Abstract
Regulatory T cells (Treg cells) maintain host self-tolerance but are a major barrier to effective cancer immunotherapy. Treg cells subvert beneficial anti-tumor immunity by modulating inhibitory receptor expression on tumor-infiltrating lymphocytes (TILs); however, the underlying mediators and mechanisms have remained elusive. Here, we found that the cytokines IL-10 and IL-35 (Ebi3–IL-12α heterodimer) were divergently expressed by Treg cell subpopulations in the tumor microenvironment (TME) and cooperatively promoted intratumoral T cell exhaustion by modulating several inhibitory receptor expression and exhaustion-associated transcriptomic signature of CD8+ TILs. While expression of BLIMP1 (encoded by Prdm1) was a common target, IL-10 and IL-35 differentially affected effector T cell versus memory T cell fates, respectively, highlighting their differential, partially overlapping but non-redundant regulation of anti-tumor immunity. Our results reveal previously unappreciated cooperative roles for Treg cell-derived IL-10 and IL-35 in promoting BLIMP1-dependent exhaustion of CD8+ TILs that limits effective anti-tumor immunity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Bulk RNAseq datasets of CD8+ T cells and Treg cell subpopulations have been deposited in the GEO with the accession code GSE127735. Single-cell RNAseq dataset has been deposited in the GEO with the accession code GSE126184. The RNAseq data sets reported by other studies used to cross-examine with our sequencing data in this study were obtained from GSE9650 and GSE84105. The main data supporting the findings of this study are available in the article and its Supplementary Figures. Data are available from the corresponding authors upon appropriate and reasonable request.
Code availability
Computational and mathematical codes used in the RNAseq analyses supporting the findings of this study are available in the article. Additional information is available from corresponding author on reasonable and appropriate request.
References
Vignali, D. A., Collison, L. W. & Workman, C. J. How regulatory T cells work. Nat. Rev. Immunol. 8, 523–532 (2008).
Sawant, D. V. & Vignali, D. A. Once a treg, always a treg? Immunol. Rev. 259, 173–191 (2014).
Tanaka, A. & Sakaguchi, S. Regulatory T cells in cancer immunotherapy. Cell Res. 27, 109–118 (2017).
Liu, C., Workman, C. J. & Vignali, D. A. Targeting regulatory T cells in tumors. FEBS J. 283, 2731–2748 (2016).
Curiel, T. J. et al. Specific recruitment of regulatory T cells in ovarian carcinoma fosters immune privilege and predicts reduced survival. Nat. Med. 10, 942–949 (2004).
Shimizu, J., Yamazaki, S. & Sakaguchi, S. Induction of tumor immunity by removing CD25+ CD4+ T cells: a common basis between tumor immunity and autoimmunity. J. Immunol. 163, 5211–5218 (1999).
Turnis, M. E. et al. Interleukin-35 limits anti-tumor immunity. Immunity 44, 316–329 (2016).
Shitara, K. & Nishikawa, H. Regulatory T cells: a potential target in cancer immunotherapy. Annals NY Acad. Sci. 1417, 104–115 (2018).
Sawant, D. V., Hamilton, K. & Vignali, D. A. Interleukin-35: expanding its job profile. J. Interferon Cytokine Res. 35, 499–512 (2015).
Brooks, D. G. et al. Interleukin-10 determines viral clearance or persistence in vivo. Nat. Med. 12, 1301–1309 (2006).
Ejrnaes, M. et al. Resolution of a chronic viral infection after interleukin-10 receptor blockade. J. Exp. Med. 203, 2461–2472 (2006).
Tinoco, R. et al. Cell-intrinsic transforming growth factor-beta signaling mediates virus-specific CD8+ T cell deletion and viral persistence in vivo. Immunity 31, 145–157 (2009).
Brooks, D. G. et al. IL-10 and PD-L1 operate through distinct pathways to suppress T-cell activity during persistent viral infection. Proc. Natl Acad. Sci. USA 105, 20428–20433 (2008).
Penaloza-MacMaster, P. et al. Interplay between regulatory T cells and PD-1 in modulating T cell exhaustion and viral control during chronic LCMV infection. J. Exp. Med. 211, 1905–1918 (2014).
Collison, L. W. et al. IL-35-mediated induction of a potent regulatory T cell population. Nat. Immunol. 11, 1093–1101 (2010).
Bettini, M. et al. Prevention of autoimmune diabetes by ectopic pancreatic beta-cell expression of interleukin-35. Diabetes 61, 1519–1526 (2012).
Madan, R. et al. Nonredundant roles for B cell-derived IL-10 in immune counter-regulation. J. Immunol. 183, 2312–2320 (2009).
Rubtsov, Y. P. et al. Regulatory T cell-derived interleukin-10 limits inflammation at environmental interfaces. Immunity 28, 546–558 (2008).
Wei, X. et al. Reciprocal expression of IL-35 and IL-10 defines two distinct effector treg subsets that are required for maintenance of immune tolerance. Cell Rep. 21, 1853–1869 (2017).
Kheradmand, F. et al. A protease-activated pathway underlying Th cell type 2 activation and allergic lung disease. J. Immunol. 169, 5904–5911 (2002).
Moran, A. E. & Hogquist, K. A. T-cell receptor affinity in thymic development. Immunology 135, 261–267 (2012).
Chen, G. et al. Sequence and structural analyses reveal distinct and highly diverse human CD8(+) TCR repertoires to immunodominant viral antigens. Cell Rep. 19, 569–583 (2017).
Levine, A. G., Arvey, A., Jin, W. & Rudensky, A. Y. Continuous requirement for the TCR in regulatory T cell function. Nat. Immunol. 15, 1070–1078 (2014).
Noy, R. & Pollard, J. W. Tumor-associated macrophages: from mechanisms to therapy. Immunity 41, 49–61 (2014).
Genard, G., Lucas, S. & Michiels, C. Reprogramming of tumor-associated macrophages with anticancer therapies: radiotherapy versus chemo- and immunotherapies. Front. Immunol. 8, 828 (2017).
Collison, L. W. et al. The composition and signaling of the IL-35 receptor are unconventional. Nat. Immunol. 13, 290–299 (2012).
Spencer, S. D. et al. The orphan receptor CRF2-4 is an essential subunit of the interleukin 10 receptor. J. Exp. Med. 187, 571–578 (1998).
Wherry, E. J. et al. Molecular signature of CD8+ T cell exhaustion during chronic viral infection. Immunity 27, 670–684 (2007).
Rutishauser, R. L. et al. Transcriptional repressor Blimp-1 promotes CD8(+) T cell terminal differentiation and represses the acquisition of central memory T cell properties. Immunity 31, 296–308 (2009).
Im, S. J. et al. Defining CD8+ T cells that provide the proliferative burst after PD-1 therapy. Nature 537, 417–421 (2016).
Xin, A., Nutt, S. L., Belz, G. T. & Kallies, A. Blimp1: driving terminal differentiation to a T. Adv. Exp. Med. Biol. 780, 85–100 (2011).
Shin, H. et al. A role for the transcriptional repressor Blimp-1 in CD8(+) T cell exhaustion during chronic viral infection. Immunity 31, 309–320 (2009).
Chihara, N. et al. Induction and transcriptional regulation of the co-inhibitory gene module in T cells. Nature 558, 454–459 (2018).
Poholek, A. C. et al. IL-10 induces a STAT3-dependent autoregulatory loop in TH2 cells that promotes Blimp-1 restriction of cell expansion via antagonism of STAT5 target genes. Sci. Immunol. 1, eaaf8612 (2016).
Fontenot, J. D., Gavin, M. A. & Rudensky, A. Y. Foxp3 programs the development and function of CD4+ CD25+ regulatory T cells. Nat. Immunol. 4, 330–336 (2003).
Hori, S., Nomura, T. & Sakaguchi, S. Control of regulatory T cell development by the transcription factor Foxp3. Science 299, 1057–1061 (2003).
Shen, P. et al. IL-35-producing B cells are critical regulators of immunity during autoimmune and infectious diseases. Nature 507, 366–370 (2014).
Roers, A. et al. T cell-specific inactivation of the interleukin 10 gene in mice results in enhanced T cell responses but normal innate responses to lipopolysaccharide or skin irritation. J. Exp. Med. 200, 1289–1297 (2004).
Seo, W., Muroi, S., Akiyama, K. & Taniuchi, I. Distinct requirement of Runx complexes for TCRbeta enhancer activation at distinct developmental stages. Sci. Rep. 7, 41351 (2017).
Roederer, M., Nozzi, J. L. & Nason, M. C. SPICE: exploration and analysis of post-cytometric complex multivariate datasets. Cytometry A 79, 167–174 (2011).
Zheng, G. X. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Angerer, P. et al. destiny: diffusion maps for large-scale single-cell data in R. Bioinformatics 32, 1241–1243 (2016).
Haghverdi, L. et al. Diffusion pseudotime robustly reconstructs lineage branching. Nat. Methods 13, 845–848 (2016).
Yu, D., Huber, W. & Vitek, O. Shrinkage estimation of dispersion in negative binomial models for RNA-seq experiments with small sample size. Bioinformatics 29, 1275–1282 (2013).
Klipper-Aurbach, Y. et al. Mathematical formulae for the prediction of the residual beta cell function during the first two years of disease in children and adolescents with insulin-dependent diabetes mellitus. Med. Hyp. 45, 486–490 (1995).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Pruitt, K. D., Tatusova, T. & Maglott, D. R. NCBI reference sequences (RefSeq): a curated non-redundant sequence database of genomes, transcripts and proteins. Nucleic Acids Res. 35, D61–D65 (2007).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Law, C. W., Chen, Y., Shi, W. & Smyth, G. K. voom: precision weights unlock linear model analysis tools for RNA-seq read counts. Genome Biol. 15, R29 (2014).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Wu, D. & Smyth, G. K. Camera: a competitive gene set test accounting for inter-gene correlation. Nucleic Acids Res. 40, e133 (2012).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Acknowledgements
The authors wish to thank H. Shen, D. Falkner and A. Yates from the Immunology Flow Core for cell sorting; E. Brunazzi and the staff of the Division of Laboratory Animals for animal husbandry; A. Cillo for helpful suggestions regarding scRNAseq analysis; W. Horne, J. Kolls and the University of Pittsburgh HSCRF Genomics Research Core for assistance with sequencing and A. Menk and G. Delgoffe at the Universtiy of Pittsburgh for generation and provision of the BrafPten (clone 24) cell line for tumor growth experiments. The authors also wish to thank the Department of Cardiothoracic Surgery at the University of Pittsburgh, in particular, J. Ward for her help in coordination and gathering patient consents, as well as the Department of Cardiothoracic Surgery at the University of Colorado and the University of Colorado SPORE for providing some samples. This work was supported by the National Institutes of Health (grant nos. R01 CA203689 and P01 AI108545 to D.A.A.V.), NCI Comprehensive Cancer Center Support CORE grant (no. CA047904 to D.A.A.V.) and an SRA from Tizona Therapeutics. This work also benefitted from the Immunology Department Flow Cytometry Core SPECIAL BD LSR FORTESSA funded by NIH grant no. 1S10OD011925-01 (L. Borghesi, Department of Immunology). This project also used the Hillman Cancer Center Immunologic Monitoring and Cellular Products Laboratory that is supported in part by award no. P30 CA047904.
Author information
Authors and Affiliations
Contributions
D.A.A.V. conceived, directed and obtained funding for the project. D.V.S., H.Y. and D.A.A.V. conceptualized, designed, analyzed the experiments and wrote the manuscript. D.V.S. and H.Y. performed the experiments with help from Q.Z. for Treg cell TCRseq and Treg cell subpopulation analysis with multiple organs. M.C. analyzed RNAseq data. C.L., T.T. and R.L. performed single-cell RNAseq experiments and contributed critical reagents and experiment design. Z.S., T.S. and W.C. analyzed single-cell RNAseq data. A.C.P. contributed critical reagents and helped with experimental design and analysis with ChIP–qPCR experiments. A.P. and J.D.L. obtained the NSCLC lung specimens. M.L. and T.C.B. processed and analyzed human healthy donor samples and NSCLC lung specimens. A.P. contributed critical reagents and helped with experimental design and analysis with the NSCLC lung specimens. D.V.S. and D.J.C. performed the allergic airway model experiments. D.B.C. contributed critical reagents and helped with experimental design and analysis for the allergic airway model experiments. C.J.W. contributed to experimental design, analysis and developing mouse strains. All authors provided feedback and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
D.A.A.V. and C.J.W. have submitted patents covering IL-35 that are pending and are entitled to a share in net income generated from licensing of these patent rights for commercial development.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Distribution and characterization of IL-10- and IL-35-expressing Treg cells in inflammatory environments.
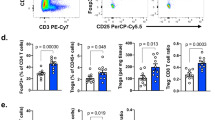
a, Validation of fluorescent reporters by real-time RT-PCR. Splenic Treg cell subpopulations were sorted from Il10GFP.Ebi3Tom.Foxp3Cre-YFP mice, and Il10 and Ebi3 transcripts were analyzed. Relative fold change expression was determined by normalizing to GFP–Tom– double-negative Treg cells. Data averaged from 3 independent experiments. Statistical significance was determined by One-way ANOVA with Holm-Sidak multiple comparisons (P values as indicated). b, Representative flow plots depicting the expression of IL-10 and Ebi3 in Treg cells isolated from the indicated tissues of 8wks old naive Il10GFP.Ebi3Tom.Foxp3Cre-YFP mice (top). Stacked-bar graph depicting the percent distribution of subpopulations (bottom). Recirculating (CD62L+/–Nrp1+) and de novo (CD62L+Nrp1–) Treg cells for Thymus. Data were averaged from 2 independent experiments with n=5 mice per tissue, except for thymus, blood, and lamina propria (n = 3). Error bars represent s.e.m. Statistical significance was determined by comparing tissues to de novo Thymus with Two-way ANOVA with Holm-Sidak multiple comparisons (****p<0.0001 and other P values as indicated). c, Representative flow plots depicting expression of IL-10+ and Tom+ cells within Foxp3-YFP+ Treg cells isolated from day 14 B16 tumor-bearing Il10GFP.Ebi3L/L-Tom.Foxp3Cre-YFP mice. CD4+YFP+ Treg cells were assessed for the expression of IL-10 and Tom in NDLN, DLN and TIL. d, SPICE plots depicting co-expression pattern of IL-10 and Tom on TIL Treg cells from B16 tumor-bearing Il10GFP.Ebi3L/L-Tom.Foxp3Cre-YFP mice as in (c). Data averaged from 3 independent experiments with n=9 mice total. e, Scatter-bar plot depicting percent distribution of Treg cell subpopulations as in (c). Bars represent mean values. f, Representative flow plots depicting the distribution of IL-10+ and Tom+ cells within Foxp3-YFP+ Treg cells isolated from d14 allergic inflammation induced Il10GFP.Ebi3L/L-Tom.Foxp3Cre-YFP mice. g, SPICE plots depicting co-expression pattern of IL-10 and Tom on Treg cells as described in (f). Data averaged from 3 independent experiments with n=10 mice total. h, Scatter-bar plot depicting percent distribution of Treg cell subpopulations as in (f). Bars represent mean values. (e and h) Statistical significance was determined by Two-way ANOVA with Holm-Sidak multiple comparisons (****p<0.0001 and other P values as indicated).
Supplementary Figure 2 Substantial TCR clonal overlap among intratumoral Treg cell subpopulations, consistent with lack of distinct transcriptomic signatures.
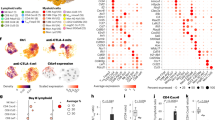
a, scRNAseq tSNE plots of bulk Treg cells from naive LNs and day 14 B16 Tumors (TIL Treg cells) from Foxp3Cre-YFP mice, with unsupervised K-means clustering (left) and a heat map depicting the top 10 differentially expressed genes for each of the 6 clusters identified (right). b, Stacked-bar graph depicting the percent distribution of Treg cell subpopulations within each cluster identified as in (a). c, Clonality was determined by examining the degree of TCR diversity within each population by TCRseq. Productive CDR3 sequences were used to define the TCR diversity. The indicated Teff control and Treg cell subpopulations were FACS sorted from day 14 B16 tumor-bearing Il10GFP.Ebi3Tom.Foxp3Cre-YFP mice for genomic DNA purification. Bars represent mean values. Statistical significance was determined by One-way ANOVA with with Holm-Sidak multiple comparisons (****p<0.0001). d, Line graphs representing the mean length of productive CDR3 locus sequence comparing the Treg cells subpopulations and effector T cells. Error bars represent s.e.m. e, TCR clonal similarity was determined by Morisita-Horn similarity index cross-examining the productive CDR3 sequences of Teff control and Treg cell subpopulations. Representative of three independent mouse samples is shown. (c-e) Data combined from TCR-sequencing data from three separate, individual Il10GFP.Ebi3Tom.Foxp3Cre-YFP mice. Error bars represent s.e.m. f, Representative flow plots depicting the BLIMP1YFP expression level plotted against CD4. (left): BLIMP1YFP– and BLIMP1YFP+ fractions of CD4+ TILs were purified by FACS sorting from d14 B16 tumor-bearing Prdm1YFP transgenic mice, followed by intracellular Foxp3 staining of the sorted cells. Data averaged from three sets of tumors with three tumors per set pooled for sorting. (right): Scatter plot depicting the percent distribution of BLIMP1YFP+ Teffs and Treg cells.
Supplementary Figure 3 Global transcriptomic analysis of intratumoral Treg cell subpopulations.
a, (left) Representative gating strategy for Treg cell subpopulation sorting for RNAseq. Each population was double-sorted. (right) PCA plot demonstrating the segregation of naive LN, NDLN, and B16 tumor derived TIL Treg cell subpopulations isolated from B16 tumor-bearing Il10GFP.Ebi3Tom.Foxp3Cre-YFP mice defined by their IL-10 and IL-35 expression pattern (IL-10–Ebi3–, IL-10+Ebi3–, and IL-10–Ebi3+) as well as naive LN effector T cell control (left PCA). A separate PCA plot was generated depicting only TIL Treg cell populations (right PCA). Data generated from three independent sequencing runs. Samples pooled from 4 mice per experiment to sort Treg cell subpopulations. Each dot in PCA plots represent individual sequencing run. For statistical analysis q-value FDR of 0.05 was used to determine the confidence of adjusted P values. b, Heat map of top 5,000 differentially expressed genes representing cross-examination of Treg cell subpopulations as in (a). c, Heat map of differentially expressed co-signaling molecules contrasting naive LN Teff cells and Treg cell subpopulations. Only genes that showed significant overall changes are listed. d, Representative flow plots depicting the expression of CD226 and CD27. Scatter-bar graphs demonstrating Treg cells differentially distributed based on the reciprocal expression of CD226 and CD27. e, Representative flow plots depicting the expression of TIM3 and LAG3. Scatter-bar graphs with demonstrating Treg cells differentially distributed between TIM3 single positive and TIM3+LAG3+ expression pattern. (d-e) Data averaged from 2 independent experiments with n=7 mice. Statistical significance was determined by Two-way ANOVA with Holm-Sidak multiple comparisons (****p<0.0001 and other P values indicated). f, Microsuppression assay comparing the suppressive function of Treg cell subpopulations (top: NDLN Treg cells, bottom: TIL Treg cells). Representative half-offset histograms show the proliferation of CD4+Foxp3– Teff responder (Tresponder) cells assessed by the decay of Cell Trace Violet proliferation dye at 1:4 Treg:Tresponder ratio. Line graphs depicting the summary of 3 independent experiments. Two-way ANOVA with Holm-Sidak multiple comparisons (P values indicated in the graphs).
Supplementary Figure 4 Modulation of inhibitory receptor expression in CD4+ and CD8+ TILs by Treg cell-derived IL-10 and IL-35.
a, Representative histograms depicting PD-1 expression on B16 tumor-infiltrating CD8+ TILs from Foxp3Cre-YFP (n=13), Il10L/L.Foxp3Cre-YFP (n=9), Ebi3L/L-Tom.Foxp3Cre-YFP (n=11), and Il10L/L.Ebi3L/L-Tom.Foxp3Cre-YFP (n=10) mice at day 14 post-tumor inoculation. Scatter-bar graphs depicting percent distribution of PD-1neg, PD-1int, and PD-1hi CD8+ TILs. b, Representative histograms depicting the PD-1 expression on CD4+Foxp3– T cells isolated from B16 tumor-bearing mice as in (a). Scatter-bar graphs depicting percent distribution of PD-1neg, PD-1int, and PD-1hi CD4+Foxp3– Teff cells. (a-b) Data averaged from 3 independent experiments. Statistical significance was determined by One-way ANOVA with Holm-Sidak multiple comparison (****p<0.0001 and other P values as indicated). c, Representative flow plots depicting the expression of inhibitory receptors (PD-1, TIM3, LAG3, TIGIT, and 2B4) on CD8+ T cells from day 20 B16 tumor-bearing Foxp3Cre-YFP (n=15), Il10L/L.Foxp3Cre-YFP (n=20), Ebi3L/L-Tom.Foxp3Cre-YFP YFP (n=17), and Il10L/L.Ebi3L/L-Tom.Foxp3Cre-YFP YFP (n=13) mice. SPICE plots depicting co-expression pattern of multiple inhibitory receptors. d, Scatter-bar graphs representing the percent distribution of inhibitory receptor-negative (0 & 1 inhibitory receptor-expressing effector-like) and multi-inhibitory receptor+ (3-5 inhibitory receptor-expressing exhausted) CD8+ T cells as in (c). e, Representative flow plots depicting expression of inhibitory receptors (PD-1, TIM3, LAG3, TIGIT, and 2B4) on CD4+Foxp3– Teff cells from day 20 B16 tumor-bearing mice as in (c). SPICE plots depicting co-expression pattern of multiple inhibitory receptors. f, Scatter-bar graphs representing the percent distribution of inhibitory receptor-negative (0 & 1 inhibitory receptor-expressing effector-like) and multi-inhibitory receptor+ (3–5 inhibitory receptor-expressing exhausted) CD4+Foxp3– TILs as in (e). Statistical significance was determined by One-way ANOVA with Holm-Sidak multiple comparison (****p<0.0001 and other P values as indicated).
Supplementary Figure 5 Differential regulation of the tumor microenvironment by Treg cell-derived IL-10 and IL-35.
a, Scatter-bar graphs depicting the absolute number of CD8+ TILs normalized to tumor volumes (either total, 0inhibitory receptor+, or 5inhibitory receptors+) isolated from day 14 B16 tumor-bearing Foxp3Cre-YFP (n=8), Il10L/L.Foxp3Cre-YFP (n=9), Ebi3L/L-Tom.Foxp3Cre-YFP, (n=10) and Il10L/L.Ebi3L/L-Tom.Foxp3Cre-YFP (n=12) mice. b, Scatter-bar graphs depicting absolute number of CD8+ TILs (either total CD8+ TILs, 0inhibitory receptor+, or 5inhibitory receptors+) normalized to tumor volume, CD8/Treg cell ratio, and Brdu+Ki67+ proliferating CD8+ TILs from day 20 B16 tumor-bearing Foxp3Cre-YFP (n=15), Il10L/L.Foxp3Cre-YFP (n=20), Ebi3L/L-Tom.Foxp3Cre-YFP (n=17), and Il10L/L.Ebi3L/L-Tom.Foxp3Cre-YFP (n=13) mice. (a-b) Data averaged from 3 independent experiments. Bars represent mean values. Statistical significance was determined by One-way ANOVA with Holm-Sidak multiple comparisons (****p<0.0001 and other P values as indicated). c, Representative histograms depicting expression of indicated markers on splenic and tumor-infiltrating DCs from B16 tumor-bearing Foxp3Cre-YFP (n=6), Il10L/L.Foxp3Cre-YFP (n=8), Ebi3L/L-Tom.Foxp3Cre-YFP (n=6), and Il10L/L.Ebi3L/L-Tom.Foxp3Cre-YFP (n=9) mice. d, Scatter-bar graphs depicting the Mean Fluorescent Intensity (MFI) of the markers examined as in (c). e, Scatter-bar graphs depicting the percent distribution of indicated markers on splenic and tumor-infiltrating DCs examined as in (c). f, Scatter-bar graphs demonstrating the M1 vs M2 subpopulation ratio within the tumor associated macrophage (TAM) population. Statistical significance was determined by One-way ANOVA with Holm-Sidak multiple comparisons (P values as indicated). (c-f) Data averaged from 2 independent experiments. Bars represent mean values. (c-e) Statistical significance was determined by Two-way ANOVA with Holm-Sidak multiple comparisons (****p<0.0001 and other p-values as indicated). g, Representative flow plots depicting the expression of inhibitory receptors and SPICE plots showing the multi-inhibitory receptor co-expression pattern on the donor CD8+ T cells infiltrating B16 tumor-bearing reconstituted Rag1–/– mice 14 days post-tumor inoculation as described in Fig. 5a. h, Scatter-bar graphs representing the percent distribution of donor CD8+ TILs based on the number of inhibitory receptors expressed. (WT: n=10, IL-10R–/–: n=6, IL-35R–/–: n=7). Bars represent mean values. Statistical significance was determined by One-way ANOVA with Holm-Sidak multiple comparisons (****p<0.0001 and other P values indicated). (g-h) Data averaged from 2 independent experiments.
Supplementary Figure 6 Transcriptional analysis of CD8+ TIL subsets from IL-10/IL-35 Treg cell-deficient tumor microenvironments.
a, Double-sorting scheme used to fractionate the CD8+ TIL subsets from the control Foxp3Cre-YFP and Treg cell cytokine-deficient environments into the NEG, INT, SP and DP fractions. Total CD8+ T cells were sorted from the NDLN for each of the 4 genotypes. Data represent at least 5 independent RNAseq experiments. b, PCA depicting the NDLN and CD8+ TILs subsets (NEG, INT, SP and DP) isolated from day 14 B16 tumor-bearing Foxp3Cre-YFP (n=3), Il10L/L.Foxp3Cre-YFP (n=2), Ebi3L/L-Tom.Foxp3Cre-YFP (n=3), and Il10 L/L.Ebi3L/L-Tom.Foxp3Cre-YFP (n=4) mice. Each of symbol represent an independent RNAseq run replicate. c, Heat map of top 5,000 differentially expressed genes representing CD8+ TILs from NDLN and NEG, INT, SP, and DP subsets. Genes were considered differentially expressed if their q-value FDR was below 0.05. d, Volcano plots contrasting CD8+ TIL subpopulations (NEG = PD-1neg, INT = PD-1int, SP = PD-1hiTIM3–, DP = PD-1hiTIM3+) isolated from Il10L/L.Foxp3Cre-YFP, Ebi3L/L-Tom.Foxp3Cre-YFP, and Il10L/L.Ebi3L/L-Tom.Foxp3Cre-YFP mice against the corresponding populations from Foxp3Cre-YFP control mice. Colored dots represent genes with q-value false discovery rate of <0.05. e, GSEA plot depicting the comparison of the exhaustion program in chronic LCMV infection versus the tumor exhaustion signature (derived by PD-1hi to PD-1neg CD8+ T cell comparison), with P value as indicated. f, GSEA plot depicting the gene signature of PD-1 checkpoint blockade responsive TCF-1+CXCR5+ CD8+ T cell subset in SP CD8+ TILs (left) and DP CD8+ TILs (right) from cytokine-deficient Treg cell environments (Il10L/L.Foxp3Cre-YFP, Ebi3L/L-Tom.Foxp3Cre-YFP, and Il10L/L.Ebi3L/L-Tom.Foxp3Cre-YFP); P values as indicated. (e-f) Statistical significance was determined by the rankSumTestWithCorrelation test implemented in the limma package with correction for loss in degrees of freedom due to correlation among genes. All P values are two sided. No correction for multiple hypothesis was performed. (a-f) Data are averaged from at least 5 independent RNASeq experiments with genotype-specific CD8+ T cell replicates as indicated - Foxp3Cre-YFP (n=3), Il10L/L.Foxp3Cre-YFP (n=2), Ebi3L/L-Tom.Foxp3Cre-YFP (n=3), and Il10 L/L.Ebi3L/L-Tom.Foxp3Cre-YFP CD8s (n=4).
Supplementary Figure 7 IL-10+ and IL-35+ Treg cells drive BLIMP1 induction on TILs in vivo.
a, Representative flow plots depicting the expression of BLIMP1YFP expression levels in NDLN and TIL CD8+ T cells from B16 tumor-bearing Prdm1YFP transgenic reporter mice (n=13). CD8+ TILs representative flow and SPICE plots demonstrating the expression of inhibitory receptors on the BLIMP1YFPhi, YFPint and YFPneg CD8+ T cell fractions. b, Scatter plots showing the percent distribution of YFPhi subpopulation among CD8+ T cells as in (a). Statistical significance was determined by paired two-tailed Student’s t-test (****p<0.0001). c, Scatter-bar graphs depicting the percent distribution of CD8+ TILs based on the number of inhibitory receptors expressed as in (a). Bars represent mean values. Statistical significance was determined by One-way ANOVA with Holm-Sidak multiple comparison (****p<0.0001 and other P values as indicated). (a-c) Data averaged from 3 independent experiments, with n=13 mice/group. d, Representative flow plots depicting the protein level expression pattern of PD-1 and BLIMP1 on CD8+ T cells from B16 tumor-bearing Foxp3Cre-YFP (n=14), Il10L/L.Foxp3Cre-YFP (n=11), Ebi3L/L-Tom.Foxp3Cre-YFP (n=12) and Il10L/L.Ebi3L/L-Tom.Foxp3Cre-YFP (n=14) mice, with stacked bar graph depicting percent distribution of PD-1hi, PD-1int, and PD-1neg CD8+ T cells co-expressing PD-1 and BLIMP1. Data averaged from 3 independent experiments. e, Scatter-bar graphs representing the percent distribution of CD8+ T cells expressing 0-5 inhibitory receptors from B16 tumor-bearing E8ICre-GFP (n=9), Prdm1L/+.E8ICre-GFP (n=12), and Prdm1L/L.E8ICre-GFP (n=10) mice as in Fig. 7d. Data averaged from 3 independent experiments. Bars represent mean values. Statistical significance was determined by One-way ANOVA with Holm-Sidak multiple comparison (****p<0.0001 and other P values as indicated). g, Graphical summary of the regulatory functions of Treg-derived IL-10 and IL35 depicted by their common and non-redundant effects on TILs. Treg-derived IL35 predominantly regulated IR-expression and TCM differentiation while Treg-derived IL-10 played a greater role in limiting cytokine production and proliferation of CD8+ TILs.
Supplementary information
Supplementary Information
Supplementary Figures 1–7
Rights and permissions
About this article
Cite this article
Sawant, D.V., Yano, H., Chikina, M. et al. Adaptive plasticity of IL-10+ and IL-35+ Treg cells cooperatively promotes tumor T cell exhaustion. Nat Immunol 20, 724–735 (2019). https://doi.org/10.1038/s41590-019-0346-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-019-0346-9
This article is cited by
-
MicroRNAs as regulators of immune checkpoints in cancer immunotherapy: targeting PD-1/PD-L1 and CTLA-4 pathways
Cancer Cell International (2024)
-
Targeting inflammation as cancer therapy
Journal of Hematology & Oncology (2024)
-
Nanoparticles in tumor microenvironment remodeling and cancer immunotherapy
Journal of Hematology & Oncology (2024)
-
Regulatory T cell-mediated immunosuppression orchestrated by cancer: towards an immuno-genomic paradigm for precision medicine
Nature Reviews Clinical Oncology (2024)
-
Stroma-infiltrating T cell spatiotypes define immunotherapy outcomes in adolescent and young adult patients with melanoma
Nature Communications (2024)