor protein to instigate IL-1α-mediated inflammatory disease. Immunity 46, 635–648 (2017).
Thrall, R. S., Vogel, S. N., Evans, R. & Shultz, L. D. Role of tumor necrosis factor-α in the spontaneous development of pulmonary fibrosis in viable motheaten mutant mice. Am. J. Pathol. 151, 1303–1310 (1997).
Bossaller, L. et al. Cutting edge: FAS (CD95) mediates noncanonical IL-1β and IL-18 maturation via caspase-8 in an RIP3-independent manner. J. Immunol. 189, 5508–5512 (2012).
Rickard, J. A. et al. RIPK1 regulates RIPK3-MLKL-driven systemic inflammation and emergency hematopoiesis. Cell 157, 1175–1188 (2014).
Wicki, S. et al. Loss of XIAP facilitates switch to TNFα-induced necroptosis in mouse neutrophils. Cell Death Dis. 7, e2422 (2016).
Miwa, K. et al. Caspase 1-independent IL-1β release and inflammation induced by the apoptosis inducer Fas ligand. Nat. Med. 4, 1287–1292 (1998).
Dillon, C. P. et al. RIPK1 blocks early postnatal lethality mediated by caspase-8 and RIPK3. Cell 157, 1189–1202 (2014).
Kaiser, W. J. et al. RIP1 suppresses innate immune necrotic as well as apoptotic cell death during mammalian parturition. Proc. Natl Acad. Sci. USA 111, 7753–7758 (2014).
Oberst, A. et al. Catalytic activity of the caspase-8-FLIPL complex inhibits RIPK3-dependent necrosis. Nature 471, 363–367 (2011).
Kaiser, W. J. et al. RIP3 mediates the embryonic lethality of caspase-8-deficient mice. Nature 471, 368–372 (2011).
Alvarez-Diaz, S. et al. The pseudokinase MLKL and the kinase RIPK3 have distinct roles in autoimmune disease caused by loss of death-receptor-induced apoptosis. Immunity 45, 513–526 (2016).
Lukens, J. R. et al. RIP1-driven autoinflammation targets IL-1α independently of inflammasomes and RIP3. Nature 498, 224–227 (2013).
Roderick, J. E. et al. Hematopoietic RIPK1 deficiency results in bone marrow failure caused by apoptosis and RIPK3-mediated necroptosis. Proc. Natl Acad. Sci. USA 111, 14436–14441 (2014).
Zargarian, S. et al. Phosphatidylserine externalization, ‘necroptotic bodies’ release, and phagocytosis during necroptosis. PLoS Biol. 15, e2002711 (2017).
Gong, Y. N. et al. ESCRT-III acts downstream of MLKL to regulate necroptotic cell death and its consequences. Cell 169, 286–300.e16 (2017).
Greten, F. R. et al. NF-κB is a negative regulator of IL-1β secretion as revealed by genetic and pharmacological inhibition of IKKβ. Cell 130, 918–931 (2007).
Kaiser, W. J. et al. Toll-like receptor 3-mediated necrosis via TRIF, RIP3 and MLKL. J. Biol. Chem. 288, 31268–31279 (2013).
D’Cruz, A. A. et al. The pseudokinase MLKL activates PAD4-dependent NET formation in necroptotic neutrophils. Sci. Signal 11, eaao1716 (2018).
Lalaoui, N. et al. Targeting p38 or MK2 enhances the anti-leukemic activity of SMAC-mimetics. Cancer Cell 29, 145–158 (2016).
Jaco, I. et al. MK2 phosphorylates RIPK1 to prevent TNF-induced cell death. Mol. Cell 66, 698–710 (2017).
Dondelinger, Y. et al. MK2 phosphorylation of RIPK1 regulates TNF-mediated cell death. Nat. Cell Biol. 19, 1237–1247 (2017).
Menon, M. B. et al. p38MAPK/MK2-dependent phosphorylation controls cytotoxic RIPK1 signalling in inflammation and infection. Nat. Cell Biol. 19, 1248–1259 (2017).
Tartey, S., Gurung, P., Dasari, T. K., Burton, A. & Kanneganti, T. D. ASK1/2 signaling promotes inflammation in a mouse model of neutrophilic dermatosis. J. Clin. Invest. 128, 2042–2047 (2018).
De Graaf, C. A. et al. Haemopedia: an expression atlas of murine hematopoietic cells. Stem Cell Rep. 7, 571–582 (2016).
Mandal, P. et al. RIP3 induces apoptosis independent of pronecrotic kinase activity. Mol. Cell 56, 481–495 (2014).
Cook, W. D. et al. RIPK1- and RIPK3-induced cell death mode is determined by target availability. Cell Death Differ. 21, 1600–1612 (2014).
Lawlor, K. E. et al. XIAP loss triggers RIPK3- and caspase-8-driven IL-1β activation and cell death as a consequence of TLR-MyD88-induced cIAP1-TRAF2 degradation. Cell Rep. 20, 668–682 (2017).
Yang, Y., Fang, S., Jensen, J. P., Weissman, A. M. & Ashwell, J. D. Ubiquitin protein ligase activity of IAPs and their degradation in proteasomes in response to apoptotic stimuli. Science 288, 874–877 (2000).
Jia, S. H., Parodo, J., Kapus, A., Rotstein, O. D. & Marshall, J. C. Dynamic regulation of neutrophil survival through tyrosine phosphorylation or dephosphorylation of caspase-8. J. Biol. Chem. 283, 5402–5413 (2008).
Anderton, H. et al. RIPK1 prevents TRADD-driven, but TNFR1 independent, apoptosis during development. Cell Death Differ. 26, 877–889 (2019).
Dowling, J. P., Alsabbagh, M., Del Casale, C., Liu, Z. G. & Zhang, J. TRADD regulates perinatal development and adulthood survival in mice lacking RIPK1 and RIPK3. Nat. Commun. 10, 705 (2019).
Newton, K. et al. RIPK1 inhibits ZBP1-driven necroptosis during development. Nature 540, 129–133 (2016).
Moulin, M. et al. IAPs limit activation of RIP kinases by TNF receptor 1 during development. EMBO J. 31, 1679–1691 (2012).
Zhang, J. et al. Ubiquitin ligases cIAP1 and cIAP2 limit cell death to prevent inflammation. Cell Rep. 27, 2679–2689 (2019).
Matalon, O. et al. Actin retrograde flow controls natural killer cell response by regulating the conformation state of SHP-1. EMBO J. 37, e96264 (2018).
Pao, L. I. et al. B cell-specific deletion of protein-tyrosine phosphatase Shp1 promotes B-1a cell development and causes systemic autoimmunity. Immunity 27, 35–48 (2007).
Shultz, L. D., Coman, D. R., Bailey, C. L., Beamer, W. G. & Sidman, C. L. “Viable motheaten,” a new allele at the motheaten locus. I. Pathology. Am. J. Pathol. 116, 179–192 (1984).
Kelliher, M. A. et al. The death domain kinase RIP mediates the TNF-induced NF-κB signal. Immunity 8, 297–303 (1998).
Dannappel, M. et al. RIPK1 maintains epithelial homeostasis by inhibiting apoptosis and necroptosis. Nature 513, 90–94 (2014).
Polykratis, A. et al. Cutting edge: RIPK1 kinase inactive mice are viable and protected from TNF-induced necroptosis in vivo. J. Immunol. 193, 1539–1543 (2014).
Newton, K., Sun, X. & Dixit, V. M. Kinase RIP3 is dispensable for normal NF-κBs, signaling by the B-cell and T-cell receptors, tumor necrosis factor receptor 1, and Toll-like receptors 2 and 4. Mol. Cell Biol. 24, 1464–1469 (2004).
Salmena, L. et al. Essential role for caspase 8 in T-cell homeostasis and T-cell-mediated immunity. Genes Dev. 17, 883–895 (2003).
Murphy, J. M. et al. The pseudokinase MLKL mediates necroptosis via a molecular switch mechanism. Immunity 39, 443–453 (2013).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Andrews, S. FastQC: a quality control tool for high throughput sequence data. Babraham Bioinformatics https://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (2010).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-Seq data with DESeq2. Genome Biol. 15, 550 (2014).
Newman, A. M. et al. Robust enumeration of cell subsets from tissue expression profiles. Nat. Methods 12, 453–457 (2015).
Pedregosa, F. et al. Scikit-learn: machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Acknowledgements
Mlkl−/− mice were provided by W. S. Alexander (Walter and Eliza Hall Institute of Medical Research). Ripk3−/− mice were provided by V. Dixit (Genentech). Ripk1−/−, conditional Ripk1fl/fl and Ripk1D138N mice were provided by M. Kelliher and M. Pasparakis, supported by NIH/AID grant RO1 AI075118. Casp8 antibody was provided by A. Strasser (Walter and Eliza Hall Institute of Medical Research). This work was supported by NIH grant 5RO1HL124209 (to B.A.C.), the American Asthma Foundation (to B.A.C.), ISF grants 1416/15 and 818/18 (to M.G.), Alpha-1 Foundation grant 615533 (to M.G.), the Recanati Foundation and Varda and Boaz Dotan Research Center (to M.G.), United States–Israel Binational Science Foundation grant 2017176 (to M.G. and B.A.C.), the Australian National Health and Medical Research Council (NHMRC) Dora Lush Scholarship (to J.A.O.) and NHMRC grants 637367, 1145788 and 1162765. This work was supported by a NHMRC Independent Research Institutes Infrastructure Support Scheme grant (9000220), and a Victorian State Government Operational Infrastructure Support grant, support from the Novo Nordisk Foundation provided to the Center for Biosustainability at the Technical University of Denmark (NNF10CC1016517 to N.E.L.) and NIGMS (R35 GM119850 to I.S.). Live-cell imaging performed at Boston Children’s Hospital Intellectual Developmental Disabilities Research Center is supported by grant 1U54HD0902565.
Author information
Authors and Affiliations
Contributions
M.S., A.A.C., J.A.O., A.A.D., J.J.B., R.S.L., M.B.-M., I.S., K.E.L., A.I.S., A.W.R., M.G. and B.A.C. designed the project. M.S., C.J.N., A.A.C., J.A.O., I.S.S., P.R.L., A.A.D., R.O.B., J.J.B., R.S.L., M.B.-M., I.S., S.W., L.H.C., A.I.S., H.P., K.E.L., E.W., N.E.L., A.W.R., M.G. and B.A.C. analyzed the results. W.R., E.W., N.E.L., K.E.L., R.H., M.A.K., J.J.B., C.J.N. and B.A.C. secured funding. M.S., A.A.C., J.A.O., A.A.D., R.O.B., J.J.B., R.S.L., M.B.-M., I.S., S.W., L.H.C., A.I.S., H.P., L.A.O., K.E.L., M.G. and B.A.C. performed the experiments. C.J.N. developed the software. M.S., A.A.C., J.J.B., E.W., N.E.L., K.E.L., A.W.R., M.G. and B.A.C. supervised the project. M.S., J.A.O., K.E.L., N.E.L., I.S., M.G. and B.A.C. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Laurie A. Dempsey was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Ptpn6 prevents an inflammatory signature in bone marrow neutrophils.
a) Gene Set Enrichment Analysis of RNA-Seq transcriptomic data derived from bone marrow neutrophils of female Ptpn6ΔPMNcaspase-8ΔPMNRipk3-/-Mlkl-/- mice and controls. b) Age-dependent effects on the neutrophil transcriptome in Ptpn6ΔPMNcaspase-8ΔPMNRipk3-/-Mlkl-/- mice and controls. Values are gene-wise z-scores of counts normalized using the variance stabilized transformation from DESeq2.
Extended Data Fig. 2 Flow cytometry gating strategy for hematopoietic stem and progenitor cells (HSPC).
For gating of HSPC in Fig. 3, bone marrow cells were sorted after enrichment for hematopoietic progenitor cells by magnetic bead-based depletion of lineage-positive hematopoietic cells. a) Definition and gating strategy for lineage restricted progenitors (LRP), multi-potent progenitors (MPP), and hematopoietic stem cells (HSC), (b) common myeloid progenitors (CMP), megakaryocyte erythroid progenitors (MEP), granulocyte macrophage progenitors (GMP), and (c) common lymphoid progenitors. Lineage-negative cells and gating were defined using isotype control antibodies.
Extended Data Fig. 3 Loss of RIPK1 sensitizes neutrophils to TNF-mediated cell death.
a-b) Live-cell imaging of CTG-labeled wild-type, Ripk1ΔPMN, and Ripk1D138N/D138N neutrophils treated with 2 μM BPT and/or 10 μM z-VAD-fmk + /- 100 ng/mL TNFα. PI and Annexin V were used to monitor changes in viability. Mean and SEM, n = 3 biologically independent samples, and triplicate fields of view per independent biological sample. c) Live-cell imaging of CTG-labeled wild-type, Ptpn6ΔPMN, Ripk1ΔPMN, and Ptpn6ΔPMNRipk1ΔPMN neutrophils treated with saline or 100 ng/mL TNFα. PI and Annexin V were used to monitor changes in viability. Mean and SEM of technical replicates shown. Data are representative of two independent experiments.
Extended Data Fig. 4 TNF induces caspase activation in the absence of Ripk1 in neutrophils.
Live-cell imaging of CellTracker Orange-labeled wild-type, Ripk3-/-, Ripk1ΔPMN, and Ptpn6∆PMNRipk1ΔPMN neutrophils treated with 2 μM BPT and/or 10 μM z-VAD-fmk + /- 100 ng/mL TNFα. CellEvent caspase-3/7 Green Detection Reagent and Draq7 were used to monitor changes in caspase activation and viability. Mean and SEM, n = 3 technical triplicate samples from triplicate fields of view.
Extended Data Fig. 5 Generalized linear mixed effects models (logit link).
Comparison of CTG-positive proportion of a) z-VAD-fmk and b) birinapant treatment of TNF-stimulated Ripk1ΔPMN and wild-type neutrophils. Red curves indicate predicted profiles for wild-type, blue for Ripk1ΔPMN, with line patterns indicating predicted profile for treatment.
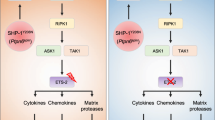
Extended Data Fig. 6 A model illustrating the role of Ptpn6 in regulation of cell death signaling in neutrophils.
Ptpn6 function is controlled in part by Y208−dependent anchoring to the actin−myosin−9 cytoskeleton. In the absence of Ptpn6, the negative regulatory functions of Ripk1 are lost but the kinase domain remains active to influence apoptotic and necroptotic cell death. In Ptpn6ΔPMN neutrophils lacking RIPK1 kinase activity or RIPK1, necroptotic and apoptotic cell death proceed unabated.
Supplementary information
Supplementary Information
Supplementary Tables 1 and 2.
Source data
Source Data Fig. 2
Unprocessed western blots
Source Data Fig. 4
Unprocessed western blots
Source Data Fig. 5
Unprocessed western blots
Source Data Fig. 6
Unprocessed western blots
Source Data Fig. 7
Unprocessed western blots
Rights and permissions
About this article
Cite this article
Speir, M., Nowell, C.J., Chen, A.A. et al. Ptpn6 inhibits caspase-8- and Ripk3/Mlkl-dependent inflammation. Nat Immunol 21, 54–64 (2020). https://doi.org/10.1038/s41590-019-0550-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-019-0550-7
This article is cited by
-
Targeting protein phosphatases in cancer immunotherapy and autoimmune disorders
Nature Reviews Drug Discovery (2023)
-
Dysregulation of testis mRNA expression levels in hatchery-produced vs wild greater amberjack Seriola dumerili
Scientific Reports (2023)
-
The intricate relationship between autoimmunity disease and neutrophils death patterns: a love-hate story
Apoptosis (2023)
-
Radiotherapy modulates tumor cell fate decisions: a review
Radiation Oncology (2022)
-
Targeting protein phosphatases for the treatment of inflammation-related diseases: From signaling to therapy
Signal Transduction and Targeted Therapy (2022)