Abstract
Muscle damage elicits a sterile immune response that facilitates complete regeneration. Here, we used mass spectrometry–based lipidomics to map the mediator lipidome during the transition from inflammation to resolution and regeneration in skeletal muscle injury. We observed temporal regulation of glycerophospholipids and production of pro-inflammatory lipid mediators (for example, leukotrienes and prostaglandins) and specialized pro-resolving lipid mediators (for example, resolvins and lipoxins) that were modulated by ibuprofen. These time-dependent profiles were recapitulated in sorted neutrophils and Ly6Chi and Ly6Clo muscle-infiltrating macrophages, with a distinct pro-resolving signature observed in Ly6Clo macrophages. RNA sequencing of macrophages stimulated with resolvin D2 showed similarities to transcriptional changes found during the temporal transition from Ly6Chi macrophage to Ly6Clo macrophage. In vivo, resolvin D2 increased Ly6Clo macrophages and functional improvement of the regenerating muscle. These results reveal dynamic lipid mediator signatures of innate immune cells and provide a proof of concept for their exploitable effector roles in muscle regeneration.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Shotgun lipidomics and targeted lipidomics raw data are provided in Supplementary Table 1. These datasets are relevant to Figs. 1–4 and Supplementary Figs. 1 and 4. The accession codes for the RNA-seq reported in this paper are SRP145076 (SRA) and GSE114291 (GEO). They have been assigned to Bioproject PRJNA466152 and are relevant to Figs. 4 and 5 and Supplementary Figs. 3 and 5. The RNA-seq analysis data (including complete lists with gene ontology analysis terms) for BMDMs and muscle-infiltrating macrophages are provided in Supplementary Tables 2 and 3.
Change history
02 May 2019
In the version of this article initially published, two arrows in the far right plot of Fig. 3c were aimed incorrectly, and the error bars were missing in Fig. 6e,f. In Fig. 3c, the arrow labeled ‘5-LOX’ should be aimed at the plot measuring LXB4, and the arrow labeled ‘LTA4H’ should be aimed at the plot measuring LTB4. The errors have been corrected in the HTML and PDF versions of the article.
References
Kelly, B. & O’Neill, L. A. J. Metabolic reprogramming in macrophages and dendritic cells in innate immunity. Cell Res. 25, 771–784 (2015).
Dadgar, S. et al. Asynchronous remodeling is a driver of failed regeneration in Duchenne muscular dystrophy. J. Cell Biol. 207, 139–158 (2014).
Tidball, J. G. & Villalta, S. A. Regulatory interactions between muscle and the immune system during muscle regeneration. Am. J. Physiol. Regul. Integr. Comp. Physiol. 298, R1173–R1187 (2010).
Chazaud, B. Macrophages: supportive cells for tissue repair and regeneration. Immunobiology 219, 172–178 (2014).
Yona, S. et al. Fate mapping reveals origins and dynamics of monocytes and tissue macrophages under homeostasis. Immunity 38, 79–91 (2013).
Buckley, C. D., Gilroy, D. W. & Serhan, C. N. Proresolving lipid mediators and mechanisms in the resolution of acute inflammation. Immunity 40, 315–327 (2014).
Tidball, J. G. Regulation of muscle growth and regeneration by the immune system. Nat. Rev. Immunol. 17, 165–178 (2017).
Varga, T. et al. Tissue LyC6- macrophages are generated in the absence of circulating LyC6-monocytes and Nur77 in a model of muscle regeneration. J. Immunol. 191, 5695–5701 (2013).
Arnold, L. et al. Inflammatory monocytes recruited after skeletal muscle injury switch into antiinflammatory macrophages to support myogenesis. J. Exp. Med. 204, 1057–1069 (2007).
Wang, H. et al. Altered macrophage phenotype transition impairs skeletal muscle regeneration. Am. J. Pathol. 184, 1167–1184 (2014).
Serhan, C. N. Pro-resolving lipid mediators are leads for resolution physiology. Nature 510, 92–101 (2014).
Dennis, E. A. & Norris, P. C. Eicosanoid storm in infection and inflammation. Nature Rev. Immunol. 15, 511–523 (2015).
Samuelsson, B. Role of basic science in the development of new medicines: examples from the eicosanoid field. J. Biol. Chem. 287, 10070–10080 (2012).
Spite, M. et al. Resolvin D2 is a potent regulator of leukocytes and controls microbial sepsis. Nature 461, 1287–1291 (2009).
Chiang, N. et al. Infection regulates pro-resolving mediators that lower antibiotic requirements. Nature 484, 524–528 (2012).
Serhan, C. N. Discovery of specialized pro-resolving mediators marks the dawn of resolution physiology and pharmacology. Mol. Asp. Med. 58, 1–11 (2017).
Motwani, M. P. et al. Pro-resolving mediators promote resolution in a human skin model of UV-killed Escherichia coli-driven acute inflammation. JCI Insight 3, e94463 (2018).
Rathod, K. S. et al. Accelerated resolution of inflammation underlies sex differences in inflammatory responses in humans. J. Clin. Invest. 127, 169–182 (2017).
Spite, M., Clària, J. & Serhan, C. N. Resolvins, specialized proresolving lipid mediators, and their potential roles in metabolic diseases. Cell Metabol. 19, 21–36 (2014).
Dalli, J. & Serhan, C. N. Identification and structure elucidation of the proresolving mediators provides novel leads for resolution pharmacology. Br. J. Pharmacol. https://doi.org/10.1111/bph.14336 (2018).
Hellmann, J. et al. Biosynthesis of D-series resolvins in skin provides insights into their role in tissue repair. J. Invest. Dermatol. 138, 2051–2060 (2018).
Serhan, C. N. et al. Macrophage proresolving mediator maresin 1 stimulates tissue regeneration and controls pain. FASEB J. 26, 1755–1765 (2012).
Gronert, K. et al. A role for the mouse 12/15-lipoxygenase pathway in promoting epithelial wound healing and host defense. J. Biol. Chem. 280, 15267–15278 (2005).
Hardy, D. et al. Comparative study of injury models for studying muscle regeneration in mice. PLoS ONE 11, e0147198 (2016).
Childers, M. K., Grange, R. W. & Kornegay, J. N. In vivo canine muscle function assay. J. Vis. Exp. 50, e2623 (2011).
Kornegay, J., Bogan, J. & Bogan, D. Canine models of Duchenne muscular dystrophy and their use in therapeutic strategies. Mamm. Genome 23, 85–108 (2012).
Varga, T. et al. Highly dynamic transcriptional signature of distinct macrophage subsets during sterile inflammation, resolution, and tissue repair. J. Immunol. 196, 4771–4782 (2016).
Chiang, N., Dalli, J., Colas, R. A. & Serhan, C. N. Identification of resolvin D2 receptor mediating resolution of infections and organ protection. J. Exp. Med. 212, 1203–1217 (2015).
Vaidyanathan, S., Patel, C. N., Scarsbrook, A. F. & Chowdhury, F. U. FDG PET/CT in infection and inflammation—current and emerging clinical applications. Clin. Radiol. 70, 787–800 (2015).
Pirooznia, M., Nagarajan, V. & Deng, Y. GeneVenn—a web application for comparing gene lists using Venn diagrams. Bioinformation 1, 420–422 (2007).
Sager, H. B., Kessler, T. & Schunkert, H. Monocytes and macrophages in cardiac injury and repair. J. Thorac. Dis. 9, S30–S35 (2017).
Patsalos, A. et al. In situ macrophage phenotypic transition is affected by altered cellular composition prior to acute sterile muscle injury. J. Physiol. 595, 5815–5842 (2017).
Glaudemans, A. W. et al. The use of (18)F-FDG-PET/CT for diagnosis and treatment monitoring of inflammatory and infectious diseases. Clin. Dev. Immunol. 2013, 623036 (2013).
Kasuga, K. et al. Rapid appearance of resolvin precursors in inflammatory exudates: novel mechanisms in resolution. J. Immunol. 181, 8677–8687 (2008).
Levy, B. D., Clish, C. B., Schmidt, B., Gronert, K. & Serhan, C. N. Lipid mediator class switching during acute inflammation: signals in resolution. Nat. Immunol. 2, 612–619 (2001).
Dalli, J. et al. Resolvin D3 and aspirin-triggered resolvin D3 are potent immunoresolvents. Chem. Biol. 20, 188–201 (2013).
Fredman, G. et al. An imbalance between specialized pro-resolving lipid mediators and pro-inflammatory leukotrienes promotes instability of atherosclerotic plaques. Nat. Commun. 7, 12859 (2016).
Thul, S., Labat, C., Temmar, M., Benetos, A. & Back, M. Low salivary resolvin D1 to leukotriene B4 ratio predicts carotid intima media thickness: a novel biomarker of non-resolving vascular inflammation. Eur. J. Prev. Cardiol. 24, 903–906 (2017).
Markworth, J. F. et al. Human inflammatory and resolving lipid mediator responses to resistance exercise and ibuprofen treatment. AJP Regul. Integr. Comp. Physiol. 305, R1281–R1296 (2013).
Halade, G. V., Norris, P. C., Kain, V., Serhan, C. N. & Ingle, K. A. Splenic leukocytes define the resolution of inflammation in heart failure. Sci. Signal. 11, eaao1818 (2018).
Stables, M. J. et al. Transcriptomic analyses of murine resolution-phase macrophages. Blood 118, e192–e208 (2011).
Dalli, J. & Serhan, C. N. Specific lipid mediator signatures of human phagocytes: microparticles stimulate macrophage efferocytosis and pro-resolving mediators. Blood 120, e60–e72 (2012).
Chiang, N., de la Rosa, X., Libreros, S. & Serhan, C. N. Novel resolvin D2 receptor axis in infectious inflammation. J. Immunol. 198, 842–851 (2017).
Zhang, M. J. et al. Resolvin D2 enhances postischemic revascularization while resolving inflammation. Circulation 134, 666–680 (2016).
Soki, F. N. et al. Polarization of prostate cancer-associated macrophages is induced by milk fat globule-EGF factor 8 (MFG-E8)-mediated efferocytosis. J. Biol. Chem. 289, 24560–24572 (2014).
Inoue, Y. et al. Resolvin D2 limits secondary tissue necrosis after burn wounds in rats. J. Burn Care Res. 39, 423–432 (2017).
Ho, A. T. V. et al. Prostaglandin E2 is essential for efficacious skeletal muscle stem-cell function, augmenting regeneration and strength. Proc. Natl Acad. Sci. USA 114, 6675–6684 (2017).
Baker, L. A. et al. Resolvin E1 (RvE1) attenuates LPS induced inflammation and subsequent atrophy in C2C12 myotubes. J. Cell Biochem. 119, 6094–6103 (2018).
Guardiola, O. et al. Induction of acute skeletal muscle regeneration by cardiotoxin injection. J. Vis. Exp. 119, e54515 (2017).
Wang, M. & Han, X. Multi-dimensional mass spectrometry-based shotgun lipidomics. Meth. Mol. Biol. 1198, 203–220 (2014).
Yang, K., Cheng, H., Gross, R. W. & Han, X. Automated lipid identification and quantification by multidimensional mass spectrometry-based shotgun lipidomics. Anal. Chem. 81, 4356–4368 (2009).
Wang, M., Wang, C., Han, R. H. & Han, X. Novel advances in shotgun lipidomics for biology and medicine. Progr. Lipid Res. 61, 83–108 (2016).
Dalli, J., Colas, R. A., Walker, M. E. & Serhan, C. N. Lipid mediator metabolomics via LC–MS/MS profiling and analysis . Meth. Mol. Biol. 1730, 59–72 (2018).
English, J. T., Norris, P. C., Hodges, R. R., Dartt, D. A. & Serhan, C. N. Identification and profiling of specialized pro-resolving mediators in human tears by lipid mediator metabolomics. Prostaglandins, Leukot. Essent. Fat. Acids 117, 17–27 (2017).
Xia, J., Sinelnikov, I. V., Han, B. & Wishart, D. S. MetaboAnalyst 3.0––making metabolomics more meaningful. Nucleic Acid. Res. 43, 251–257 (2015).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Acknowledgements
The authors are grateful for the outstanding technical contribution of M. Peloquin and acknowledge the discussions and comments on the manuscript from members of the Nagy and Spite laboratories. The authors also thank L. Thomas for excellent administrative assistance. N.G. and L.N. are supported by ‘Chromatin3D’ ITN funded by the European Union under the Horizon2020 Framework Programme (Grant Agreement no. 622934). A.P. and L.N. are supported by ‘NR-NET’ ITN PITN-GA-2013-606806 from the EU-FP7 PEOPLE-2013 program, by grants from the Hungarian Scientific Research Fund (numbers K124298, K126885, K116855 to L.N.) and by the NIH (no. R01DK115924). M.S. acknowledges the support of NIH grant numbers GM095467 (Project 3 and Core B) and HL106173. B.E.S. is supported by an NRSA from the NIH (no. HL136044).
Author information
Authors and Affiliations
Contributions
A.P. and B.E.S. carried out the experiments and performed the measurements. A.P., N.G. and B.E.S. performed the analysis, and designed the figures. N.G., A.P., B.E.S., L.N. and M.S. drafted the manuscript. Computational analyses were performed by N.G. and A.P. C.O.R. and B.E.S. performed targeted lipidomics and in vitro treatments. A.P. produced the muscle samples and characterized the in vivo experiments. X.H. performed the shotgun lipidomics characterization. T.H. aided in mouse colony management. L.N. and M.S. planned the project and supervised the work. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Integrated supplementary information
Supplementary Figure 1 Time-dependent lipid mediator profiles of mice undergoing eccentric exercise-induced muscle injury.
Heatmap displaying the relative abundance of individual lipid mediators at the indicated day post eccentric exercise-induced injury. Each column represents the average of n = 3 biologically independent muscles at the indicated time points post EE.
Supplementary Figure 2 Kinetics of leukocyte accumulation after CTX and EE injury.
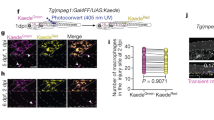
(a) and (b) FACS gating strategy for the analysis and sorting of (a) PMNs and (b) macrophage subsets (Ly6Chi F4/80lo and Ly6Clo F4/80hi cells) from CTX injured muscles. Insets shows representative frequencies (from at least 5 independent experiments) for each gated population. (c) Absolute number of CD45+ cells isolated from gastrocnemius (GAST) muscles at indicated timepoints following eccentric exercise-induced injury (EE). p < 0.05=*, p < 0.001=***, p < 0.0001=**** by Sidak’s multiple comparisons test in two-way ANOVA. Data are shown as mean ± SEM and n = at least 4 biologically independent experiments per time point. (d) Frequencies of PMNs, inflammatory (Ly6Chi F4/80lo) and repair (Ly6Clo F4/80hi) macrophages at indicated timepoints following EE-injury. Data are shown as mean ± SEM and n = at least 3 biologically independent experiments per time point. (e) and (f) Representative flow cytometry pseudocolor density plots (with 2 alternative gatings) for PMNs (Ly6Ghi Ly6Cint F4/80neg), inflammatory (Ly6Chi F4/80lo) and repair (Ly6Clo F4/80hi) macrophages at indicated timepoints post EE injury. Insets shows representative frequencies (from at least 5 independent experiments) for each gated population.
Supplementary Figure 3 Expression of phospholipase A2 isoforms in sorted immune cells following CTX injury.
Heatmap representation of normalized expression of phospholipase a2 isoforms transcripts from sorted neutrophils, Ly6Chi and Ly6Clo macrophage populations at the indicated day post CTX. Hierarchical clustering analysis was performed by Ward’s clustering algorithm and Euclidean measure distance; n = 3 biologically independent samples for each cell populations at the indicated time points. Expression values are visualized as log10 of the normalized expression values from the RNA-seq analysis.
Supplementary Figure 4 COX inhibitor ibuprofen alters the lipid profile and phenotype of macrophages following CTX injury.
(a) Graphical scheme representing the workflow and downstream lipidomic and FACS analyses in the CTX model after ibuprofen (IBP) treatment. (b) Heatmap showing the fold change in abundance of COX-derived pro-inflammatory lipid mediators in TA muscles from untreated and IBP-treated mice at day 1 after CTX (Day 1 group). Each column represents a biologically independent sample per treatment group. (c) Interaction network pathway analyses of the docosahexaenoic (DHA), arachidonic (AA) and eicosapentaenoic (EPA) acid bioactive metabolomes in TA muscles at day 2 (top panel) and day 4 (bottom panel) post CTX injury (Day 2 and Day 4 group). The networks depict both the relative changes of each lipid mediator in untreated (control) compared to IBP treated mice (color of circle) and the absolute abundance of the mediators in IBP treated mice (size of circle). Not detected compounds are shown in black and compounds not included in the analysis in gray. n = 3 biologically independent samples per treatment group per time point. (d) Absolute number of CD45+ cells in TA injured muscles of untreated (control) and IBP treated mice at days 2 and 4 after CTX. Data are shown as mean ± SEM and n = at least 5 biologically independent samples per treatment group per time point. (e) and (f) Frequencies of inflammatory (Ly6Chi F4/80lo) and repair (Ly6Clo F4/80hi) macrophages from untreated (control) and IBP treated animals at days 2 (e) and 4 (f) following CTX injury. p < 0.05=*, p < 0.01=**, p < 0.001=***, p < 0.0001=**** by Sidak’s multiple comparisons test in two-way ANOVA. Data are shown as mean ± SEM, n = at least 5 biologically independent samples per treatment group per time point.
Supplementary Figure 5 Gene ontology analysis of differentially expressed genes in Ly6Chi and Ly6Clo macrophage subsets and effect of RvD2 on gene-expression changes in bone marrow–derived macrophages.
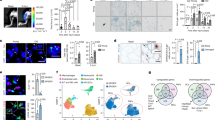
(a) and (b) Gene ontology (GO) analysis of the genes that are differentially expressed in (a)Ly6Chi macrophages at days 2 and 4 post CTX, and (b) Ly6Clo macrophages at days 2 and 4 post CTX. Fold enrichment threshold was set at >2 and Bonferroni-corrected for p value < 0.05. (c) GO analysis of the 172 common differentially expressed genes modulated by RvD2 in BMDMs and during the inflammatory (Ly6Chi) to repair (Ly6Clo) transition during CTX-induced muscle injury. Fold enrichment threshold was set at >2 and Bonferroni-corrected for p value < 0.05. (d) . BMDMs were incubated with 0.1, 1.0 or 10 nM Resolvin D2 for 3 or 4 hours. The expression of genes chosen from the previous RNA-seq analysis were then measured and validated by qRT-PCR. Data are shown as mean ± SEM and n = 3 biologically independent experiments per group per time point. For each gene there was a significant (p < 0.05) relationship based on time but not between doses as measured by two-way ANOVA with Tukey’s multiple comparisons test.
Supplementary Figure 6 RvD2 administration improves regeneration and function in a model of delayed muscle regeneration.
(a) Deuterium-labeled resolvin D2 (d5-RvD2; 4 µg/kg) was injected into the TA muscle of mice. Tissue was collected at the indicated times and subjected to LC-MS/MS analysis. Representative MS/MS fragmentation spectra (from at least 3 independent experiments) of d5-RvD2 identified with diagnostic ion assignments. The structure and fragmentation pattern are shown as insets. (b) Levels of d5-RvD2 measured in the TA muscle 1, 2 or 3 hours after injection. Data are shown as mean ± SEM and n = 3 biologically independent experiments per time point. The dashed line indicates the maximum level of endogenous RvD2 that was detected in vivo in response to cardiotoxin (see Fig. 2e). (c) Ratio between Ly6Clo and Ly6Chi macrophages at day 4 post CTX, in mice receiving saline or RvD2 at days 2 and 3 post CTX. p < 0.01=**, p < 0.0001=**** by Tukey’s multiple comparison test in ordinary one-way ANOVA. Data are shown as mean ± SEM, n = at least 4 biologically independent samples per treatment group per time point. (d) Representative 2-DG uptake whole body FLI image (4 biologically independent experiments were repeated with similar results) of chimeric mice at day 4 post CTX injury receiving saline or RvD2 at day 3 post CTX injury. Contralateral leg was used a saline control. The fluorescence image correlates with levels of active inflammation. (e) TA muscle to body weight ratio between untreated and RvD2 treated mice. p < 0.01=** by Tukey’s multiple comparison test in ordinary one-way ANOVA. Data are shown as mean ± SEM and n = at least 7 biologically independent muscle samples per treatment group.
Supplementary information
Supplementary Table 1: Normalized relative concentrations of the identified lipids and lipid mediators after shotgun and LC-MS/MS lipidomics
An Excel workbook is used as a collection of the normalized relative concentrations of the identified lipids and lipid mediators. Six Excel worksheets contain the concentrations of (a) lipid molecular species after shotgun lipidomics (‘Shotgun_lipidomics’ worksheet), (b) lipid mediators after CTX-injury from whole muscle tissue (TA) (‘CTX_muscle_LC-MS’ worksheet), (c) lipid mediators after eccentric exercise-induced injury from whole muscle extracts (TA) (‘ECC_EX_muscle_LC-MS’ worksheet), (d) lipid mediators after CTX-injury from muscle-retrieved flow-sorted and isolated leukocyte populations (‘CTX_leukocytes_LC-MS’ worksheet), (e) lipid mediators at day 1 after CTX injury and ibuprofen (IBP) treatment (‘CTX_muscle_IBP_Day_1’ worksheet), and (f) lipid mediators at days 2 and 4 after CTX injury and IBP treatment (‘CTX_muscle_IBP_D2_and_D4’ worksheet). In all the worksheets, the concentrations of the identified compounds are indicated at specific time points after injury (days 0, 1, 2, 4 and 8). For the shotgun lipidomic analysis, the mass is also included at the last column. For the lipidomic analysis on whole muscle the unit is pg per gr of tissue; in sorted leukocytes unit is pg per 106 cells and the cell type is also included.
Supplementary Table 2: RNA-seq analysis of bone marrow–derived macrophages
An Excel workbook is used as a collection of the (a) normalized expression values, fold changes (‘RNA-seq Control vs RvD2 BMDMs’ worksheet) and (b) GO analysis of the RvD2 treatment in BMDMs (‘GO analysis RvD2 BMDMs’ worksheet). The top 120 differentially expressed genes on this list are shown as a heatmap in Fig. 5a.
Supplementary Table 3: RNA-seq analysis of muscle-infiltrating macrophages
An Excel workbook is used as a collection of the (a) normalized expression values and FC comparison between muscle infiltrating Ly6Chi and Ly6Clo macrophages of day 2 and day 4 post CTX (‘FC>2 muscle MFs comparisons’ worksheet), (b) GO analysis of the differentially expressed genes between day 2 Ly6Chi and day 4 Ly6Chi (‘GO analysis Ly6Chi D2 vs D4’ worksheet), (c) GO analysis of the differentially expressed genes between day 2 Ly6Clo and day 4 Ly6Clo (‘GO analysis Ly6Clo D2 vs D4 worksheet’), (d) normalized expression values of the 172 commonly differentially expressed genes shown on Fig. 5d (‘172 common genes’ worksheet), (e) GO analysis of the 172 commonly DE genes; identified on Fig. 5c and shown as a heatmap in Fig. 5d; between the following groups: day 2 vs day 4 Ly6Chi; day 2 vs day 4 Ly6Clo ; RvD2 treatment in BMDMs (‘GO analysis of 172 Common genes’ worksheet).
Rights and permissions
About this article
Cite this article
Giannakis, N., Sansbury, B.E., Patsalos, A. et al. Dynamic changes to lipid mediators support transitions among macrophage subtypes during muscle regeneration. Nat Immunol 20, 626–636 (2019). https://doi.org/10.1038/s41590-019-0356-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-019-0356-7
This article is cited by
-
Single-cell transcriptomics identifies the differentiation trajectory from inflammatory monocytes to pro-resolving macrophages in a mouse skin allergy model
Nature Communications (2024)
-
DHA alleviates diet-induced skeletal muscle fiber remodeling via FTO/m6A/DDIT4/PGC1α signaling
BMC Biology (2022)
-
Metabolism of tissue macrophages in homeostasis and pathology
Cellular & Molecular Immunology (2022)
-
Primary cilia on muscle stem cells are critical to maintain regenerative capacity and are lost during aging
Nature Communications (2022)
-
Redressing the interactions between stem cells and immune system in tissue regeneration
Biology Direct (2021)