Abstract
The fetus is thought to be protected from exposure to foreign antigens, yet CD45RO+ T cells reside in the fetal intestine. Here we combined functional assays with mass cytometry, single-cell RNA sequencing and high-throughput T cell antigen receptor (TCR) sequencing to characterize the CD4+ T cell compartment in the human fetal intestine. We identified 22 CD4+ T cell clusters, including naive-like, regulatory-like and memory-like subpopulations, which were confirmed and further characterized at the transcriptional level. Memory-like CD4+ T cells had high expression of Ki-67, indicative of cell division, and CD5, a surrogate marker of TCR avidity, and produced the cytokines IFN-γ and IL-2. Pathway analysis revealed a differentiation trajectory associated with cellular activation and proinflammatory effector functions, and TCR repertoire analysis indicated clonal expansions, distinct repertoire characteristics and interconnections between subpopulations of memory-like CD4+ T cells. Imaging mass cytometry indicated that memory-like CD4+ T cells colocalized with antigen-presenting cells. Collectively, these results provide evidence for the generation of memory-like CD4+ T cells in the human fetal intestine that is consistent with exposure to foreign antigens.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Mass cytometry data are available via Flow Repository (https://flowrepository.org/id/FR-FCM-ZYRD). Single-cell RNA-seq data are available via Gene Expression Omnibus accession code GSE122846. The remaining data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Janeway, C. A. Jr. Approaching the asymptote? Evolution and revolution in immunology. Cold Spring Harb. Symp. Quant. Biol. 54(Pt 1), 1–13 (1989).
Fan, X. & Rudensky, A. Y. Hallmarks of tissue-resident lymphocytes. Cell 164, 1198–1211 (2016).
Thome, J. J. C. et al. Early-life compartmentalization of human T cell differentiation and regulatory function in mucosal and lymphoid tissues. Nat. Med. 22, 72–77 (2015).
Bunders, M. J. et al. Memory CD4(+)CCR5(+) T cells are abundantly present in the gut of newborn infants to facilitate mother-to-child transmission of HIV-1. Blood 120, 4383–4390 (2012).
Zhang, X. et al. CD4 T cells with effector memory phenotype and function develop in the sterile environment of the fetus. Sci. Transl. Med. 6, 238ra72 (2014).
Stout, M. J. et al. Identification of intracellular bacteria in the basal plate of the human placenta in term and preterm gestations. Am. J. Obstet. Gynecol. 208, 226.e1–7 (2013).
Aagaard, K. et al. The placenta harbors a unique microbiome. Sci. Transl. Med. 6, 237ra65 (2014).
Collado, M. C., Rautava, S., Aakko, J., Isolauri, E. & Salminen, S. Human gut colonisation may be initiated in utero by distinct microbial communities in the placenta and amniotic fluid. Sci. Rep. 6, 23129 (2016).
DiGiulio, D. B. Diversity of microbes in amniotic fluid. Semin. Fetal Neonatal Med. 17, 2–11 (2012).
Ardissone, A. N. et al. Meconium microbiome analysis identifies bacteria correlated with premature birth. PLoS ONE 9, e90784 (2014).
Hornef, M. & Penders, J. Does a prenatal bacterial microbiota exist? Mucosal Immunol. 10, 598–601 (2017).
Li, N. et al. Mass cytometry reveals innate lymphoid cell differentiation pathways in the human fetal intestine. J. Exp. Med. 215, 1383–1396 (2018).
van Unen, V. et al. Visual analysis of mass cytometry data by hierarchical stochastic neighbour embedding reveals rare cell types. Nat. Commun. 8, 1740 (2017).
Maaten, Lvander & Hinton, G. Visualizing data using t-SNE. J. Mach. Learn. Res. 9, 2579–2605 (2008).
Höllt, T. et al. Cytosplore: interactive immune cell phenotyping for large single-cell datasets. Comput. Graph. Forum 35, 171–180 (2016).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Kimmig, S. et al. Two subsets of naive T helper cells with distinct T cell receptor excision circle content in human adult peripheral blood. J. Exp. Med. 195, 789–794 (2002).
Angerer, P. et al. destiny: diffusion maps for large-scale single-cell data in R. Bioinformatics 32, 1241–1243 (2015).
Samusik, N., Good, Z., Spitzer, M. H., Davis, K. L. & Nolan, G. P. Automated mapping of phenotype space with single-cell data. Nat. Methods 13, 493–496 (2016).
Jolliffe, I. Principal component analysis. in International Encyclopedia of Statistical Science (ed. Lovric, M) Ch. 427, 1094–1096 (Springer, Berlin, 2011).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014).
Qiu, X. et al. Single-cell mRNA quantification and differential analysis with Census. Nat. Methods 14, 309–315 (2017).
Azzam, H. S. et al. CD5 expression is developmentally regulated by T cell receptor (TCR) signals and TCR avidity. J. Exp. Med. 188, 2301–2311 (1998).
Mandl, J. N., Monteiro, J. P., Vrisekoop, N. & Germain, R. N. T. Cell-positive selection uses self-ligand binding strength to optimize repertoire recognition of foreign antigens. Immunity 38, 263–274 (2013).
Persaud, S. P., Parker, C. R., Lo, W.-L., Scott Weber, K. & Allen, P. M. Intrinsic CD4 T cell sensitivity and response to a pathogen are set and sustained by avidity for thymic and peripheral complexes of self peptide and MHC. Nat. Immunol. 15, 266–274 (2014).
Fulton, R. B. et al. The TCR’s sensitivity to self peptide-MHC dictates the ability of naive CD8(+) T cells to respond to foreign antigens. Nat. Immunol. 16, 107–117 (2015).
Shugay, M. et al. VDJtools: unifying post-analysis of T cell receptor repertoires. PLoS Comput. Biol. 11, e1004503 (2015).
Egorov, E. S. et al. The changing landscape of naive T cell receptor repertoire with human aging. Front. Immunol. 9, 1618 (2018).
Miyazawa, S. & Jernigan, R. L. Residue-residue potentials with a favorable contact pair term and an unfavorable high packing density term, for simulation and threading. J. Mol. Biol. 256, 623–644 (1996).
Kidera, A., Konishi, Y., Oka, M., Ooi, T. & Scheraga, H. A. Statistical analysis of the physical properties of the 20 naturally occurring amino acids. J. Protein Chem. 4, 23–55 (1985).
Rackovsky, S. Global characteristics of protein sequences and their implications. Proc. Natl Acad. Sci. USA 107, 8623–8626 (2010).
Pezzotti, N. et al. Approximated and user steerable tSNE for progressive visual analytics. IEEE Trans. Vis. Comput. Graph. 23, 1739–1752 (2017).
Fergusson, J. R. et al. CD161 defines a transcriptional and functional phenotype across distinct human T cell lineages. Cell Rep. 9, 1075–1088 (2014).
Edfors, F. et al. Gene-specific correlation of RNA and protein levels in human cells and tissues. Mol. Syst. Biol. 12, 883 (2016).
McFarland, R. D., Douek, D. C., Koup, R. A. & Picker, L. J. Identification of a human recent thymic emigrant phenotype. Proc. Natl Acad. Sci. USA 97, 4215–4220 (2000).
Staton, T. L. et al. CD8+ recent thymic emigrants home to and efficiently repopulate the small intestine epithelium. Nat. Immunol. 7, 482–488 (2006).
Michaelsson, J., Mold, J. E., McCune, J. M. & Nixon, D. F. Regulation of T cell responses in the developing human fetus. J. Immunol. 176, 5741–5748 (2006).
Feng, Y. et al. A mechanism for expansion of regulatory T-cell repertoire and its role in self-tolerance. Nature 528, 132–136 (2015).
Allenspach, E. J. et al. Absence of functional fetal regulatory T cells in humans causes in utero organ-specific autoimmunity. J. Allergy Clin. Immunol. 140, 616–619.e7 (2017).
Gomez de Agüero, M. et al. The maternal microbiota drives early postnatal innate immune development. Science 351, 1296–1302 (2016).
van Unen, V. et al. Mass cytometry of the human mucosal immune system identifies tissue- and disease-associated immune subsets. Immunity 44, 1227–1239 (2016).
Zheng, G. X. Y. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Chen, J., Bardes, E. E., Aronow, B. J. & Jegga, A. G. ToppGene Suite for gene list enrichment analysis and candidate gene prioritization. Nucleic Acids Res. 37, W305–W311 (2009).
Bean, D. M. et al. esyN: network building, sharing and publishing. PLoS ONE 9, e106035 (2014).
Ogata, H. et al. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 27, 29–34 (1999).
Quigley, M. F., Almeida, J. R., Price, D. A. & Douek, D. C. Unbiased molecular analysis of T cell receptor expression using template-switch anchored RT–PCR. Curr. Protoc. Immunol. 94, 10.33.1–10.33.16 (2011).
Lefranc, M.-P. et al. IMGT unique numbering for immunoglobulin and T cell receptor constant domains and Ig superfamily C-like domains. Dev. Comp. Immunol. 29, 185–203 (2005).
Shugay, M. et al. Towards error-free profiling of immune repertoires. Nat. Methods 11, 653–655 (2014).
Bolotin, D. A. et al. MiXCR: software for comprehensive adaptive immunity profiling. Nat. Methods 12, 380–381 (2015).
Acknowledgements
We thank the Center for Contraception, Abortion and Sexuality (Leiden and The Hague) for collection and provision of fetal material, K. Lodder, T. van Herwaarden, M. Bialecka and F. Wang for dissection of fetal tissues, and S. L. Kloet for assistance with single-cell RNA sequencing. This research was supported by Leiden University Medical Center (N.L., V.v.U., N.G., F.K.), the Netherlands Organization for Scientific Research (NWO Applied Technical Sciences grant no. 12721 and ZonMW grant no. 91112008) (T.A., T.H., J.E., B.P.F.L.), the Russian Science Foundation (grant no. 16-15-00149) (S.A.K., E.S.E., M.I., O.V.B, D.M.C.), the Wellcome Trust (grant no. 100326/Z/12/Z) (K.L., J.E.M., D.A.P), the European Commission under an MSCA-ITN award (grant no. 675743/ISPIC) (T.A.), and the China Scholarship Council (N.L. and N.G.).
Author information
Authors and Affiliations
Contributions
N.L., V.v.U. and F.K. conceived the study and wrote the manuscript. N.L. performed most of the experiments with help from V.v.U. and N.G. N.L. performed most of the data analyses with help from V.v.U., T.A. and B.P.F.L. S.A.K., K.L., J.E.M., E.S.E., M.I., D.M.C., O.V.B. and D.A.P. performed TCR repertoire analyses. D.A.P. revised the manuscript. N.F.C.C.d.M. helped with imaging mass cytometry experiments. T.H., V.v.U., J.E. and B.P.F.L. developed Cytosplore. S.M.C.d.S.L. provided human fetal tissues. All authors discussed the results and helped prepare the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Mass cytometric analysis of fetal intestinal immune cells.
(a) Biaxial plots from one fetal intestine showing the gating strategy for single, live, CD45+ cells analyzed by mass cytometry (n = 7). (b) First-level HSNE embedding of CD45+ immune cells (n = 224,286) derived from fetal intestines (n = 7). Each dot represents a landmark. The size of each landmark is proportional to the number of cells that the landmark represents. Colors indicate ArcSinh5-transformed expression values. (c) t-SNE embedding showing all CD4+ T cells (n = 110,332) derived from fetal intestines (n = 7). Colors indicate the ArcSinh5-transformed expression values.
Supplementary Figure 2 Mass cytometric analysis of the fetal CD4+ T cell compartment across tissues.
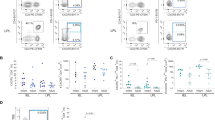
(a) t-SNE embedding of all CD4+ T cells (n = 9.7 × 104) derived from fetal intestines (n = 6.6 × 104 cells from 7 samples), fetal livers (n = 1,530 cells from 3 samples), and fetal spleens (n = 3.0 × 104 cells from 3 samples). Colors indicate tissue origin. Downsampling was performed for samples with more than 10,000 CD4+ T cells. (b) Bar graphs showing the quantification of Treg cells and TN cells, TCM cells, and TEM cells among fetal CD4+ T cells across tissues. TN cells: CD45RA+CCR7+; TCM cells: CD45RA-CCR7+; TEM cells: CD45RA-CCR7−. Fetal liver and spleen, n = 3 independent samples; fetal intestine, n = 7 independent samples. Error bars indicate mean ± SEM. **p < 0.01, Kruskal–Wallis test with Dunn’s test for multiple comparisons.
Supplementary Figure 3 Identification of CD4+ T cell clusters in fetal intestines.
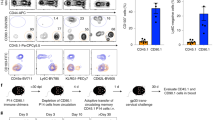
(a) Representative biaxial plots showing the gating strategy for TN, CD161− TM, CD161+CD117− TM, CD161+CD117+ TM and Treg cell clusters derived from one fetal intestine analyzed by flow cytometry for expression of CD3, CD4, CD8a, CD25, CD45RA, CD45RO, CD117, CD127, CD161 and TCRγδ (n = 10). (b) Expression (log-normalized) of KIT as determined by single-cell RNA-seq analysis, presented as violin plots. Colors indicate different cell clusters. (c) Bar plot depicting cell frequencies for clusters identified by mass cytometry and RNA-seq.
Supplementary Figure 4 Single-cell trajectories of fetal intestinal CD4+ T cells.
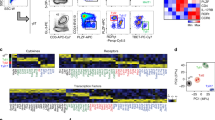
(a-b) t-SNE embeddings of all fetal intestinal CD4+ T cells analyzed in (a) the mass cytometry dataset (n = 10,436 cells, 35 proteins) and (b) the RNA-seq dataset (n = 1,743 cells, 300 variable genes) at the middle of the t-SNE computation. Colors indicate marker expression. (c) Diffusion map and (d) VorteX analysis of all fetal intestinal CD4+ T cells analyzed in mass cytometry dataset (n = 10,436 cells, 35 proteins). Colors indicate different cell clusters. (e) Diffusion map and (f) PCA analysis of all fetal intestinal CD4+ T cells analyzed in the RNA-seq dataset (n = 1,743 cells, 2,174 variable genes). Colors indicate different cell clusters.
Supplementary Figure 5 Immune-related genes in the pseudotime-dependent gene module.
(a) Table listing 106 immune-related genes kinetically enriched in the pseudotime-dependent third gene module (Fig. 4d). (b) Interaction network of the pseudotime-dependent third gene module using Esyn. Orange lines indicate literature-described physical interactions of the gene-encoded proteins. Orange nodes indicate genes enriched in immunological processes.
Supplementary Figure 6 TCR sequencing of distinct CD4+ T cell populations.
(a) Dot plot showing the percentage of reads per unique TCRβ sequence in each cluster. Number of unique TCRβ sequences per cluster: TN cells, 59; CD161- TM cells, 54; CD161+CD117- TM cells, 37; CD161+CD117+ TM cells, 51; Naive Treg cells, 29; Memory Treg cells, 50. (b) Hierarchical clustering of CD4+ T subpopulations based on individual V-J rearrangements in the TCRα (TRA) and TCRβ (TRB) datasets. Colors indicate different fetal intestines. (c) Dot plot showing weighted clonal overlaps for TCRβ amino acid sequences among clusters from two different fetal intestines analyzed using the F2 similarity metric in VDJtools. (d) Chord diagram showing clonal overlaps among different cell clusters. Numbers in the yellow circle indicate the total numbers and frequencies (parentheses) of private TCRβ clonotypes among TN cells. Numbers outside the plot border indicate the total numbers and frequencies (parentheses) of unique TCRβ clonotypes shared among TN cells and other CD4+ T cell subpopulations. (a) Conventional sequencing, (b-d) high-throughput sequencing.
Supplementary Figure 7 Cytokine production by fetal intestinal CD4+ T cells.
(a-c) Purified fetal intestinal CD4+ T cells were treated with a control antibody or stimulated with anti-CD3 and anti-CD28 for 4 h. Intracellular expression of TNF, IFN-γ, IL-2, IL-4 and Ki-67 was determined for each subpopulation by flow cytometry. (a) Dot plot showing the median fluorescence intensity (MFI) of TNF for each cell cluster relative to corresponding CD117+ TM subpopulation in each fetal intestine (n = 4) after stimulation with anti-CD3 and anti-CD28. Data represent two independent experiments. (b) Representative biaxial plots showing coexpression of cytokines vs. Ki-67 for the indicated cell clusters after stimulation with anti-CD3 and anti-CD28. (c) Pie charts depicting coexpression profiles of IFN-γ and IL-2 for the indicated cell clusters from fetal intestines (n = 2) after stimulation with anti-CD3 and anti-CD28. Data represent two independent experiments.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Tables 1–4
Rights and permissions
About this article
Cite this article
Li, N., van Unen, V., Abdelaal, T. et al. Memory CD4+ T cells are generated in the human fetal intestine. Nat Immunol 20, 301–312 (2019). https://doi.org/10.1038/s41590-018-0294-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-018-0294-9
This article is cited by
-
The maternal gut microbiome in pregnancy: implications for the developing immune system
Nature Reviews Gastroenterology & Hepatology (2024)
-
High-dimensional analysis reveals an immune atlas and novel neutrophil clusters in the lungs of model animals with Actinobacillus pleuropneumoniae-induced pneumonia
Veterinary Research (2023)
-
Immune cell identifier and classifier (ImmunIC) for single cell transcriptomic readouts
Scientific Reports (2023)
-
Immune subset-committed proliferating cells populate the human foetal intestine throughout the second trimester of gestation
Nature Communications (2023)
-
CXCR5+PD-1++ CD4+ T cells colonize infant intestines early in life and promote B cell maturation
Cellular & Molecular Immunology (2023)