Abstract
The cytokine IL-6 controls the survival, proliferation and effector characteristics of lymphocytes through activation of the transcription factors STAT1 and STAT3. While STAT3 activity is an ever-present feature of IL-6 signaling in CD4+ T cells, prior activation via the T cell antigen receptor limits IL-6’s control of STAT1 in effector and memory populations. Here we found that phosphorylation of STAT1 in response to IL-6 was regulated by the tyrosine phosphatases PTPN2 and PTPN22 expressed in response to the activation of naïve CD4+ T cells. Transcriptomics and chromatin immunoprecipitation–sequencing (ChIP-seq) of IL-6 responses in naïve and effector memory CD4+ T cells showed how the suppression of STAT1 activation shaped the functional identity and effector characteristics of memory CD4+ T cells. Thus, tyrosine phosphatases induced by the activation of naïve T cells determine the way activated or memory CD4+ T cells sense and interpret cytokine signals.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
Microarray, ChiP-seq and RNA-seq data have been deposited in ArrayExpress under accession codes E-MTAB-7682, E-MTAB-6273 and E-MTAB-6141, respectively. Available p300 ChIP-seq fastq files from CD4 T cells, Th1, Th2 (GSE40463) and Th17 (GSE60482), and Stat1−/− and Stat3−/− T cells (GSE65621) were obtained from GEO (https://www.ncbi.nlm.nih.gov/geo/). Access to interactive datasets can be found at www.jones-cytokinelab.co.uk (addition information in relevant figure legends).
References
MacLeod, M. K., Kappler, J. W. & Marrack, P. Memory CD4 T cells: generation, reactivation and re-assignment. Immunology 130, 10–15 (2010).
O’Shea, J. J. et al. The JAK-STAT pathway: impact on human disease and therapeutic intervention. Annu. Rev. Med. 66, 311–328 (2015).
Hunter, C. A. & Jones, S. A. IL-6 as a keystone cytokine in health and disease. Nat. Immunol. 16, 448–457 (2015).
Nish, S. A. et al. T cell-intrinsic role of IL-6 signaling in primary and memory responses. eLife 3, e01949 (2014).
Longhi, M. P. et al. Interleukin-6 is crucial for recall of influenza-specific memory CD4 T cells. PLoS Pathog. 4, e1000006 (2008).
Strutt, T. M. et al. Direct IL-6 signals maximize protective secondary CD4 T cell responses against influenza. J. Immunol. 197, 3260–3270 (2016).
Jones, G. W. et al. Loss of CD4+ T cell IL-6R expression during inflammation underlines a role for IL-6 trans signaling in the local maintenance of Th17 cells. J. Immunol. 184, 2130–2139 (2010).
Fielding, C. A. et al. Interleukin-6 signaling drives fibrosis in unresolved inflammation. Immunity 40, 40–50 (2014).
Harker, J. A., Lewis, G. M., Mack, L. & Zuniga, E. I. Late interleukin-6 escalates T follicular helper cell responses and controls a chronic viral infection. Science 334, 825–829 (2011).
Briso, E. M., Dienz, O. & Rincon, M. Cutting edge: soluble IL-6R is produced by IL-6R ectodomain shedding in activated CD4 T cells. J. Immunol. 180, 7102–7106 (2008).
Teague, T. K. et al. Activation-induced inhibition of interleukin 6-mediated T cell survival and signal transducer and activator of transcription 1 signaling. J. Exp. Med. 191, 915–926 (2000).
Hong, C. et al. Interleukin-6 expands homeostatic space for peripheral T cells. Cytokine 64, 532–540 (2013).
Chtanova, T. et al. Identification of T cell-restricted genes, and signatures for different T cell responses, using a comprehensive collection of microarray datasets. J. Immunol. 175, 7837–7847 (2005).
Curnow, S. J. et al. Inhibition of T cell apoptosis in the aqueous humor of patients with uveitis by IL-6/soluble IL-6 receptor trans-signaling. J. Immunol. 173, 5290–5297 (2004).
Nowell, M. A. et al. Therapeutic targeting of IL-6 trans signaling counteracts STAT3 control of experimental inflammatory arthritis. J. Immunol. 182, 613–622 (2009).
Atreya, R. et al. Blockade of interleukin 6 trans signaling suppresses T-cell resistance against apoptosis in chronic intestinal inflammation: evidence in Crohn disease and experimental colitis in vivo. Nat. Med. 6, 583–588 (2000).
Liao, W., Lin, J. X., Wang, L., Li, P. & Leonard, W. J. Modulation of cytokine receptors by IL-2 broadly regulates differentiation into helper T cell lineages. Nat. Immunol. 12, 551–559 (2011).
Villarino, A. V., Kanno, Y. & O’Shea, J. J. Mechanisms and consequences of Jak-STAT signaling in the immune system. Nat. Immunol. 18, 374–384 (2017).
Costa-Pereira, A. P. et al. Mutational switch of an IL-6 response to an interferon-γ-like response. Proc. Natl Acad. Sci. USA 99, 8043–8047 (2002).
Hirahara, K. et al. Asymmetric action of STAT transcription factors drives transcriptional outputs and cytokine specificity. Immunity 42, 877–889 (2015).
Peters, A. et al. IL-27 induces Th17 differentiation in the absence of STAT1 signaling. J. Immunol. 195, 4144–4153 (2015).
Jones, G. W. et al. Exacerbated inflammatory arthritis in response to hyperactive gp130 signalling is independent of IL-17A. Ann. Rheum. Dis. 72, 1738–1742 (2013).
Anderson, A. E. et al. IL-6-driven STAT signalling in circulating CD4+ lymphocytes is a marker for early anticitrullinated peptide antibody-negative rheumatoid arthritis. Ann. Rheum. Dis. 75, 466–473 (2016).
Pratt, A. G. et al. A CD4 T cell gene signature for early rheumatoid arthritis implicates interleukin 6-mediated STAT3 signalling, particularly in anti-citrullinated peptide antibody-negative disease. Ann. Rheum. Dis. 71, 1374–1381 (2012).
Hnisz, D. et al. Super-enhancers in the control of cell identity and disease. Cell 155, 934–947 (2013).
Whyte, W. A. et al. Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell 153, 307–319 (2013).
Vahedi, G. et al. Super-enhancers delineate disease-associated regulatory nodes in T cells. Nature 520, 558–562 (2015).
Ciofani, M. et al. A validated regulatory network for Th17 cell specification. Cell 151, 289–303 (2012).
Humby, F. et al. Evaluation of minimally invasive, ultrasound-guided synovial biopsy techniques by the OMERACT filter–determining validation requirements. J. Rheumatol. 43, 208–213 (2016).
Lu, D. et al. The phosphatase DUSP2 controls the activity of the transcription activator STAT3 and regulates T17 differentiation. Nat. Immunol. 16, 1263–1273 (2015).
Bohmer, F. D. & Friedrich, K. Protein tyrosine phosphatases as wardens of STAT signaling. JAKSTAT 3, e28087 (2014).
ten Hoeve, J. et al. Identification of a nuclear Stat1 protein tyrosine phosphatase. Mol. Cell Biol. 22, 5662–5668 (2002).
Wu, T. R. et al. SHP-2 is a dual-specificity phosphatase involved in Stat1 dephosphorylation at both tyrosine and serine residues in nuclei. J. Biol. Chem. 277, 47572–47580 (2002).
Hinks, A. et al. Association between the PTPN22 gene and rheumatoid arthritis and juvenile idiopathic arthritis in a UK population: further support that PTPN22 is an autoimmunity gene. Arthritis Rheum. 52, 1694–1699 (2005).
Sharp, R. C., Abdulrahim, M., Naser, E. S. & Naser, S. A. Genetic variations of PTPN2 and PTPN22: role in the pathogenesis of type 1 diabetes and Crohn’s disease. Front. Cell Infect. Microbiol. 5, 95 (2015).
Pitzalis, C., Jones, G. W., Bombardieri, M. & Jones, S. A. Ectopic lymphoid-like structures in infection, cancer and autoimmunity. Nat. Rev. Immunol. 14, 447–462 (2014).
Wiede, F., Sacirbegovic, F., Leong, Y. A., Yu, D. & Tiganis, T. PTPN2-deficiency exacerbates T follicular helper cell and B cell responses and promotes the development of autoimmunity. J. Autoimmun. 76, 85–100 (2017).
Wiede, F. et al. T cell protein tyrosine phosphatase attenuates T cell signaling to maintain tolerance in mice. J. Clin. Invest. 121, 4758–4774 (2011).
Salmond, R. J., Brownlie, R. J., Morrison, V. L. & Zamoyska, R. The tyrosine phosphatase PTPN22 discriminates weak self peptides from strong agonist TCR signals. Nat. Immunol. 15, 875–883 (2014).
Wiede, F., La Gruta, N. L. & Tiganis, T. PTPN2 attenuates T-cell lymphopenia-induced proliferation. Nat. Commun. 5, 3073 (2014).
Look, D. C., Pelletier, M. R., Tidwell, R. M., Roswit, W. T. & Holtzman, M. J. Stat1 depends on transcriptional synergy with Sp1. J. Biol. Chem. 270, 30264–30267 (1995).
Cantwell, C. A., Sterneck, E. & Johnson, P. F. Interleukin-6-specific activation of the C/EBPdelta gene in hepatocytes is mediated by Stat3 and Sp1. Mol. Cell Biol. 18, 2108–2117 (1998).
Zhang, Z., Jones, S., Hagood, J. S., Fuentes, N. L. & Fuller, G. M. STAT3 acts as a co-activator of glucocorticoid receptor signaling. J. Biol. Chem. 272, 30607–30610 (1997).
Zhu, M., John, S., Berg, M. & Leonard, W. J. Functional association of Nmi with Stat5 and Stat1 in IL-2- and IFNγ-mediated signaling. Cell 96, 121–130 (1999).
Kasahara, Y. et al. Role of interleukin 6 for differential responsiveness of naive and memory CD4+ T cells in CD2-mediated activation. J. Exp. Med. 172, 1419–1424 (1990).
Beagley, K. W. et al. Peyer’s patch B cells with memory cell characteristics undergo terminal differentiation within 24 hours in response to interleukin-6. Cytokine 3, 107–116 (1991).
Gentleman, R. C. et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol. 5, R80 (2004).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Lee, Y. K. et al. Late developmental plasticity in the T helper 17 lineage. Immunity 30, 92–107 (2009).
Haines, C. J. et al. Autoimmune memory T helper 17 cell function and expansion are dependent on interleukin-23. Cell Rep. 3, 1378–1388 (2013).
Mohrs, M., Lacy, D. A. & Locksley, R. M. Stat signals release activated naive Th cells from an anergic checkpoint. J. Immunol. 170, 1870–1876 (2003).
van Panhuys, N., Klauschen, F. & Germain, R. N. T-cell-receptor-dependent signal intensity dominantly controls CD4+ T cell polarization in vivo. Immunity 41, 63–74 (2014).
Jones, G. W. et al. Interleukin-27 inhibits ectopic lymphoid-like structure development in early inflammatory arthritis. J. Exp. Med. 212, 1793–1802 (2015).
Kelly, S. et al. Ultrasound-guided synovial biopsy: a safe, well-tolerated and reliable technique for obtaining high-quality synovial tissue from both large and small joints in early arthritis patients. Ann Rheum Dis. 74, 611–617 (2015).
Humby, F. et al. Ectopic lymphoid structures support ongoing production of class-switched autoantibodies in rheumatoid synovium. PLoS Med. 6, e1 (2009).
Jones, G. W. et al. Naive and activated T cells display differential responsiveness to TL1A that affects Th17 generation, maintenance, and proliferation. FASEB J. 25, 409–419 (2011).
Greenhill, C. J. et al. Interleukin-10 regulates the inflammasome-driven augmentation of inflammatory arthritis and joint destruction. Arthritis Res. Ther. 16, 419 (2014).
Gautier, L., Cope, L., Bolstad, B. M. & Irizarry, R. A. affy–analysis of Affymetrix GeneChip data at the probe level. Bioinformatics 20, 307–315 (2004).
Carvalho, B. S. & Irizarry, R. A. A framework for oligonucleotide microarray preprocessing. Bioinformatics 26, 2363–2367 (2010).
Krzywinski, M. et al. Circos: an information aesthetic for comparative genomics. Genome Res. 19, 1639–1645 (2009).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, (15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15, 550 (2014).
Bray, N. L., Pimentel, H., Melsted, P. & Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nature Biotechnol. 34, 525–527 (2016).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat Methods 9, 357–359 (2012).
Robertson, G. et al. Genome-wide profiles of STAT1 DNA association using chromatin immunoprecipitation and massively parallel sequencing. Nat Methods 4, 651–657 (2007).
Robinson, J. T. et al. Integrative genomics viewer. Nature Biotechnol. 29, 24–26 (2011).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Mahony, S. & Benos, P. V. STAMP: a web tool for exploring DNA-binding motif similarities. Nucleic Acids Res. 35, W253–W258 (2007).
Mathelier, A. et al. JASPAR 2016: a major expansion and update of the open-access database of transcription factor binding profiles. Nucleic Acids Res. 44, D110–D115 (2016).
Acknowledgements
This manuscript is dedicated in fond memory of Javier Uceda Fernandez, a dearly loved friend and colleague who was tragically taken from us on August 29, 2018. Remembered forever. Versus Arthritis (Reference 19796,20770 awarded to S.A.J., N.M.W. and G.W.J.), Hoffmann la Roche (to S.A.J.), Kidney Research UK (to S.A.J.) and the National Health and Medical Research Council of Australia (to T.T.) provided grant support for this project. G.W.J. is recipient of a Versus Arthritis Career Development Fellowship (Reference 20305). J.U.F. held a la Caixa PhD Studentship administered through the British Council, and B.C.C. is supported by a PhD studentship from the Systems Immunity University Research Institute at Cardiff. J.L. is recipient of a Rutherford Fellowship Grant. Analysis of synovial tissues was conducted with support from the Medical Research Council (to C.J.P.; Reference 36661) and the Versus Arthritis-funded Experimental Arthritis Treatment Centre (to C.J.P.; Reference 20022). P.R.T. is recipient of a Wellcome Trust Investigator Award (107964/Z/15/Z) and receives funding through the UK Dementia Research Institute. Bioinformatic analysis was developed with support from the Systems Immunity University Research Institute in Cardiff. The authors would like to thank Y. Kanno, C. Hunter, S. Turner, T. Nakayama, M. Czubala, R. Errington, A. Onodera, K. Hirahara and J. O’Shea for their advice, constructive opinions and kind support.
Author information
Authors and Affiliations
Contributions
S.A.J., G.W.J., A.C.F. and J.P.T. wrote the text and prepared the figures for the paper. All authors reviewed and approved the final manuscript draft. S.A.J., G.W.J., T.T., N.M.W., C.P., P.R.T., C.J.P., J.P.T., and A.C.F. designed the study. B.C.C., R.A., M.J.L., M.J.T., B.S., A.C.F., J.P.T. and N.M.W. conducted bioinformatic, biostatistical and molecular pathway analyses. Laboratory based studies were performed by J.P.T., A.C.F., F.W., M.J.T., X.L., J.U.F., A.D.S., J.L., D.G.H. and D.M.
Corresponding author
Ethics declarations
Competing interests
S.A.J. has received funding support from Hoffman-La Roche, GlaxoSmithKline, Ferring Pharmaceuticals and NovImmune SA and during the past 5 years has acted as an advisory consultant for Roche, Chugai Pharmaceuticals, NovImmune SA, Genentech, Sanofi Regeneron, Johnson & Johnson, Janssen Pharmaceuticals and Eleven Biotherapeutics.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Phenotypical and functional characteristics of CD4+ T cell populations.
(a) Flow cytometry analysis for IFNγ, IL-4, IL-17A and IL-21 in naïve (TN) and effector memory (TEM) CD4+ T cells. Representative plot of n=3. (b) Representative histogram plots for IL-6RA (CD126), Gp130, CD62L and CD44 analysis in CD4+ TN (CD25−CD44lo-intCD62LhiCD127hi), central memory (CD25−CD44hiCD62LhiCD127hi) (TCM), effector (CD25−CD44lo-intCD62LloCD127lo-int) (TEff), and effector memory (CD25−CD44hiCD62LloCD127int-hi) (TEM) T cells. Bottom heat map shows the intensity expression of Sell (CD62L) and Il6ra genes in CD4+ TN and TEM cells. (c) Relative expression of pY-STAT1 (n=3), pY-STAT3 (n=12), pY-STAT5 (n=3), pS-STAT1 (n=3) and pS-STAT3 (n=3) in CD4+ TN, TCM, TEff and TEM cells sorted from human blood after 30 min stimulation with 20 ng/ml IL-6. **** P<0.0001, *** P<0.001 (One-way ANOVA test with Tukey multiple comparison test. Data are shown as mean ± s.e.m).
Supplementary Figure 2 Activation of CD4+ T cell TCRs changes IL-6 response through STAT1.
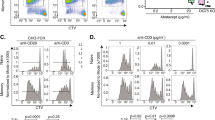
(a) Experimental design for TCR experienced effector-like CD4+ T cells (TEXP). CD4+ TN cells were activated with anti-CD3/CD28 (1 μgr/ml and 5 μgr/ml, respectively) for 3 days after 2 days rest in normal media (RPMI). (b) Representative plot of STAT1 and STAT3 activity of CD4+ TEXP cells exposed to IL-6 (20ng/ml) for 30 minutes (n=3). (c) Analysis of pY-STAT1 and pY-STAT3 (MFI) in CD4+ TN cells (Day 1; n=3) versus CD4+ TEXP cells (Day 5; n=4) derived from WT and IL6ra −/− mice in response to IL-6 (20 ng/ml), HDS (30.74ng/mL; equimolar concentration equivalent to 20 ng/ml IL-6) or IL-27 (20 ng/ml) for 30 minutes as indicated. (d) Mean fluorescence intensity (MFI) for pY-STAT1 in CD4+ TN, TCM, TEM and TEff cells after 20 ng/ml of IL-6, IL-7 or IL-10 stimulation for 30min (n=3). **** P<0.0001, *** P<0.001, ** P<0.01, * P<0.05 (One-way ANOVA with Tukey multiple comparison test (c,d). Data are shown as mean ± s.e.m).
Supplementary Figure 3 Transcriptional response to IL-6 in naïve and differentiated CD4+ T cells.
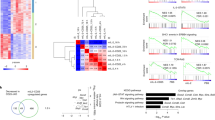
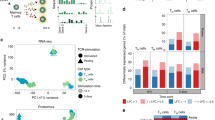
(a) CD4+ TN cells were cultured for 3 days with varying concentrations of CD3 (0.1-10.0 μg/ml) and CD28 antibodies (0.5-15.0 μg/ml). Cells were rested for 2 days before activation for 30 min with 20ng/ml IL-6. Intracellular flow cytometry for pY-STAT1 and pY-STAT3 is presented as a fold change in MFI (n=4). (b) Real-time PCR for Ahr, Bcl3, Bcl6, Il10, Il21, Kat2b, Pim1 and Stat3 in CD4+ TN and TEM cells after 30 min stimulation with 20 ng/ml IL-6 (n=3). (c) Hierarchical clustering, using the complete linkage method (row 1-spearman rank correlation, of all significant transcriptomic data (P<0.05, relative signal intensity of >150, and a >1.5 fold) in each CD4+ T cells population with and without treatment with 20ng/ml IL-6 for 6h. (d) Relative number of genes (up-regulated in red and down-regulated in blue), individual fold change and gene intensity expression for genes differentially regulated by IL-6 in CD4+ TN, TEM and TEXP cells (see Supplementary Fig. 2a) at 6h after stimulation in the presence or absence of CD3/CD28 antibodies. Analysis was confined to genes displaying both a relative signal intensity of >150 and >1.5-fold alteration in expression following IL-6 treatment (P<0.05). n.s – not significant. (One-way ANOVA (a). Data are shown as mean ± s.d).
Supplementary Figure 4 Role of protein tyrosine phosphatases in the regulation of Stat1 activity.
(a) Intensity expression of selected genes involved in the IL-6 signaling cascade in CD4+ T cells subsets after 6h treatment with 20 ng/ml IL-6 or control (PBS). (b) Ptpn2 (n = 2), pY-STAT1 (n = 2) and pY-STAT3 (n = 3) positive cells in CD4+ TN and TEM cells after 30 min IL-6 stimulation (20ng/ml) determined by flow cytometry. (c) Intracellular cytokine staining for IL-21 and IL-17A in CD4+ TEM cells from Ptpn2fl/fl, Lck-Cre Ptpn2fl/fl and Ptpn22−/− mice (n = 3). *** P < 0.0001, ** P < 0.01 (One-way ANOVA (c). Data are shown as mean ± s.d).
Supplementary Figure 5 Validation of STAT1 and STAT3 ChIP-Seq data.
(a) Representative peak alignment for STAT1 (top) an STAT3 (bottom) for selected target genes including: Irf9 (Chr14:55,602,633-55,611,556), Stat3 (Chr11:100,913,375-100,948,267), Nfat5 (Chr8:107,292,133-107,303,956, Il27ra (Chr8: 84,028,287-84,044,575), Stat3 (Chr11:100,931,206-100,942,360) and Cmtm6 (Chr9:114,729,054-114,740,438). (b) Panther analysis (http://pantherdb.org) for the differential expressed genes with a STAT1 or STAT3 binding in the promoter region for CD4+ TEM cells. (c) Enrichment of STAT1 and STAT3 to consensus binding motifs for specific transcription factors. Sequences were identified using MEME and STAMP/Jaspar software. For each condition, the two mostly highly predicted motifs are presented. Statistical significance of the motif prediction is shown as an E-value (MEME tool) (d) ChIP-qPCR analysis of STAT1, STAT3 and SP1 binding to promoter sequences from Irf1 and Socs3. CD4+ TN and TEM cells were treated with 20ng/ml IL-6 for 1 hour. For STAT1 and STAT3, ChIP-qPCR was normalised against binding to a control DNA sequence (NBS: non-binding sequence). To control for constitutive SP1 binding, analysis was normalised to untreated sample (UT) as described in Methods.
Supplementary Figure 6 Overlap of p300 with STAT1 and STAT3 ChIP peaks.
(a) Canonical pathway analysis of IL-6 regulated signatures in CD4+ TN and TEM cells. Statistical analysis shows the profile of canonical pathways that are common and distinct to both population after treatment with 20 ng/ul IL-6 for 30 min. The statistical significance assigned to each pathway is depicted by the size of the dot. A full list of the pathways identified is presented in Supplemental Table 3, and an interactive figure of additional information (http://jones-cytokinelab.co.uk/NI2019/sfigure6a.shtml). (b) ChIP-seq tracks shows p300 peaks for TH1(orange), TH2 (green) and TH17 (purple) CD4+ T cells overlapping with STAT1 and STAT3 peaks in sorted CD4+ TN (blue) and/or TEM (red) cells for the 8 distinct patters of STATs binding. Chromatin marks for Osm (chr11:4,219,039-4,241,349), Stat5b (chr11:100,796,667-100,852,538), Il4ra (chril217:125,551,189-125,572,632), Il21 (chr3:37,220,308-37,234,185), Plgrkt (chr19:29,352,142-29,370,256), Junb (chr:84,975,439-84,980,207), Socs1 (chr16:10,770,927-10,785,406) and Treml2 (chr17:48,289,565-48,303,569) are shown. (c) Prediction of consensus binding motifs for STAT1 or STAT3 transcription factors. Sequences were identified using MEME and STAMP/Jaspar software. Statistical significance of the motif prediction is shown as an E-value (MEME tool). Due to the small number of genes in some of the patterns, not significant motifs were found.
Supplementary Figure 7 Gating strategy for flow cytometry.
(a) FSC vs SSC gate was used to selected live lymphocyte population. Doublet discrimination was applied when we use FSC-H vs FSC-A. After selecting CD4 positive CD25 negative population, we used CD62L and CD44 difference to select CD4+ T cells subsets. (b) After sorting a sample of each population were run to check population purity.
Supplementary information
Rights and permissions
About this article
Cite this article
Twohig, J.P., Cardus Figueras, A., Andrews, R. et al. Activation of naïve CD4+ T cells re-tunes STAT1 signaling to deliver unique cytokine responses in memory CD4+ T cells. Nat Immunol 20, 458–470 (2019). https://doi.org/10.1038/s41590-019-0350-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-019-0350-0
This article is cited by
-
Impaired JAK-STAT pathway signaling in leukocytes of the frail elderly
Immunity & Ageing (2022)