Abstract
Foxp3+ regulatory T cells (Treg cells) are the central component of peripheral immune tolerance. Whereas a dysregulated Treg cytokine signature has been observed in autoimmune diseases, the regulatory mechanisms underlying pro- and anti-inflammatory cytokine production are elusive. Here, we identify an imbalance between the cytokines IFN-γ and IL-10 as a shared Treg signature present in patients with multiple sclerosis and under high-salt conditions. RNA-sequencing analysis on human Treg subpopulations revealed β-catenin as a key regulator of IFN-γ and IL-10 expression. The activated β-catenin signature was enriched in human IFN-γ+ Treg cells, as confirmed in vivo with Treg-specific β-catenin-stabilized mice exhibiting lethal autoimmunity with a dysfunctional Treg phenotype. Moreover, we identified prostaglandin E receptor 2 (PTGER2) as a regulator of IFN-γ and IL-10 production under a high-salt environment, with skewed activation of the β-catenin–SGK1–Foxo axis. Our findings reveal a novel PTGER2–β-catenin loop in Treg cells linking environmental high-salt conditions to autoimmunity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
RNA-seq data are available in the GEO repository under accession code GSE116283. The remaining data that support the findings of this study are available from the corresponding authors upon reasonable request.
Change history
07 June 2019
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Sakaguchi, S. et al. Foxp3+ CD25+ CD4+ natural regulatory T cells in dominant self-tolerance and autoimmune disease. Immunol. Rev. 212, 8–27 (2006).
Viglietta, V., Baecher-Allan, C., Weiner, H. L. & Hafler, D. A. Loss of functional suppression by CD4+CD25+ regulatory T cells in patients with multiple sclerosis. J. Exp. Med. 199, 971–979 (2004).
Buckner, J. H. Mechanisms of impaired regulation by CD4+CD25+FOXP3+ regulatory T cells in human autoimmune diseases. Nat. Rev. Immunol. 10, 849–859 (2010).
Miyara, M. et al. Functional delineation and differentiation dynamics of human CD4+ T cells expressing the FoxP3 transcription factor. Immunity 30, 899–911 (2009).
Dominguez-Villar, M., Baecher-Allan, C. M. & Hafler, D. A. Identification of T helper type 1-like, Foxp3+ regulatory T cells in human autoimmune disease. Nat. Med. 17, 673–675 (2011).
McClymont, S. A. et al. Plasticity of human regulatory T cells in healthy subjects and patients with type 1 diabetes. J. Immunol. 186, 3918–3926 (2011).
Lowther, D. E. et al. PD-1 marks dysfunctional regulatory T cells in malignant gliomas. JCI Insight 1, e85935 (2016).
Rubtsov, Y. P. et al. Regulatory T cell-derived interleukin-10 limits inflammation at environmental interfaces. Immunity 28, 546–558 (2008).
Laidlaw, B. J. et al. Production of IL-10 by CD4+ regulatory T cells during the resolution of infection promotes the maturation of memory CD8+ T cells. Nat. Immunol. 16, 871–879 (2015).
Marson, A., Housley, W. J. & Hafler, D. A. Genetic basis of autoimmunity. J. Clin. Invest. 125, 2234–2241 (2015).
Hafler, D. A. et al. Risk alleles for multiple sclerosis identified by a genomewide study. N. Engl. J. Med. 357, 851–862 (2007).
International Multiple SclerosisGenetics Consortium et al. The Multiple Sclerosis Genomic Map: role of peripheral immune cells and resident microglia in susceptibility. Preprint at https://www.biorxiv.org/content/early/2017/07/13/143933 (2017).
Olsson, T., Barcellos, L. F. & Alfredsson, L. Interactions between genetic, lifestyle and environmental risk factors for multiple sclerosis. Nat. Rev. Neurol. 13, 25–36 (2017).
Kleinewietfeld, M. et al. Sodium chloride drives autoimmune disease by the induction of pathogenic TH17 cells. Nature 496, 518–522 (2013).
Wu, C. et al. Induction of pathogenic TH17 cells by inducible salt-sensing kinase SGK1. Nature 496, 513–517 (2013).
Hernandez, A. L. et al. Sodium chloride inhibits the suppressive function of FOXP3+ regulatory T cells. J. Clin. Invest. 125, 4212–4222 (2015).
Wei, Y. et al. High salt diet stimulates gut Th17 response and exacerbates TNBS-induced colitis in mice. Oncotarget 8, 70–82 (2017).
Tubbs, A. L., Liu, B., Rogers, T. D., Sartor, R. B. & Miao, E. A. Dietary salt exacerbates experimental colitis. J. Immunol. 199, 1051–1059 (2017).
Paling, D. et al. Sodium accumulation is associated with disability and a progressive course in multiple sclerosis. Brain 136, 2305–2317 (2013).
Fitzgerald, K. C. et al. Sodium intake and multiple sclerosis activity and progression in BENEFIT. Ann. Neurol. 82, 20–29 (2017).
Clevers, H. Wnt/beta-catenin signaling in development and disease. Cell 127, 469–480 (2006).
Naito, A. T. et al. Complement C1q activates canonical Wnt signaling and promotes aging-related phenotypes. Cell 149, 1298–1313 (2012).
Staal, F. J., van Noort, M., Strous, G. J. & Clevers, H. C. Wnt signals are transmitted through N-terminally dephosphorylated beta-catenin. EMBO Rep. 3, 63–68 (2002).
Duhen, T., Duhen, R., Lanzavecchia, A., Sallusto, F. & Campbell, D. J. Functionally distinct subsets of human FOXP3+ Treg cells that phenotypically mirror effector Th cells. Blood 119, 4430–4440 (2012).
Kitz, A. et al. AKT isoforms modulate Th1-like Treg generation and function in human autoimmune disease. EMBO Rep. 17, 1169–1183 (2016).
Wing, K. et al. CTLA-4 control over Foxp3+ regulatory T cell function. Science 322, 271–275 (2008).
Harada, N. et al. Intestinal polyposis in mice with a dominant stable mutation of the beta-catenin gene. EMBO J. 18, 5931–5942 (1999).
Keerthivasan, S. et al. β-Catenin promotes colitis and colon cancer through imprinting of proinflammatory properties in T cells. Sci. Transl. Med. 6, 225ra28 (2014).
Sebastian, M. et al. Helios controls a limited subset of regulatory T cell functions. J. Immunol. 196, 144–155 (2016).
Essers, M. A. et al. Functional interaction between beta-catenin and FOXO in oxidative stress signaling. Science 308, 1181–1184 (2005).
Okada, K. et al. Wnt/β-catenin signaling contributes to skeletal myopathy in heart failure via direct interaction with Forkhead box O. Circ Heart Fail 8, 799–808 (2015).
Ouyang, W. et al. Novel Foxo1-dependent transcriptional programs control Treg cell function. Nature 491, 554–559 (2012).
Dehner, M., Hadjihannas, M., Weiske, J., Huber, O. & Behrens, J. Wnt signaling inhibits Forkhead box O3a-induced transcription and apoptosis through up-regulation of serum- and glucocorticoid-inducible kinase 1. J. Biol. Chem. 283, 19201–19210 (2008).
Wang, R. et al. Global discovery of high-NaCl-induced changes of protein phosphorylation. Am. J. Physiol. Cell. Physiol. 307, C442–C454 (2014).
Irarrazabal, C. E., Burg, M. B., Ward, S. G. & Ferraris, J. D. Phosphatidylinositol 3-kinase mediates activation of ATM by high NaCl and by ionizing radiation: role in osmoprotective transcriptional regulation. Proc. Natl. Acad. Sci. USA 103, 8882–8887 (2006).
Fang, D. et al. Phosphorylation of beta-catenin by AKT promotes beta-catenin transcriptional activity. J. Biol. Chem. 282, 11221–11229 (2007).
Lund, R. J. et al. Genome-wide identification of novel genes involved in early Th1 and Th2 cell differentiation. J. Immunol. 178, 3648–3660 (2007).
Boniface, K. et al. Prostaglandin E2 regulates Th17 cell differentiation and function through cyclic AMP and EP2/EP4 receptor signaling. J. Exp. Med. 206, 535–548 (2009).
Kofler, D. M. et al. Decreased RORC-dependent silencing of prostaglandin receptor EP2 induces autoimmune Th17 cells. J. Clin. Invest. 124, 2513–2522 (2014).
Li, X. et al. Divergent requirement for Gαs and cAMP in the differentiation and inflammatory profile of distinct mouse Th subsets. J. Clin. Invest. 122, 963–973 (2012).
Sreeramkumar, V., Fresno, M. & Cuesta, N. Prostaglandin E2 and T cells: friends or foes? Immunol. Cell Biol. 90, 579–586 (2012).
Yao, C. et al. Prostaglandin E2 promotes Th1 differentiation via synergistic amplification of IL-12 signalling by cAMP and PI3-kinase. Nat. Commun. 4, 1685 (2013).
Shin, H. et al. Identification of transcriptional targets of Wnt/beta-catenin signaling in dermal papilla cells of human scalp hair follicles: EP2 is a novel transcriptional target of Wnt3a. J. Dermatol. Sci. 58, 91–96 (2010).
Castellone, M. D., Teramoto, H., Williams, B. O., Druey, K. M. & Gutkind, J. S. Prostaglandin E2 promotes colon cancer cell growth through a Gs-axin-beta-catenin signaling axis. Science 310, 1504–1510 (2005).
Ding, Y., Shen, S., Lino, A. C., Curotto de Lafaille, M. A. & Lafaille, J. J. Beta-catenin stabilization extends regulatory T cell survival and induces anergy in nonregulatory T cells. Nat. Med. 14, 162–169 (2008).
van Loosdregt, J. et al. Canonical Wnt signaling negatively modulates regulatory T cell function. Immunity 39, 298–310 (2013).
Pappalardo, J. L. & Hafler, D. A. The Human Functional Genomics Project: understanding generation of diversity. Cell 167, 894–896 (2016).
Wilck, N. et al. Salt-responsive gut commensal modulates TH17 axis and disease. Nature 551, 585–589 (2017).
Mahic, M., Yaqub, S., Johansson, C. C., Taskén, K. & Aandahl, E. M. FOXP3+CD4+CD25+ adaptive regulatory T cells express cyclooxygenase-2 and suppress effector T cells by a prostaglandin E2-dependent mechanism. J. Immunol. 177, 246–254 (2006).
Kihara, Y. et al. Targeted lipidomics reveals mPGES-1-PGE2 as a therapeutic target for multiple sclerosis. Proc. Natl. Acad. Sci. USA 106, 21807–21812 (2009).
Acknowledgements
We thank L. Devine, C. Wang, and H. Tomita for cell sorting; E. Cheng and M. Zhang for preparation of the RNA-seq library; K. Tanaka for microscopic imaging; and M. Shimizu, H. Taniwaki, and N. Yamanaka for technical support. FIC mice were a generous gift from S. Sakaguchi (Osaka University). This work was supported by grants from the Uehara Memorial Foundation Research Fellowship, MSD Life Science Foundation Research Fellowship, LEGEND Study Abroad Grant from BioLegend, and Tomy Digital Biology (to T.S.); the Ministry of Education, Culture, Sports, Science and Technology (MEXT), JSPS KAKENHI (grant 21229010), and the Core Research for Evolutional Science and Technology (CREST) program from the Japan Science and Technology Agency (to I.K.); and the National Institutes of Health (grants P01 AI073748, P01 AI045757, U19 AI046130, U19 AI070352, and P01 AI039671) and the Nancy Taylor Foundation for Chronic Diseases (to D.A.H.). RNA-seq was conducted at the Yale Stem Cell Center Genomics Core facility, which is supported by the Connecticut Regenerative Medicine Research Fund and the Li Ka Shing Foundation.
Author information
Authors and Affiliations
Contributions
T.S., D.M.R., and C.M.U. performed in vitro experiments with the help of M.R.L. and M.D.V.; T.S. and A.T.N. performed in vivo experiments with the help of H.A. and T.N.; T.S. and M.R.L. analyzed the RNA-seq data under the supervision of D.A.H.; T.S. performed data analysis and wrote the manuscript under the supervision of A.T.N., I.K., M.R.L., M.D.-V., and D.A.H.; A.T.N., I.K., M.D.-V., and D.A.H. provided supervision in the overall study.
Corresponding author
Ethics declarations
Competing interests
D.A.H. has consulted for the following companies: Bayer Pharmaceuticals, Biohaven Pharmaceuticals, Bristol Myers Squibb, Compass Therapeutics, Eisai Pharmaceuticals, EMD Serono, Genentech, Juno Therapeutics, McKinsey & Co., MedImmune/AstraZeneca, Mylan, Pharmaceuticals, Neurophage Pharmaceuticals, NKT Therapeutics, Novartis Pharmaceuticals, Proclara Biosciences, Questcor Pharmaceuticals, Roche, Sage Therapeutics, Sanofi Genzyme, Toray Industries, and Versant Venture.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Memory Treg cells are the main source of effector cytokines IFN-γ and IL-10.
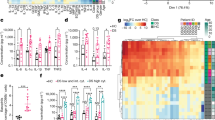
(a) Sorting strategy for memory and naive Treg cells from circulating human CD4+ T cells. (b) mRNA expression of IFNG and IL10 gene on memory and naive Treg cells (memory Treg cells; n = 35 subjects, naive Treg cells; n = 16 subjects). P values were calculated by two-sided Student’s t-test. Data were represented as mean + /- SD.
Supplementary Figure 2 β-catenin signaling is activated in the IFN-γ-producing human Treg subset.
(a) GSEA enrichment plots of four different Wnt signaling pathway gene sets between IFN-γSP vs. DN. (b) Flow cytometric analysis of TCF-1 expression on ex vivo Treg subpopulations relative to DN (n = 7 subjects). *P < 0.05, **P < 0.01 (one-way ANOVA with Tukey’s multiple comparisons test). (c) AXIN2 and TCF7 mRNA expression in IFN-γ+ and IFN-γ– human Treg populations assessed by DNA microarray (n = 8 subjects). P values were calculated by two-sided Student’s t-test. (d) CTNNB1 gene expression on Treg cells transduced with a non-targeted shRNA or a CTNNB1 shRNA and cultured for 5 days (n = 10 subjects). P values were calculated by two-sided Student’s t-test. (e) Frequency of IFN-γ and IL-10 positive cell number. Treg cells were stimulated with anti-CD3 and anti-CD28 in the presence of SGK1 inhibitor GSK650394 (SGK1-i), IL-12 (TH1), or IL-12 and GSK650394 (TH1 + SGK1-i) (n = 6 subjects) ***P < 0.001 (one-way ANOVA with Tukey’s multiple comparisons test). Data were represented as mean + /- SD.
Supplementary Figure 3 Treg-specific activation of β-catenin induces an IFN-γ-secreting dysfunctional phenotype with Scurfy-like autoimmunity.
(a) Schematic of the wild-type and targeted CTNNB1 allele. (b) Flow cytometric analysis of β-catenin and Foxp3 in peripheral lymph nodes, spleen, and thymus CD4+ T cells from Foxp3Cre and Ctnnb1ΔEx3; Foxp3Cre mice. (c) Expression for classical helper cytokines and transcription factors in both Treg cells and T effector cells (CD4+ CD25neg; non-Treg) assessed by qPCR (n = 4 mice). *P < 0.05, **P < 0.01 (two-sided Student’s t-test). (d) Flow cytometric analysis of Foxp3 and Helios expression on CD4+ T cells in peripheral lymph nodes and spleen from Foxp3Cre and Ctnnb1ΔEx3; Foxp3Cre mice at the age of 3 weeks. Percentages of Foxp3+ and/or Helios+ CD4+ T cells isolated from lymph nodes are shown at the bottom. (n = 5-6 mice) ***P < 0.001 (two-way ANOVA with Sidak’s multiple comparisons test). (e) GSEA enrichment plot between Foxp3Cre and Ctnnb1ΔEx3; Foxp3Cre Treg cells using the gene set that is positively regulated by Foxo1 (left) and negatively regulated by Foxo1 (right) identified from the comparison between wild-type vs. Foxo1 KO Treg cells (GSE40655). Normalized enrichment score (NES) and false discovery rate (FDR) are indicated. Data were represented as mean + /- SD.
Supplementary Figure 4 High salt activates the β-catenin–SGK1–Foxo axis in IFN-γ-producing human Treg cells.
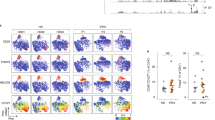
(a) Flow cytometric analysis of p-SGK1 (Thr256) (left) and p-Foxo1 (Ser256) (right) level in human Treg subsets. Treg cells were stimulated with anti-CD3 and anti-CD28 for 96 h followed by 4 h PMA plus iomomycin stimulation, and the expression of p-SGK1 and p-Foxo1 were determined by intracellular staining in each subset. (p-SGK1; n = 12, p-Foxo1; n = 10 subjects) (b) Representative immunofluorescence images of human Treg cells with PLA signal for β-catenin-Foxo1 interaction (red) and IFN-γ staining (green). Nuclei were stained with DAPI (blue). PLA signal (arrowheads) was observed in an IFN-γ positive cell (arrow). Scale bar 5 μm. Data were representative of three experiments. (c) Flow cytometric analysis of active β-catenin, p-SGK1, and p-Foxo1 expression in human IL-10 producing Treg cells. Treg cells were stimulated with anti-CD3 and anti-CD28 in the presence (NaCl) or absence (Control) of additional 40 mM NaCl for 96 h followed by PMA plus iomomycin stimulation for 4 h (active β-catenin; n = 15, p-SGK1; n = 13, p-Foxo1; n = 10 subjects). (d) Flow cytometric analysis of TCF-1 expression in human IFN-γ producing Treg cells. Treg cells were cultured as in (c) (n = 3 subjects). P values were calculated by two-sided Student’s t-test for (c, d). (e) Relative frequency of IFN-γ and IL-10 positive cell number (fold of control condition) in human Treg cells cultured as in Fig. 6c (n = 11 subjects). *P < 0.05, **P < 0.01 (one-way ANOVA with Tukey’s multiple comparisons test). (f) Relative frequency of IFN-γ and IL-10 positive cell number (fold of control condition) in human Treg cells cultured as in Fig. 6d (n = 9 subjects). *P < 0.05, **P < 0.01 (one-way ANOVA with Tukey’s multiple comparisons test). (g) Flow cytometric analysis of p-SGK1 expression in human IFN-γ producing Treg cells. Treg cells were cultured as in Fig. 6d (n = 6 subjects). *P < 0.05, **P < 0.01 (one-way ANOVA with Tukey’s multiple comparisons test). (h) Relative frequency of IFN-γ and IL-10 positive cell number in human Treg cells stimulated in the presence (NaCl) or absence (Control) of additional 40 mM NaCl with and without Wnt/inhibitor GSK650394 (SGK1-i) for 96 h (n = 11 subjects). *P < 0.05, **P < 0.01 (one-way ANOVA with Tukey’s multiple comparisons test). (i) Flow cytometric analysis of p-Foxo1 expression in human IFN-γ producing Treg cells. Treg cells were cultured as in (h) (n = 8 subjects). *P < 0.05 (one-way ANOVA with Tukey’s multiple comparisons test). Data were represented as mean + /- SD.
Supplementary Figure 5 The β-catenin–SGK1–Foxo axis is also activated in Teff and human Jurkat T cells under high-salt conditions.
(a) Flow cytometric analysis of active β-catenin expression in human T effector cells (Teffs) and human Jurkat T cells. Human Teff were stimulated as well as Treg cells for 96 h (n = 8 subjects). Human Jurkat T cells were cultured without TCR stimulation for 120 h (n = 12). Both were cultured in the presence (NaCl) or absence (Control) of additional 40 mM NaCl. P values were calculated by two-sided Student’s t-test. (b) Relative frequency of IFN-γ and IL-10 positive cell number (fold of control condition) in human Teffs cultured as in (a) (n = 8 subjects). P values were calculated by two-sided Student’s t-test. (c, d) Flow cytometric analysis of p-SGK1 (Thr256) (c) and p-Foxo1 (Ser256) (d) in human Jurkat T cells. Human Jurkat T cells were transduced with scramble gRNA (CRISPR/Scramble) or CTNNB1 targeted gRNA (CRISPR/CTNNB1) with Cas9. Both cell lines were cultured in the presence (NaCl) or absence (Control) of additional 40 mM NaCl without TCR stimulation for 120 h (n = 4). **P < 0.01 (one-way ANOVA with Tukey’s multiple comparisons test). Data were represented as mean + /- SD.
Supplementary Figure 6 High-salt-induced β-catenin activation via AKT is independent of Wnt ligands.
(a) Relative frequency of IFN-γ and IL-10 positive cell number (fold of control condition) in human Treg cells. Treg cells were stimulated with anti-CD3 and anti-CD28 in the presence of Fzd8-FC (Fzd), additional 40 mM NaCl (NaCl), or Fzd8-FC and NaCl (NaCl + Fzd) (n = 7 subjects). *P < 0.05, **P < 0.01 (one-way ANOVA with Tukey’s multiple comparisons test). (b) Relative expression level of active β-catenin in human Treg cells cultured as in (a). (n = 7 subjects) *P < 0.05, **P < 0.01 (one-way ANOVA with Tukey’s multiple comparisons test). (c) GSEA enrichment plots of PI3K/AKT pathway gene sets between IFN-γSP vs. IL10SP. Normalized enrichment score (NES) and false discovery rate (FDR) are indicated at the bottom of each plot. (d) Relative expression level of p-AKT on Treg subsets. Treg cells were stimulated with anti-CD3 and anti-CD28 for 4 days, followed by 4 h PMA/iomomycin stimulation and intracellular cytokine staining for IFN-γ and IL-10 (n = 5 subjects). *P < 0.05, **P < 0.01 (one-way ANOVA with Tukey’s multiple comparisons test). (e) Flow cytometric analysis of β-catenin phosphorylation at s522 in human Jurkat T cells. Human Jurkat T cells were stimulated in the presence of AKT inhibitor MK2206 (AKT-i), additional 40 mM NaCl (NaCl), or MK2206 and NaCl (NaCl + AKT-i) (n = 4). (f) Flow cytometric analysis of GSK3β phosphorylation at s9 and AKT phosphorylation at s473 in human Jurkat T cells. Human Jurkat T cells were prepared as in Supplementary Fig. 6c. (n = 4). *P < 0.05, **P < 0.01 (one-way ANOVA with Tukey’s multiple comparisons test). Data were represented as mean + /- SD.
Supplementary Figure 7 The PTGER2–β-catenin loop is activated by high-salt stimulation.
(a) PTGER2 expression assessed by RNA-seq on ex vivo Treg subpopulations (n = 8 subjects). (b) Flow cytometric analysis of PTGER2 in human Jurkat T cells. Human Jurkat T cells were prepared as in Supplementary Fig. 6c. (n = 4). **P < 0.01, ***P < 0.001 (one-way ANOVA with Tukey’s multiple comparisons test). (c) Flow cytometric analysis of active β-catenin in human Jurkat T cells. Human Jurkat T cells were transduced with a scramble shRNA or a PTGER2 shRNA and cultured in normal media (Control) or media supplemented with additional 40 mM NaCl (NaCl) for 120 h. (n = 4) *P < 0.05, **P < 0.01 (one-way ANOVA with Tukey’s multiple comparisons test). (d) Representative flow cytometric analysis of IFN-γ and IL-10 production in human Treg cells cultured in the normal media (Control) or media supplemented with additional 40 mM NaCl (NaCl) with anti CD3 (2 μg/ml) and different concentration of anti CD28 (1, 2, 5 μg/ml) for 96 h. Relative frequency of IFN-γ and IL-10 producing Treg cells are shown at the bottom (n = 4 subjects). *P < 0.05, **P < 0.01 (two-way ANOVA with Sidak’s multiple comparisons test). Data were represented as mean + /- SD.
Supplementary Figure 8 Schematic model of the role of PTGER2 and the AKT–β-catenin–SGK1–Foxo axis in the production of IFN-γ and IL-10 in Treg cells.
AKT-β-catenin signaling balances between IFN-γ and IL-10 production in Treg cells. Under high salt conditions, PTGER2 was increased and established the positive feed forward loop with β-catenin, resulted in amplified activation of the β-catenin-SGK1-Foxo axis in IFN-γ-producing Treg cells.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8
Supplementary Table 1: Upstream regulator analysis in IPA between each Treg subpopulation
Lists of the top-ranking genes identified by IPA analysis as upstream regulators between each Treg subpopulations. Tables show statistically significant (overlap P value <0.05) upstream regulators in each comparison (genes that could not be calculated for fold change were blank). CTNNB1 gene, which encodes β-catenin protein, was highlighted in red.
Rights and permissions
About this article
Cite this article
Sumida, T., Lincoln, M.R., Ukeje, C.M. et al. Activated β-catenin in Foxp3+ regulatory T cells links inflammatory environments to autoimmunity. Nat Immunol 19, 1391–1402 (2018). https://doi.org/10.1038/s41590-018-0236-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-018-0236-6
This article is cited by
-
The dynamic shifts of IL-10-producing Th17 and IL-17-producing Treg in health and disease: a crosstalk between ancient "Yin-Yang" theory and modern immunology
Cell Communication and Signaling (2024)
-
The regulation and differentiation of regulatory T cells and their dysfunction in autoimmune diseases
Nature Reviews Immunology (2024)
-
Dysregulation of Wnt/β-catenin signaling contributes to intestinal inflammation through regulation of group 3 innate lymphoid cells
Nature Communications (2024)
-
Salt sensitivity includes effects on immune cell signalling and metabolism
Nature Reviews Immunology (2023)
-
Regulatory T cells in the face of the intestinal microbiota
Nature Reviews Immunology (2023)