Abstract
Foxo transcription factors play an essential role in regulating specialized lymphocyte functions and in maintaining T cell quiescence. Here, we used a system in which Foxo1 transcription-factor activity, which is normally terminated upon cell activation, cannot be silenced, and we show that enforcing Foxo1 activity disrupts homeostasis of CD4 conventional and regulatory T cells. Despite limiting cell metabolism, continued Foxo1 activity is associated with increased activation of the kinase Akt and a cell-intrinsic proliferative advantage; however, survival and cell division are decreased in a competitive setting or growth-factor-limiting conditions. Via control of expression of the transcription factor Myc and the IL-2 receptor β-chain, termination of Foxo1 signaling couples the increase in cellular cholesterol to biomass accumulation after activation, thereby facilitating immunological synapse formation and mTORC1 activity. These data reveal that Foxo1 regulates the integration of metabolic and mitogenic signals essential for T cell competitive fitness and the coordination of cell growth with cell division.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Manning, B. D. & Toker, A. AKT/PKB signaling: navigating the network. Cell 169, 381–405 (2017).
Liao, W., Lin, J. X. & Leonard, W. J. Interleukin-2 at the crossroads of effector responses, tolerance, and immunotherapy. Immunity 38, 13–25 (2013).
Smith-Garvin, J. E., Koretzky, G. A. & Jordan, M. S. T cell activation. Annu. Rev. Immunol. 27, 591–619 (2009).
Kerdiles, Y. M. et al. Foxo1 links homing and survival of naive T cells by regulating L-selectin, CCR7 and interleukin 7 receptor. Nat. Immunol. 10, 176–184 (2009).
Ouyang, W., Beckett, O., Flavell, R. A. & Li, M. O. An essential role of the Forkhead-box transcription factor Foxo1 in control of T cell homeostasis and tolerance. Immunity 30, 358–371 (2009).
Kerdiles, Y. M. et al. Foxo transcription factors control regulatory T cell development and function. Immunity 33, 890–904 (2010).
Ouyang, W. et al. Foxo proteins cooperatively control the differentiation of Foxp3+ regulatory T cells. Nat. Immunol. 11, 618–627 (2010).
Ouyang, W. et al. Novel Foxo1-dependent transcriptional programs control Treg cell function. Nature 491, 554–559 (2012).
Zeng, H. et al. mTORC1 couples immune signals and metabolic programming to establish Treg-cell function. Nature 499, 485–490 (2013).
Yang, K. et al. T cell exit from quiescence and differentiation into Th2 cells depend on Raptor-mTORC1-mediated metabolic reprogramming. Immunity 39, 1043–1056 (2013).
Kuczma, M. et al. Foxp3-deficient regulatory T cells do not revert into conventional effector CD4+ T cells but constitute a unique cell subset. J. Immunol. 183, 3731–3741 (2009).
Kousteni, S. FoxO1, the transcriptional chief of staff of energy metabolism. Bone 50, 437–443 (2012).
Webb, A. E., Kundaje, A. & Brunet, A. Characterization of the direct targets of FOXO transcription factors throughout evolution. Aging Cell 15, 673–685 (2016).
Wilhelm, K. et al. FOXO1 couples metabolic activity and growth state in the vascular endothelium. Nature 529, 216–220 (2016).
Sinclair, L. V. et al. Control of amino-acid transport by antigen receptors coordinates the metabolic reprogramming essential for T cell differentiation. Nat. Immunol. 14, 500–508 (2013).
Wang, R. et al. The transcription factor Myc controls metabolic reprogramming upon T lymphocyte activation. Immunity 35, 871–882 (2011).
Lord, J. D., McIntosh, B. C., Greenberg, P. D. & Nelson, B. H. The IL-2 receptor promotes proliferation, bcl-2 and bcl-x induction, but not cell viability through the adapter molecule Shc. J. Immunol. 161, 4627–4633 (1998).
Preston, G. C. et al. Single cell tuning of Myc expression by antigen receptor signal strength and interleukin-2 in T lymphocytes. EMBO J. 34, 2008–2024 (2015).
Buck, M. D., O’Sullivan, D. & Pearce, E. L. T cell metabolism drives immunity. J. Exp. Med. 212, 1345–1360 (2015).
MacIver, N. J., Michalek, R. D. & Rathmell, J. C. Metabolic regulation of T lymphocytes. Annu. Rev. Immunol. 31, 259–283 (2013).
Dustin, M. L. The immunological synapse. Cancer Immunol. Res. 2, 1023–1033 (2014).
Yang, W. et al. Potentiating the antitumour response of CD8+ T cells by modulating cholesterol metabolism. Nature 531, 651–655 (2016).
Choudhuri, K. et al. Polarized release of T-cell-receptor-enriched microvesicles at the immunological synapse. Nature 507, 118–123 (2014).
Brunet, A. et al. Protein kinase SGK mediates survival signals by phosphorylating the forkhead transcription factor FKHRL1 (FOXO3a). Mol. Cell. Biol. 21, 952–965 (2001).
Feng, J., Park, J., Cron, P., Hess, D. & Hemmings, B. A. Identification of a PKB/Akt hydrophobic motif Ser-473 kinase as DNA-dependent protein kinase. J. Biol. Chem. 279, 41189–41196 (2004).
Xu, C. et al. Regulation of T cell receptor activation by dynamic membrane binding of the CD3epsilon cytoplasmic tyrosine-based motif. Cell 135, 702–713 (2008).
Hui, E. et al. T cell costimulatory receptor CD28 is a primary target for PD-1-mediated inhibition. Science 355, 1428–1433 (2017).
Sancak, Y. et al. Ragulator-Rag complex targets mTORC1 to the lysosomal surface and is necessary for its activation by amino acids. Cell 141, 290–303 (2010).
Castellano, B. M. et al. Lysosomal cholesterol activates mTORC1 via an SLC38A9-Niemann-Pick C1 signaling complex. Science 355, 1306–1311 (2017).
Marschner, K., Kollmann, K., Schweizer, M., Braulke, T. & Pohl, S. A key enzyme in the biogenesis of lysosomes is a protease that regulates cholesterol metabolism. Science 333, 87–90 (2011).
Hecht, V. C. et al. Biophysical changes reduce energetic demand in growth factor-deprived lymphocytes. J. Cell Biol. 212, 439–447 (2016).
Sancak, Y. et al. The Rag GTPases bind raptor and mediate amino acid signaling to mTORC1. Science 320, 1496–1501 (2008).
Efeyan, A. et al. Regulation of mTORC1 by the Rag GTPases is necessary for neonatal autophagy and survival. Nature 493, 679–683 (2013).
Verbist, K. C. et al. Metabolic maintenance of cell asymmetry following division in activated T lymphocytes. Nature 532, 389–393 (2016).
Laplante, M. & Sabatini, D. M. mTOR signaling in growth control and disease. Cell 149, 274–293 (2012).
Brennan, P. et al. Phosphatidylinositol 3-kinase couples the interleukin-2 receptor to the cell cycle regulator E2F. Immunity 7, 679–689 (1997).
Chen, C. C. et al. FoxOs inhibit mTORC1 and activate Akt by inducing the expression of Sestrin3 and Rictor. Dev. Cell 18, 592–604 (2010).
Delgoffe, G. M. et al. The kinase mTOR regulates the differentiation of helper T cells through the selective activation of signaling by mTORC1 and mTORC2. Nat. Immunol. 12, 295–303 (2011).
Zoncu, R., Efeyan, A. & Sabatini, D. M. mTOR: from growth signal integration to cancer, diabetes and ageing. Nat. Rev. Mol. Cell Biol. 12, 21–35 (2011).
Luo, C. T., Liao, W., Dadi, S., Toure, A. & Li, M. O. Graded Foxo1 activity in Treg cells differentiates tumour immunity from spontaneous autoimmunity. Nature 529, 532–536 (2016).
Aghajani, K., Keerthivasan, S., Yu, Y. & Gounari, F. Generation of CD4CreER(T2) transgenic mice to study development of peripheral CD4-T-cells. Genesis 50, 908–913 (2012).
Yang, K., Neale, G., Green, D. R., He, W. & Chi, H. The tumor suppressor Tsc1 enforces quiescence of naive T cells to promote immune homeostasis and function. Nat. Immunol. 12, 888–897 (2011).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740 (2011).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Huang, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Acknowledgements
We thank M. Li (Memorial Sloan Kettering Cancer Center) for Foxo1AAA mice; F. Gounari (University of Chicago) for CD4Cre–ERT2 mice; M. Farrar (University of Minnesota) for a constitutively active STAT5b retroviral construct; Z. Herbert and K. Bi for assistance with transcriptome analysis; the MGH-CNY FACS core facility for cell sorting; and members of the laboratory of L.A.T. for discussions. This work was supported by the US National Institutes of Health (P01HL018646 to L.A.T.; R21AI126143 to L.A.T. and R.H.N.; R01AI124693 to D.J.C.; R01HL011879 and P01AI056299 to B.R.B.; AI101407 and CA176624 to H.C.), the Wellcome Trust (PRF 100262 to M.L.D.) and the European Research Council (ERC-2014-AdG_670930 to M.L.D. and E.B.C.).
Author information
Authors and Affiliations
Contributions
R.H.N. and L.A.T. designed the study, interpreted data and wrote the manuscript. R.H.N. performed experiments. S.S. performed experiments with Rictor flox/flox animals. J.M.S. performed experiments to help characterize autoimmunity in CD4Cre Foxo1AAA/+ animals. K.B.Y. assisted with RNA-seq data analysis. E.B.C. performed immunological synapse imaging experiments. N.R.-H., B.R.B., S.J.B., W.N.H., M.L.D., D.J.C. and H.C. assisted with data analysis and interpretation.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 T cell and Treg phenotype in CD4Cre Foxo1AAA/+ mice.
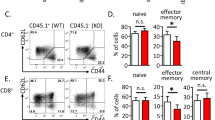
(a,b) Total numbers of lymph node cells (a) and splenocytes (b) from 4-8 week old WT (n = 12) and AAA (n = 16) mice. (c-e) Flow cytometric analysis of CD4+Foxp3+ Tregs from CD4Cre (WT; n = 6) and CD4Cre Foxo1AAA/+ (AAA; n = 9) littermates (top), and quantification of MFI of antibody staining (bottom) in 4-12 week old mice. (f) Flow cytometric analysis of CD4 + (upper) and CD8 + (lower) T cells from lymph nodes of 8 week old WT and AAA littermates. Representative of 12 mice, with similar results. (g,h) Flow cytometric analysis of Ifn-γ production in CD4+ T cells (h; n = 6) and Gzm-B production in CD8+ T cells (i; WT, n = 5; AAA, n = 3) from 4-8 week old WT and AAA littermates stimulated directly ex vivo for 4 hrs with a leucocyte activation cocktail containing Brefeldin A. For all graphs, quantification used a two-tailed Student’s t-test, with no adjustments made for multiple comparisons; centre value is the mean, with error bars indicating standard deviation. * P < 0.05; ** P < 0.005; *** P < 0.0005; **** P < 0.0001; ns = not significant.
Supplementary Figure 2 Characterization of B cell autoimmunity in CD4Cre Foxo1AAA/+ mice.
(a) Percentage of CD3+ and CD19+ cells in lymph nodes from 4-8 week old WT (n = 7) and AAA (n = 9) littermates. (b) Flow cytometric analysis of CD3-CD19+ lymph node cells from 4 week old WT and AAA littermates. The GL-7+Fas+ population (upper panels) represents germinal center B cells, and the IgM-IgD- population (lower panels) represents class-switched B cells. Representative of 4 independent experiments, with similar results. (c) Flow cytometric analysis of CD3-CD19+ B cells from 4 week old WT and AAA littermates, representative of three independent experiments, with similar results. (d) Flow cytometric analysis and quantification of CD4+CD19-Foxp3-Bcl6+CXCR5+ Tfh cells in 4-8 week old WT (n = 5) and AAA (n = 7) littermates. (e) Intracellular cytokine staining from splenocytes stimulated with PMA/ionomycin/monensin for 4 hours from 4-8 week old WT and AAA littermates (n = 4). (f-h) Serum antibody levels measured by ELISA in 6-12 week old mice (n = 12). (i,j) Representative microscopy staining of HEp-2 ANA slides stained with serum diluted 1:50 and fluorophore conjugated secondary antibody as indicated. (i): IgM (n = 2); (j): Total Ig (anti-mouse H + L; WT, n = 2; AAA, n = 3). 40× oil magnification. Serum from NZB x NZW mice was used as a positive control. (k) Representative microscopy staining of frozen kidney sections. Sections were stained for total Ig (anti-mouse IgG H + L). (l) Blood glucose levels measured by saphenous vein bleeding in 6-12 week old WT (n = 5) and AAA littermates (n = 6). (m) Percent red blood cells in blood collected from 6-12 week old WT and AAA littermates (n = 5). For all graphs, quantification used a two-tailed Student’s t-test, with no adjustments made for multiple comparisons; centre value is the mean, with error bars indicating standard deviation. * P < 0.05; ** P < 0.005; **** P < 0.0001; ns = not significant.
Supplementary Figure 3 Bone marrow chimera strategy and gene expression profiling from mixed and nonmixed bone marrow chimeras.
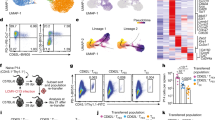
(a) Strategy for generating bone marrow chimeras. (b) Schematic of populations used for transcriptome analysis by RNAseq. Population 1: CD4+CD45.1+CD62LhiCD44lo from mixed bone marrow chimeras 10-12 weeks post-transfer; Population 2: CD4+CD45.2+CD62LhiCD44lo from mixed bone marrow chimeras 10-12 weeks post-transfer (taken from same recipients as Population 1); Population 3: CD4+CD45.2+CD62LloCD44hi from bone marrow chimeras receiving only AAA bone marrow, 10-12 weeks post-transfer. 2-3 mice for each population were used for RNAseq. (c-e) Heat maps showing DESeq2-generated normalized global expression values for transcripts isolated from all 3 populations, as described in Supplementary Figure 3; Gene sets shown are representative of genes involved in T cell activation, including cytokines and cytokine receptors (c), chemokine signaling (d), and activation induced cell death (e).
Supplementary Figure 4 Pathway enrichment analysis of CD4+ T cells from Scurfy mice versus CD4+ T cells from Foxo1AAA chimeric mice.
Differentially expressed genes in CD4+ T cells from Scurfy mice vs CD4+ T cells from wild-type mice were obtained from PubMed GEO Dataset GSE11775 analyzed using GEO2R, and were compared to differentially expressed genes from AAA CD4+ T cells from Foxo1AAA single-chimeric animals vs AAA CD4+ T cells from mixed bone marrow chimeras (populations 3 vs 2, as described in Supplementary Figure 3; n = 2). Three lists of genes were generated from this comparison: Genes that were unique to CD4 T cells in Scurfy mice versus WT mice (and not upregulated in activated versus quiescent AAA CD4 T cells), genes that were upregulated in both Scurfy and activated AAA CD4 T cells, and genes that were unique to activated AAA CD4 T cells. (a,b) Comparison of upregulated gene sets, based on log2 fold change > 1, adjusted P value < 0.05. Each gene set was analyzed using the DAVID database, and the top two enriched clusters were presented with P values (-log10) for each annotation term within the clusters, which themselves were generated by selecting the following annotation categories: GOTERM_BP_DIRECT, GOTERM_CC_DIRECT, GOTERM_MF_DIRECT and KEGG_PATHWAY (b). (c,d) Comparison of downregulated gene sets (log2 fold change < -1, adj P < 0.05), analyzed as in a and b. Arrows point to notable clusters, as mentioned in the text, within each group. For b and d, P value is a function of DAVID based on a selected gene set of differentially expressed genes, and not determined based on sample size.
Supplementary Figure 5 Dysregulation of metabolic pathways in activated Foxo1AAA CD4 T cells.
Heat maps indicating fold change in transcripts involved in metabolic pathways in “Activated” vs “Quiescent” AAA CD4 + T cells (left), and expression levels of each gene across each replicate (n = 2; scaled based on values for the entire row) within these populations. Asterisks indicate genes with log2 fold change of < -1 or > 1, and adjusted P value < 0.05. The log10 Benjamin-Hochberg correction adjusted p-values were used, corrected for the direction of fold change, to rank genes. For any adjusted p value cutoff, the Benjamin-Hochberg correction was used to calculate.
Supplementary Figure 6 Kinetics and regulation of Myc and CD122 expression.
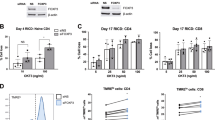
(a) Flow cytometric analysis of CellTrace Violet dilutions of GFP- (WT) and GFP+ (AAA) CD4+ T cells from CD4Cre-ERT2 Foxo1AAA/+ (iAAA) mice following 1 and 2 days in culture with plate-bound anti-CD3 and soluble anti-CD28. Representative of 4 independent experiments, with similar results (b) Flow cytometric analysis on days 1-3 of isolated cultures stimulated as in a. Open histograms are GFP- and GFP+ populations as indicated, lightly shaded histograms show isotype control staining for each population. Representative of 4 independent experiments, with similar results (c) Real-time quantitative PCR from iso-cultures of Myc at 0 and 24 hrs post-activation with plate-bound anti-CD3 and soluble anti-CD28 (normalized to beta-Actin; n = 5). (d) Real-time quantitative PCR from iso- and co-cultures of Myc at 0 (n = 2) and 72 hrs (n = 3) post-activation with plate-bound anti-CD3 and soluble anti-CD28 (normalized to beta-Actin). Following 3 days in culture, RNA was extracted directly from iso-cultures, while co-cultures were sorted once again for GFP- and GFP+ populations, and RNA was subsequently extracted. (e) Real-time quantitative PCR from iso-cultures of IL-2Rb at 0 and 48 hrs post-activation stimulated as in a (normalized to beta-Actin; n = 4). (f) Western blot analysis of iso-cultures stimulated for 1 and 2 days as in a, representative of three independent experiments. After sorting, a proportion of GFP- and GFP+ cells were immediately lysed (Day 0). (g) Flow cytometric analysis of surface (non-permeablized cells) and total levels (intracellular and surface levels of fixed and permeablized cells; ICS) of CD122 at 0 and 48 hrs post-activation of iso-cultures stimulated as in a, representative of two independent experiments. For all graphs, quantification used a two-tailed Student’s t-test, with no adjustments made for multiple comparisons; centre value is the mean, with error bars indicating standard deviation. * P < 0.05; ** P < 0.005; ns = not significant.
Supplementary Figure 7 Multiple potential mechanisms to account for hyperactivation of Akt.
(a) Western blot analysis and quantitation (normalized to beta-Actin) of iso-cultures stimulated 2 days with plate-bound anti- CD3 and soluble anti-CD28 (n = 8). (b,c) Western blot analysis of iso-cultures stimulated for 1 to 2 days with platebound anti-CD3 and soluble anti-CD28, representative of three independent experiments, with similar results. After sorting, a proportion of GFP- and GFP+ cells were immediately lysed (Day 0). (d) Western blot analysis and quantitation of iso-cultures stimulated for 1 and 2 days, as in b and c, with either vehicle or an inhibitor (NU7117; 10 mm) of DNA-PK. Representative of two independent experiments, with similar results. (e,f) Flow cytometric analysis and quantitation of GFP- and GFP+ iso-cultures stimulated for 1 and 2 days (representative flow plots shown on day 2) with anti-CD3/28 (n = 8). (g) Western blot analysis and quantitation of iso-cultures stimulated for 1 and 2 days, as in b and c. Quantitation is for PTEN expression normalized to beta-Actin on day 2 (n = 7). For all graphs, quantification used a two-tailed Student’s t-test, with no adjustments made for multiple comparisons; centre value is the mean, with error bars indicating standard deviation. **P < 0.005; *** P < 0.0005; **** P < 0.0001.
Supplementary Figure 8 Altered lysosome and mTORC1 colocalization in nutrient-deplete and nutrient-replete conditions.
Representative immunofluorescence images of mTOR, Lamp-1 and DAPI staining on day 2 of isocultures stimulated with plate-bound anti-CD3 and soluble anti-CD28. 2 hrs prior to fixation and subsequent staining and imaging, cells were adhered to CellTak coated coverslips and incubated in RPMI media with (“Complete media”) or without amino acids, glucose and 10% fetal calf serum (“Base media”). Representative of three independent experiments, with similar results. Scale bars, 10 μm.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8
Rights and permissions
About this article
Cite this article
Newton, R.H., Shrestha, S., Sullivan, J.M. et al. Maintenance of CD4 T cell fitness through regulation of Foxo1. Nat Immunol 19, 838–848 (2018). https://doi.org/10.1038/s41590-018-0157-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-018-0157-4
This article is cited by
-
A genome-wide in vivo CRISPR screen identifies essential regulators of T cell migration to the CNS in a multiple sclerosis model
Nature Neuroscience (2023)
-
The effects of MYC on tumor immunity and immunotherapy
Cell Death Discovery (2023)
-
FOXO1 regulates Th17 cell-mediated hepatocellular carcinoma recurrence after hepatic ischemia-reperfusion injury
Cell Death & Disease (2023)
-
Ionic mitigation of CD4+ T cell metabolic fitness, Th1 central nervous system autoimmunity and Th2 asthmatic airway inflammation by therapeutic zinc
Scientific Reports (2022)
-
Hyperactive PI3Kδ predisposes naive T cells to activation via aerobic glycolysis programs
Cellular & Molecular Immunology (2021)