Abstract
Cyclic GMP-AMP synthase (cGAS) is a key sensor responsible for cytosolic DNA detection. Here we report that GTPase-activating protein SH3 domain–binding protein 1 (G3BP1) is critical for DNA sensing and efficient activation of cGAS. G3BP1 enhanced DNA binding of cGAS by promoting the formation of large cGAS complexes. G3BP1 deficiency led to inefficient DNA binding by cGAS and inhibited cGAS-dependent interferon (IFN) production. The G3BP1 inhibitor epigallocatechin gallate (EGCG) disrupted existing G3BP1–cGAS complexes and inhibited DNA-triggered cGAS activation, thereby blocking DNA-induced IFN production both in vivo and in vitro. EGCG administration blunted self DNA–induced autoinflammatory responses in an Aicardi–Goutières syndrome (AGS) mouse model and reduced IFN-stimulated gene expression in cells from a patient with AGS. Thus, our study reveals that G3BP1 physically interacts with and primes cGAS for efficient activation. Furthermore, EGCG-mediated inhibition of G3BP1 provides a potential treatment for cGAS-related autoimmune diseases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The MS-identified cGAS-interacting protein list is provided in Supplementary Tables 1 and 2. The primer sequences used for qPCR are provided in Supplementary Table 3. The information on antibodies used in our study is provided in Supplementary Table 4. We also provide full scans of all the blots and gels as Supplementary Dataset 1. The data that support the findings of this study are available from the corresponding authors upon reasonable request.
References
Takeuchi, O. & Akira, S. Pattern recognition receptors and inflammation. Cell 140, 805–820 (2010).
Wu, J. & Chen, Z. J. Innate immune sensing and signaling of cytosolic nucleic acids. Annu. Rev. Immunol. 32, 461–488 (2014).
Gurtler, C. & Bowie, A. G. Innate immune detection of microbial nucleic acids. Trends Microbiol. 21, 413–420 (2013).
Gao, P. et al. Cyclic [G(2′,5′)pA(3′,5′)p] is the metazoan second messenger produced by DNA-activated cyclic GMP-AMP synthase. Cell 153, 1094–1107 (2013).
Wu, J. et al. Cyclic GMP-AMP is an endogenous second messenger in innate immune signaling by cytosolic DNA. Science 339, 826–830 (2013).
Gao, P. et al. Structure-function analysis of STING activation by c[G(2′,5′)pA(3′,5′)p] and targeting by antiviral DMXAA. Cell 154, 748–762 (2013).
Morita, M. et al. Gene-targeted mice lacking the Trex1 (DNase III) 3′-->5′ DNA exonuclease develop inflammatory myocarditis. Mol. Cell. Biol. 24, 6719–6727 (2004).
Crow, Y. J. & Manel, N. Aicardi–Goutières syndrome and the type I interferonopathies. Nat. Rev. Immunol. 15, 429–440 (2015).
Aicardi, J. & Goutières, F. A progressive familial encephalopathy in infancy with calcifications of the basal ganglia and chronic cerebrospinal fluid lymphocytosis. Ann. Neurol. 15, 49–54 (1984).
Gao, D. et al. Activation of cyclic GMP-AMP synthase by self-DNA causes autoimmune diseases. Proc. Natl Acad. Sci. USA 112, E5699–E5705 (2015).
Gray, E. E., Treuting, P. M., Woodward, J. J. & Stetson, D. B. Cutting Edge: cGAS is required for lethal autoimmune disease in the Trex1-deficient mouse model of Aicardi–Goutières syndrome. J. Immunol. 195, 1939–1943 (2015).
Protter, D. S. & Parker, R. Principles and properties of stress granules. Trends Cell Biol. 26, 668–679 (2016).
Onomoto, K., Yoneyama, M., Fung, G., Kato, H. & Fujita, T. Antiviral innate immunity and stress granule responses. Trends Immunol. 35, 420–428 (2014).
Luecke, S. et al. cGAS is activated by DNA in a length-dependent manner. EMBO Rep. 18, 1707–1715 (2017).
Zekri, L. et al. Control of fetal growth and neonatal survival by the RasGAP-associated endoribonuclease G3BP. Mol. Cell. Biol. 25, 8703–8716 (2005).
Burdette, D. L. et al. STING is a direct innate immune sensor of cyclic di-GMP. Nature 478, 515–518 (2011).
Sun, L., Wu, J., Du, F., Chen, X. & Chen, Z. J. Cyclic GMP-AMP synthase is a cytosolic DNA sensor that activates the type I interferon pathway. Science 339, 786–791 (2013).
Gao, D. et al. Cyclic GMP-AMP synthase is an innate immune sensor of HIV and other retroviruses. Science 341, 903–906 (2013).
Yoh, S. M. et al. PQBP1 is a proximal sensor of the cGAS-dependent innate response to HIV-1. Cell 161, 1293–1305 (2015).
Gilks, N. et al. Stress granule assembly is mediated by prion-like aggregation of TIA-1. Mol. Biol. Cell. 15, 5383–5398 (2004).
Liang, Q. et al. Crosstalk between the cGAS DNA sensor and Beclin-1 autophagy protein shapes innate antimicrobial immune responses. Cell Host Microbe 15, 228–238 (2014).
Tourrière, H. et al. The RasGAP-associated endoribonuclease G3BP assembles stress granules. J. Cell. Biol. 160, 823–831 (2003).
Jonsson, K. L. et al. IFI16 is required for DNA sensing in human macrophages by promoting production and function of cGAMP. Nat. Commun. 8, 14391 (2017).
Almine, J. F. et al. IFI16 and cGAS cooperate in the activation of STING during DNA sensing in human keratinocytes. Nat. Commun. 8, 14392 (2017).
Tao, J. et al. Nonspecific DNA binding of cGAS N terminus promotes cGAS activation. J. Immunol. 198, 3627–3636 (2017).
Shim, J. H. et al. Epigallocatechin gallate suppresses lung cancer cell growth through Ras-GTPase-activating protein SH3 domain-binding protein 1. Cancer Prev. Res. 3, 670–679 (2010).
Guo, H. et al. Clinicopathological and genetic analysis of Aicardi-Goutières syndrome. Chinese J. Neurol. 47, 96–100 (2014).
Liu, Y. et al. Activated STING in a vascular and pulmonary syndrome. N. Engl. J. Med. 371, 507–518 (2014).
Clark, K., Plater, L., Peggie, M. & Cohen, P. Use of the pharmacological inhibitor BX795 to study the regulation and physiological roles of TBK1 and IkappaB kinase epsilon: a distinct upstream kinase mediates Ser-172 phosphorylation and activation. J. Biol. Chem. 284, 14136–14146 (2009).
Civril, F. et al. Structural mechanism of cytosolic DNA sensing by cGAS. Nature 498, 332–337 (2013).
Zhou, W. et al. Structure of the human cGAS-DNA complex reveals enhanced control of immune surveillance. Cell 174, 300–311.e11 (2018).
Andreeva, L. et al. cGAS senses long and HMGB/TFAM-bound U-turn DNA by forming protein-DNA ladders. Nature 549, 394–398 (2017).
Orzalli, M. H. et al. cGAS-mediated stabilization of IFI16 promotes innate signaling during herpes simplex virus infection. Proc. Natl Acad. Sci. USA 112, E1773–E1781 (2015).
Mackenzie, K. J. et al. cGAS surveillance of micronuclei links genome instability to innate immunity. Nature 548, 461–465 (2017).
Harding, S. M. et al. Mitotic progression following DNA damage enables pattern recognition within micronuclei. Nature 548, 466–470 (2017).
Wang, H. et al. cGAS is essential for the antitumor effect of immune checkpoint blockade. Proc. Natl. Acad. Sci. USA 114, 1637–1642 (2017).
Li, X. et al. Cyclic GMP-AMP synthase is activated by double-stranded DNA-induced oligomerization. Immunity 39, 1019–1031 (2013).
Wu, C. et al. Oxygen promotes biofilm formation of Shewanella putrefaciens CN32 through a diguanylate cyclase and an adhesin. Sci. Rep. 3, 1945 (2013).
Acknowledgements
We sincerely thank the patient with AGS and his sibling for donating PBMCs. We thank J. Tazi (Université Montpellier 2) for providing the G3bp1+/– mice, Cancer Research Technology Limited for providing the Trex1+/– mice, C. Widmann (University of Lausanne) for providing HA-G3BP1 plasmid, J. W. Chin (Cambridge) for providing pCDF PylT-1 plasmid, J. U. Jung (University of Southern California) for providing cGAS cDNA construct, H. Shu (Wuhan University) for providing STING cDNA construct, J. Han (Xiamen University) for providing HSV-1, M. Yang (Tsinghua University) for help with protein purification. We thank H. Yu (UT Southwestern Medical Center), Y. Zheng (Carnegie Institution for Science) and H. Qi (Tsinghua University) for helpful discussion and critical reading of the manuscript. This work was supported by grants from the China National Natural Science Foundation (No. 81771708 to T.L., No. 81521064 to X.-M.Z.).
Author information
Authors and Affiliations
Contributions
T.L. and X.-M.Z. supervised the project and acquired funding for the study. T.L., Z.-S.L., H.C. and W.X. designed the experiments. Z.-S.L., H.C., M.W., M.Z., S.C., W.X. and J.D. performed cell experiments, qPCR analysis and western blotting. Z.-S.L., H.C., X.P. and Y.-J.H. performed the mouse studies. H.C., Z.-Y.Z., Ti. L. and T.X. performed cGAS–DNA binding and cGAMP synthesis assays. X.W., A.-L.L., S.-M.W. and F.D. performed LC–MS/MRM analysis. W.X., T.Z. and Q.X. isolated MEFs and primary human cells. Z.-S.L. and W.-J.L. performed EMSA and protein cross-linking experiments. J.-Q.X. and X.L. performed virus infection experiments. Z.-S.L., H.C., W.X., M.W., Z.-G.L. and T.L. analyzed the data. T.L. and X.-M.Z. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 G3BP1 is critical for the DNA-triggered cGAS/STING pathway.
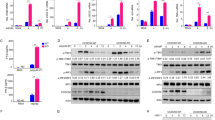
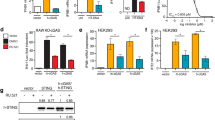
a, ELISA analysis of IFN-β production in U937 cells left untransfected (–) or transfected with HT-DNA (+) for 12 h. N.D., non-detected. b, qPCR analysis of IFNB mRNA expression in U937 cells transfected with the indicated amount of HT-DNA for 2 h. c,d, qPCR analysis of IFNB mRNA expression of WT and G3BP2–/– U937 cells transfected with HT-DNA (2 μg/ml) (c) or plasmid DNA (2 μg/ml) (d) for 2 h. e, Immunoblot analysis of G3BP2 expression. f, Immunoblot analysis of WT and G3bp1–/– primary MEF cells with the indicated antibodies. g, qPCR analysis of Ifnb mRNA expression in MEFs transfected with ISD (0.5 μg/ml) for the indicated time. h,i, Immunoblot analysis of U937 cells transfected with ISD (h) or plasmid DNA (i) with the indicated antibodies. j, Immunoblot analysis of MEFs transfected with HT-DNA with the indicated antibodies. k, qPCR analysis of IFNB mRNA expression in HT-DNA (2 μg/ml)-treated U937 cells carrying non-targeting control shRNA (sh-NC) or G3BP1-specific shRNA (sh-G3BP1) (left). G3BP1 expression was analyzed by immunoblotting (right). l, Immunoblot analysis of HeLa cells transfected with non-targeting control siRNA (si-NC) or G3BP1-specific siRNA (si-G3BP1), followed by transfection of HT-DNA (2 μg/ml) for 2 h. β-actin (e,f,h–l), loading control. **P < 0.01, ***P < 0.001, two-tailed t test (a,b,g,k). Data are representative of three experiments. Data are mean ± s.e.m. of quadruplicate samples in a or triplicate samples in b–d,g,k.
Supplementary Figure 2 Quantification of cGAMP.
a,b, Commercial cGAMP with the indicated concentrations was quantified with LC–MS/MRM. Data were fitted to generate the standard curve with GraphPad Prism using linear regression. c, cGAMP production of WT and G3bp1–/– MEFs transfected with HT-DNA (0.5 μg/ml) was analyzed by LC–MS/MRM, N.D., non-detected. d, qPCR analysis of Ifnb mRNA expression in WT and G3bp1–/– MEFs treated with cGAMP (0.5 μg/ml) for the indicated time. e, qPCR analysis of IFNB mRNA expression in WT and G3BP1–/– U937 cells transfected with c-di-GMP (4 μg/ml) for the indicated time. ***P < 0.001, two-tailed t test (c). Data are representative of three experiments, mean ± s.e.m. of triplicate samples in c–e.
Supplementary Figure 3 G3BP1 regulates cGAS-mediated antiviral responses.
a,d, ELISA analysis of secreted IFN-β (a) and qPCR analysis of specific RNA of HSV-1 (d) in MEFs left untreated (–) or infected with HSV-1 (MOI = 1) (+) for 24 h. b, qPCR analysis of the specific genomic DNA of HSV-1 in U937 cells left untreated (–) or infected with HSV-1 (MOI = 1) (+) for the indicated time (n = 2 independent experiments). c, Plaque assay shows HSV-1 titer in U937 cells left untreated (–) or infected with HSV-1 (MOI = 1) (+) for 24 h. e, qPCR analysis of ISG15 mRNA expression in U937 cells left untreated (–) or infected with HIV-1 (100 ng/ml by p24) (+) for 48 h (n = 2 independent experiments). f–h, qPCR analysis of Tnf mRNA expression in MEFs treated with Pam3CSK4 (1 μg/ml) (f), poly(I:C) (50 μg/ml) (g) or LPS (1 μg/ml) (h) for 2 h. **P < 0.01, ***P < 0.001, two-tailed t test (a,c,d). Data are representative of three experiments, mean ± s.e.m. of triplicate samples in a,c,d,f–h.
Supplementary Figure 4 SG and Beclin-1 are not involved in G3BP1-mediated cGAS activation.
a, Immunoblot analysis of G3BP1 expression in different tissues of mice. b,c, qPCR analysis of the indicated mRNA expression in U937 cells (b) and primary MEF cells (c) treated with IFN-β (20 ng/ml) for the indicated time. d,e, Immunofluorescent staining of cGAS (green) and G3BP1 (red) in ISD- or poly(I:C)-treated U937 cells (d) or HeLa cells (e), respectively. Hoechst (blue), nuclear stain. f,g, cGAS expression (g) and the percentage of cells with stress granule formation (f) in Fig. 2e were analyzed. At least 90 cells from each group were analyzed (f) (n = 2 independent experiments). h, The interaction between cGAS and Beclin-1 was analyzed by immunoprecipitation (IP) with anti-Flag M2 beads in Flag-cGAS-expressing WT or G3BP1–/– HEK293T cells. WCL, whole-cell lysate. i, Representative images of GFP-LC3-expressing HeLa cells transfected with non-targeting control siRNA (si-NC) or G3BP1-specific siRNA (si-G3BP1), followed by starvation (EBSS-treated) for 4 h (left). The GFP-LC3 puncta per cell were quantified (right); at least 100 cells from each group were analyzed (n = 2 independent experiments). Scale bars, 10 μm. β-actin (a), GAPDH (i) and α-tubulin (a,g,h), loading controls. The α-tubulin blot in the IP sample indicates the purity of IP (h). **P < 0.01, ***P < 0.001, two-tailed t test (b,c). Data are representative of three experiments (a–e,h). Data are mean ± s.e.m. of triplicate samples in b,c.
Supplementary Figure 5 G3BP1 binds to cGAS and promotes cGAS–DNA binding.
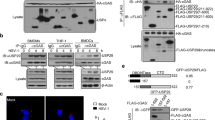
a, The cGAS–G3BP1 interaction was analyzed by immunoprecipitation with anti-cGAS antibody in HT-DNA-treated U937 cells. b, Recombinant cGAS proteins, in the presence or absence of ISD, were incubated with recombinant Flag-tagged G3BP1 proteins for 2 h. The cGAS–G3BP1 interaction was analyzed. c,d, The interaction between recombinant cGAS and the indicated G3BP1 variant proteins was analyzed by immunoprecipitation. e, U937 cells were treated with HT-DNA for the indicated time. Endogenous cGAS was immunoprecipitated and the interaction of cGAS and IFI16 was analyzed. f,g, EMSA analysis of cGAS–DNA binding in the presence or absence of Flag-tagged G3BP1FL or the mutants as indicated. (ISD)2 (1 μM) was used as DNA probe. h, The DNA-binding capacity of G3BP1 alone was analyzed by EMSA. Grayscale-inverted images are shown (f–h). i, Plasmids encoding HA-tagged G3BP1 or HA-tagged cGAS were respectively expressed in HEK293T cells, followed by transfection of biotin-ISD. The DNA-binding capacity of the proteins was analyzed by pulldown with streptavidin beads. j, The DNA-binding capacity of the proteins was analyzed by pulldown with streptavidin beads following incubating biotin-ISD with recombinant cGAS or G3BP1 proteins. WCL, whole-cell lysates. IP, immunoprecipitation. α-Tubulin (a) and GAPDH (e,i), loading controls. The α-tubulin blot (a) or anti-GAPDH blot (e) in the IP samples indicates the purity of IP. Data are representative of three experiments.
Supplementary Figure 6 G3BP1 promotes the DNA binding and activation of cGAS.
a, EMSA analysis of cGAS-DNA binding, in the presence or absence of G3BP1FL, in a high-pH (10.5) electrophoresis buffer. (ISD)2 (1 μM) was used as DNA probe. b, Image J analysis of the intensity of the free DNA bands in a; the Kd was calculated with the binding equilibrium equation. Kd, dissociation constant. c,d, EMSA analysis of cGAS–DNA binding, in the presence of G3BP1FL or the mutants as indicated, in a high-pH (10.5) electrophoresis buffer. e, Recombinant His-tagged cGAS protein, with or without recombinant Flag-tagged full length or truncated mutants of G3BP1 proteins, was respectively incubated with ISD. The production of cGAMP was analyzed by LC–MS/MRM. f, The in vitro cGAMP synthesis assay as in e was performed. The different amounts of HT-DNA as indicated were used in each group. g, Recombinant full length cGAS (FL) or the C terminus of cGAS (cGAS-C), in the presence or absence of recombinant G3BP1, was respectively incubated with HT-DNA. The production of cGAMP was analyzed by LC–MS/MRM. N.D., non-detected. ***P < 0.001, two-tailed t test (e–g). Grayscale-inverted images are shown (a,c,d). Data are representative of three experiments, mean ± s.e.m. of triplicate samples in e–g.
Supplementary Figure 7 EGCG blocks DNA-induced cGAS/STING signaling.
a,c, qPCR analysis of IFNB mRNA expression in HT-DNA-treated U937 cells (a) or human primary macrophages (c) with a 1-h pretreatment of EGCG at the indicated concentrations. b, qPCR analysis of IFNB mRNA expression in U937 cells transfected with the indicated concentrations of ISD following EGCG (20 μM) pretreatment for 1 h. d,e, qPCR analysis of IFNB mRNA expression in BMDMs (d) and human primary macrophages (e) transfected with HT-DNA following a 1-h pretreatment with EGCG at the indicated concentrations. The IC50 of EGCG for inhibiting DNA-induced IFNB production was calculated. f, Human primary macrophages were pretreated with the indicated concentrations of EGCG for 1 h, followed by HT-DNA transfection for the indicated time. Cell lysates were analyzed by immunoblotting with the indicated antibodies. g, Immunoblot analysis of cell lysates treated similarly as in a with the indicated antibodies. h, Human primary macrophages were pretreated with EGCG followed by stimulation with Pam3CSK4 (1 μg/ml) (left), poly(I:C) (50 μg/ml) (middle) or LPS (1 μg/ml) (right) for 2 h. TNF mRNA expression was analyzed by qPCR. i, HeLa cells expressing Flag-cGAS were treated with EGCG (20 μM) for 1 h, followed by immunoprecipitation (IP) with anti-Flag M2 beads. The interaction between cGAS and Beclin-1 was analyzed. WCL, whole-cell lysate. j, Becn1 was knocked down in L929 cells with siRNAs and the cells were further transfected with HT-DNA (2 μg/ml) following a 1-h pretreatment with EGCG. The mRNA expression of Ifnb was measured by qPCR (top). Knockdown of Becn1 was analyzed by immunoblotting (bottom). k, WT C57BL/6 mice were administrated EGCG (i.p., 40 mg/kg) and the blood concentration of EGCG was measured by LC–MS/MRM at the indicated time postinjection. Results are mean ± s.e.m. of n = 5 mice. l, HEK293T cells were transfected with plasmids as indicated for 6 h, followed by treatment with EGCG (20 μM) or Bx-795 (TBK1 inhibitor) for another 18 h. Cell lysates were analyzed by immunoblotting with the indicated antibodies. GAPDH (g,i), β-actin (f,l) and α-tubulin (j), loading controls. The GAPDH blot in the IP samples indicates the purity of IP (i). *P < 0.05, **P < 0.01, ***P < 0.001, two-tailed t test (a–c,j). Data are representative of three experiments (a–j,l). Data are mean ± s.e.m. of triplicate samples in a–e,h,j.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Dataset 1
Supplementary Table 1
Identification of cGAS-interacting proteins by mass spectrometry
Supplementary Table 2
High-ranking candidates of cGAS-interacting proteins
Supplementary Table 3
Primers used in the study
Supplementary Table 4
Antibodies used in the study
Rights and permissions
About this article
Cite this article
Liu, ZS., Cai, H., Xue, W. et al. G3BP1 promotes DNA binding and activation of cGAS. Nat Immunol 20, 18–28 (2019). https://doi.org/10.1038/s41590-018-0262-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-018-0262-4
This article is cited by
-
TXNRD1 drives the innate immune response in senescent cells with implications for age-associated inflammation
Nature Aging (2024)
-
SAM68 directs STING signaling to apoptosis in macrophages
Communications Biology (2024)
-
Mechanism and therapeutic potential of targeting cGAS-STING signaling in neurological disorders
Molecular Neurodegeneration (2023)
-
Meloxicam inhibits STING phosphorylation and alleviates intracellular DNA-mediated autoimmune responses
Cell & Bioscience (2023)
-
Phase-separated nucleocapsid protein of SARS-CoV-2 suppresses cGAS-DNA recognition by disrupting cGAS-G3BP1 complex
Signal Transduction and Targeted Therapy (2023)