Abstract
Interferon-γ (IFN-γ) is essential for the innate immune response to intracellular bacteria. Noncoding RNAs and RNA-binding proteins (RBPs) need to be further considered in studies of regulation of the IFN-γ-activated signaling pathway in macrophages. In the present study, we found that the microRNA miR-1 promoted IFN-γ-mediated clearance of Listeria monocytogenes in macrophages by indirectly stabilizing the Stat1 messenger RNA through the degradation of the cytoplasmic long noncoding RNA Sros1. Inducible degradation or genetic loss of Sros1 led to enhanced IFN-γ-dependent activation of the innate immune response. Mechanistically, Sros1 blocked the binding of Stat1 mRNA to the RBP CAPRIN1, which stabilized the Stat1 mRNA and, consequently, promoted IFN-γ–STAT1-mediated innate immunity. These observations shed light on the complex RNA–RNA regulatory networks involved in cytokine-initiated innate responses in host–pathogen interactions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author upon request. The RNA-seq data from the present study are deposited in the National Center for Biotechnology Information’s Gene Expression Omnibus under accession code GSE127769.
Change history
25 February 2020
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Turner, M., Galloway, A. & Vigorito, E. Noncoding RNA and its associated proteins as regulatory elements of the immune system. Nat. Immunol. 15, 484–491 (2014).
Baltimore, D., Boldin, M. P., O’Connell, R. M., Rao, D. S. & Taganov, K. D. MicroRNAs: new regulators of immune cell development and function. Nat. Immunol. 9, 839–845 (2008).
Chen, Y. G., Satpathy, A. T. & Chang, H. Y. Gene regulation in the immune system by long noncoding RNAs. Nat. Immunol. 18, 962–972 (2017).
Xiao, C. & Rajewsky, K. MicroRNA control in the immune system: basic principles. Cell 136, 26–36 (2009).
O’Connell, R. M., Rao, D. S., Chaudhuri, A. A. & Baltimore, D. Physiological and pathological roles for microRNAs in the immune system. Nat. Rev. Immunol. 10, 111–122 (2010).
Lu, L. F. et al. Function of miR-146a in controlling Treg cell-mediated regulation of Th1 responses. Cell 142, 914–929 (2010).
Wang, P. et al. The STAT3-binding long noncoding RNA lnc-DC controls human dendritic cell differentiation. Science 344, 310–313 (2014).
Carpenter, S. et al. A long noncoding RNA mediates both activation and repression of immune response genes. Science 341, 789–792 (2013).
Satpathy, A. T. & Chang, H. Y. Long noncoding RNA in hematopoiesis and immunity. Immunity 42, 792–804 (2015).
Gomez, J. A. et al. The NeST long ncRNA controls microbial susceptibility and epigenetic activation of the interferon-gamma locus. Cell 152, 743–754 (2013).
Atianand, M. K. et al. A long noncoding RNA lincRNA-EPS acts as a transcriptional brake to restrain inflammation. Cell 165, 1672–1685 (2016).
Liu, B. et al. A cytoplasmic NF-kappaB interacting long noncoding RNA blocks IkappaB phosphorylation and suppresses breast cancer metastasis. Cancer Cell 27, 370–381 (2015).
Jiang, M. et al. Self-recognition of an inducible host lncRNA by RIG-I feedback restricts innate immune response. Cell 173, 906–919 e913 (2018).
Wang, P., Xu, J., Wang, Y. & Cao, X. An interferon-independent lncRNA promotes viral replication by modulating cellular metabolism. Science 358, 1051–1055 (2017).
Hamon, M., Bierne, H. & Cossart, P. Listeria monocytogenes: a multifaceted model. Nat. Rev. Microbiol. 4, 423–434 (2006).
Freitag, N. E., Port, G. C. & Miner, M. D. Listeria monocytogenes—from saprophyte to intracellular pathogen. Nat. Rev. Microbiol. 7, 623–628 (2009).
Corr, S. C. & O’Neill, L. A. Listeria monocytogenes infection in the face of innate immunity. Cell Microbiol. 11, 703–709 (2009).
Stavru, F., Archambaud, C. & Cossart, P. Cell biology and immunology of Listeria monocytogenes infections: novel insights. Immunol. Rev. 240, 160–184 (2011).
Hu, X. & Ivashkiv, L. B. Cross-regulation of signaling pathways by interferon-gamma: implications for immune responses and autoimmune diseases. Immunity 31, 539–550 (2009).
Villarino, A. V., Kanno, Y. & O’Shea, J. J. Mechanisms and consequences of Jak-STAT signaling in the immune system. Nat. Immunol. 18, 374–384 (2017).
Boisson-Dupuis, S. et al. Inborn errors of human STAT1: allelic heterogeneity governs the diversity of immunological and infectious phenotypes. Curr. Opin. Immunol. 24, 364–378 (2012).
Chen, K. et al. Methyltransferase SETD2-mediated methylation of STAT1 is critical for interferon antiviral activity. Cell 170, 492–506 e414 (2017).
Liu, S. et al. Nuclear RNF2 inhibits interferon function by promoting K33-linked STAT1 disassociation from DNA. Nat. Immunol. 19, 41–52 (2018).
Kim, T. K. & Maniatis, T. Regulation of interferon-gamma-activated STAT1 by the ubiquitin-proteasome pathway. Science 273, 1717–1719 (1996).
Scortti, M. et al. Coexpression of virulence and fosfomycin susceptibility in Listeria: molecular basis of an antimicrobial in vitro–in vivo paradox. Nat. Med. 12, 515–517 (2006).
Bettencourt, P. et al. Actin-binding protein regulation by microRNAs as a novel microbial strategy to modulate phagocytosis by host cells: the case of N-Wasp and miR-142-3p. Front. Cell Infect. Microbiol. 3, 19 (2013).
Su, X. et al. miRNomes of haematopoietic stem cells and dendritic cells identify miR-30b as a regulator of Notch1. Nat. Commun. 4, 2903 (2013).
Heidersbach, A. et al. microRNA-1 regulates sarcomere formation and suppresses smooth muscle gene expression in the mammalian heart. eLife 2, e01323 (2013).
Wei, Y. et al. Multifaceted roles of miR-1s in repressing the fetal gene program in the heart. Cell Res. 24, 278–292 (2014).
Su, X. et al. An in vivo method to identify microRNA targets not predicted by computation algorithms: p21 targeting by miR-92a in cancer. Cancer Res. 75, 2875–2885 (2015).
Baron-Benhamou, J., Gehring, N. H., Kulozik, A. E. & Hentze, M. W. Using the lambdaN peptide to tether proteins to RNAs. Methods Mol. Biol. 257, 135–154 (2004).
Shiina, N., Shinkura, K. & Tokunaga, M. A novel RNA-binding protein in neuronal RNA granules: regulatory machinery for local translation. J. Neurosci. 25, 4420–4434 (2005).
Wu, Y., Zhu, J., Huang, X. & Du, Z. Crystal structure of a dimerization domain of human Caprin-1: insights into the assembly of an evolutionarily conserved ribonucleoprotein complex consisting of Caprin-1, FMRP and G3BP1. Acta Crystallogr. D72, 718–727 (2016).
Grill, B. et al. Activation/division of lymphocytes results in increased levels of cytoplasmic activation/proliferation-associated protein-1: prototype of a new family of proteins. J. Immunol. 172, 2389–2400 (2004).
Witte, S. & Muljo, S. A. Integrating non-coding RNAs in JAK-STAT regulatory networks. JAKSTAT 3, e28055 (2014).
Shuai, K. & Liu, B. Regulation of JAK-STAT signalling in the immune system. Nat. Rev. Immunol. 3, 900–911 (2003).
Bohmer, F. D. & Friedrich, K. Protein tyrosine phosphatases as wardens of STAT signaling. JAKSTAT 3, e28087 (2014).
Lu, D. et al. The phosphatase DUSP2 controls the activity of the transcription activator STAT3 and regulates TH17 differentiation. Nat. Immunol. 16, 1263–1273 (2015).
Liu, B. et al. PIAS1 selectively inhibits interferon-inducible genes and is important in innate immunity. Nat. Immunol. 5, 891–898 (2004).
Zhang, Q. & Cao, X. Epigenetic regulation of the innate immune response to infection. Nat. Rev. Immunol. 19, 417–432 (2019).
Hentze, M. W., Castello, A., Schwarzl, T. & Preiss, T. A brave new world of RNA-binding proteins. Nat. Rev. Mol. Cell Biol. 19, 327–341 (2018).
Katoh, H. et al. Japanese encephalitis virus core protein inhibits stress granule formation through an interaction with Caprin-1 and facilitates viral propagation. J. Virol. 87, 489–502 (2013).
Nakayama, K. et al. RNG105/caprin1, an RNA granule protein for dendritic mRNA localization, is essential for long-term memory formation. eLife 6, e29677 (2017).
Bidet, K., Dadlani, D. & Garcia-Blanco, M. A. G3BP1, G3BP2 and CAPRIN1 are required for translation of interferon stimulated mRNAs and are targeted by a dengue virus non-coding RNA. PLoS Pathog. 10, e1004242 (2014).
Solomon, S. et al. Distinct structural features of caprin-1 mediate its interaction with G3BP-1 and its induction of phosphorylation of eukaryotic translation initiation factor 2alpha, entry to cytoplasmic stress granules, and selective interaction with a subset of mRNAs. Mol. Cell Biol. 27, 2324–2342 (2007).
He, C. et al. High-resolution mapping of RNA-binding regions in the nuclear proteome of embryonic stem cells. Mol. Cell 64, 416–430 (2016).
van de Veerdonk, F. L. et al. STAT1 mutations in autosomal dominant chronic mucocutaneous candidiasis. N. Engl. J. Med. 365, 54–61 (2011).
Wu, U. I. & Holland, S. M. Host susceptibility to non-tuberculous mycobacterial infections. Lancet Infect. Dis. 15, 968–980 (2015).
Acknowledgements
This work is supported by grants from the National Natural Science Foundation of China (grant no. 81788101), Chinese Academy of Medical Sciences’ Innovation Fund for Medical Sciences (grant no. 2016-12M-1-003) and the National Key Research and Development Program of China (grant no. 2018YFA0507403). We thank H. Lin for technical help.
Author information
Authors and Affiliations
Contributions
X.C. was responsible for research supervision, coordination and strategy. H.X., Y.J., X.X., X.S. and Y.L. performed the experiments. Y.M. generated the Sros1−/− mice. Y.Z. provided miR-1-null (miR-1−/−) mice. Z.S. provided reagents. B.H. provided helpful discussion. H.X. and X.C analyzed the data and wrote the article.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Ioana Visan was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Functional screening of microRNAs in the anti−intracellular bacteria responses.
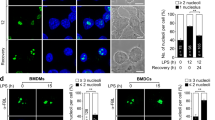
a, Listeria monocytogenes (L.m.) −induced cell death of PMs transfected with antago−NC (antg−NC) or antago−miR−1 (antg−miR−1). The survival of cells was determined 21 h later by 7−AAD uptake. b, qRT−PCR analysis of miR−1 mRNA in sorted mouse cells; results were normalized to U6. HSC, hematopoietic stem cell, NK cell, natural killer cell; macrophage. c, qRT−PCR analysis of mature miR−1 in peritoneal macrophages after intraperitoneal infection with L.m. for the indicated hours. Data were normalized to U6 expression levels. d,h, IFN−γ−induced gene expression level of Stat1 pre−mRNA after either treating with an antg−miR−1 (d) or knockout miR−1(h). e, Immunoblot analysis and quantification analysis of p−STAT1 and STAT1 and β−Actin in control or miR−1−overexpressed PMs after stimulation with IFN−γ for the indicated times. f,g, Expression levels of mature (f) or pre−mRNA transcript (g) of Stat1 in IFN−γ stimulated PMs after prior transfection with miR−1 mimics. Data are representative of three independent experiments (e) or three independent experiments with n = 3 biological replicates (a-h; shown as mean and s.e.m.). Individual data points represent individual biological replicates. *P<0.05, **P<0.01, ***P<0.001, two-tailed unpaired Student’s t−test.
Supplementary Figure 2 miR−1 protects bone marrow−reconstructed mice against L.m. infection.
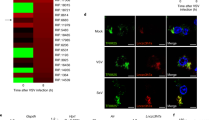
a, Flow cytometry analysis of the percentages of CD49b+NK cells(NK), CD11b+Ly6G+granulocytes, CD11c+MHC−II+DCs and F4/80+CD11b+ macrophages in splenocytes from miR−1+/+ and miR−1−/− mice at postnatal day 15. b, Bacterial load in lung from miR−1 double knockout (miR-1−/−) or littermate (miR-1+/+) − bone marrow transplanted mice after infection for indicated times. n>3 for each genotype. CFU, colony−forming units. c, Hematoxylin−and−eosin staining of lung sections from the indicated mice after infection with L.m. for 36 hr. Scale bars, 20 μm. d, qRT−PCR analysis of Stat1 mRNA in peritoneal lavages from miR−1+/+ and miR−1−/− mice. e, Immunoblot analysis of phosphorylated (p−) or total STAT1 and β−ACTIN in peritoneal lavages from miR−1+/+ and miR−1−/− mice infected with L.m. for 36 hr. f. qRT−PCR analysis of Cxcl9 and Cxcl10 mRNAs in peritoneal lavages from miR-1+/+ and miR-1−/− mice. g. ELISA of IFN−γ and IFN−β in sera. Data are representative of three independent experiments (c,e) or three independent experiments with n = 3 biological replicates (a,d,f,g; shown as mean and s.e.m.). Individual data points represent individual biological replicates. *P<0.05, **P<0.01, ***P<0.001, two-tailed unpaired Student’s t−test.
Supplementary Figure 3 Identification of Sros1 as a lncRNA.
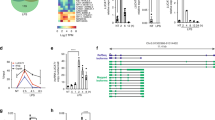
a, qRT−PCR analysis of Stat1 mRNA in BMDMs. The cells were treated with 10 μM ActD for the indicated times. Data were normalized to Gapdh mRNA. b,d, Immunoblot analysis of STAT1 and β−actin in miR−1+/+ or miR−1−/− mice derived BMDMs (b) and in miR−1−overexpressing PMs (d). Cells were treated with 10 μg/ml CHX for the indicated times. Densitometry of STAT1 relative to that of β−actin is shown. c, Stat1 mRNA decay was measured in control or miR−1−overexpressing PMs after IFN−γ treatment. e, Schematic of affinity purification for identification of target mRNAs associated with miR−1. f, Fold changes of Stat1 mRNA in PMs transfected with siRNAs for screening. g, 5′ terminal and 3′ terminal RACE analysis of the full−length Sros1 cloned from PMs. GSPR1−3 and GSPF1−3 indicate the primers used in RACE assays. h, qRT−PCR assay of Sros1 with poly(A)+ or poly(A)− RNA fraction from PMs. i, Relative distributions of RNA populations across each fraction of the sucrose gradient were determined and normalized to the sum of the mRNA signal across all gradient fractions. x axis, fraction number. j, Bioinformatics analysis of Sros1 using Coding Potential Calculator. k, Possible Open Reading Frames (ORFs) was analyzed using ORF−Finder within Sros1 were listed by nucleotide position and amino-acid sequence. Data are representative of three independent experiments (b,d,g,h) or three independent experiments with n=3 biological replicates (a−d,f,i; shown as mean and s.e.m.). Individual data points represent individual biological replicates. *P<0.05, **P<0.01, ***P<0.001, two−tailed unpaired Student’s t−test.
Supplementary Figure 4 Generation of Sros1 knockout mice via CRISPR−Cas9 approach.
a, Schematic illustration of the deleted region in Sros1−/− mice. b, RT−PCR analysis of genomic region from Sros1−/− and the littermates. c, qRT−PCR analysis of Sros1 in peritoneal lavages. Data are representative of three independent experiments (b) or three independent experiments with n=3 biological replicates (c; shown as mean and s.e.m.). Individual data points represent individual biological replicates. *P<0.05, **P<0.01, ***P<0.001, two−tailed unpaired Student’s t−test.
Supplementary Figure 5 Sros1 deficiency does not affect innate immune cell development in mice.
Flow cytometry analysis of the percentages of CD3−NK1.1+NK cells, CD11b+Ly6G+granulocytes, CD11c+MHC−II+DCs and F4/80+CD11b+ macrophages in splenocytes from Sros1−/− mice and their littermates. Data are representative of three independent experiments with n=3 biological replicates (shown as mean and s.e.m.). Individual data points represent individual biological replicates.
Supplementary Figure 6 Sros1 maintains the expression of STAT1 without affecting protein stability.
a, qRT−PCR analysis of Sros1 from Sros1−overexpressing or control RAW264.7 cells. b−c, Immunoblot analysis and quantification analysis of total STAT1 and β−actin in Sros1−silenced PMs (b), or Sros1−overexpressing RAW264.7 cells (c) and control cells. Cells were treated with 10 µg/ml CHX for the indicated times. d, Relative distributions of Stat1 mRNA populations across each fraction in the sucrose gradient were determined and normalized to the sum of the mRNA signal across all gradient fractions in Sros1−silenced PMs. x axis, fraction number. e, qRT−PCR analysis of Sros1 and Stat1 mRNA in PMs after L.m. stimulated for indicated times. f. Changes of total STAT1 protein in PMs transfected with siRNAs for functional screening. Ddx17−1 and Ddx17−2 represent different siRNAs targeting to distinct transcripts of Ddx17. g, Immunoblot analysis and quantification analysis of phosphorylated (p−) or total STAT1, total CAPRIN1 or β−Actin in siRNA transfected PMs after treatment with IFN−γ (50 U/ml) for the indicated times. h, qRT−PCR analysis of Sros1 and Stat1 mRNA in siRNA transfected PMs. Data are representative of three independent experiments (b,c,g) or three independent experiments with n=3 biological replicates (a−h; shown as mean and s.e.m.). Individual data points represent individual biological replicates. *P<0.05, **P<0.01,***P<0.001, two−tailed unpaired Student’s t−test.
Supplementary information
Supplementary Information
Supplementary Figs. 1–6, Tables 1–5 and uncropped immunoblots
Source data
Source Data Supplementary Figure 2a
Raw FACS data
Source Data Supplementary Figure 2c
Unprocessed histology images
Rights and permissions
About this article
Cite this article
Xu, H., Jiang, Y., Xu, X. et al. Inducible degradation of lncRNA Sros1 promotes IFN-γ-mediated activation of innate immune responses by stabilizing Stat1 mRNA. Nat Immunol 20, 1621–1630 (2019). https://doi.org/10.1038/s41590-019-0542-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-019-0542-7
This article is cited by
-
Disparate macrophage responses are linked to infection outcome of Hantan virus in humans or rodents
Nature Communications (2024)
-
Renal biopsies from donors with acute kidney injury show different molecular patterns according to the post-transplant function
Scientific Reports (2024)
-
Caprin-1 influences autophagy-induced tumor growth and immune modulation in pancreatic cancer
Journal of Translational Medicine (2023)
-
Nuclear lncRNA NORSF reduces E2 release in granulosa cells by sponging the endogenous small activating RNA miR-339
BMC Biology (2023)
-
LncTUG1 promotes hepatocellular carcinoma immune evasion via upregulating PD-L1 expression
Scientific Reports (2023)