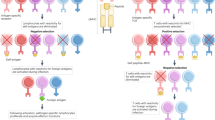
Abstract
Although the thymus has long been recognized as a key organ for T cell selection, the intricate details linking these selection events to human autoimmunity have been challenging to decipher. Over the last two decades, there has been rapid progress in understanding the role of thymic tolerance mechanisms in autoimmunity through genetics. Here we review some of the recent progress in understanding key thymic tolerance processes that are critical for preventing autoimmune disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Kappler, J. W., Roehm, N. & Marrack, P. T cell tolerance by clonal elimination in the thymus. Cell 49, 273–280 (1987).
Kisielow, P., Blüthmann, H., Staerz, U. D., Steinmetz, M. & von Boehmer, H. Tolerance in T-cell-receptor transgenic mice involves deletion of nonmature CD4+8+ thymocytes. Nature 333, 742–746 (1988).
Anderson, M. S. et al. Projection of an immunological self shadow within the thymus by the Aire protein. Science 298, 1395–1401 (2002).
Derbinski, J., Schulte, A., Kyewski, B. & Klein, L. Promiscuous gene expression in medullary thymic epithelial cells mirrors the peripheral self. Nat. Immunol. 2, 1032–1039 (2001).
Derbinski, J. et al. Promiscuous gene expression in thymic epithelial cells is regulated at multiple levels. J. Exp. Med. 202, 33–45 (2005).
Finnish-German APECED Consortium. An autoimmune disease, APECED, caused by mutations in a novel gene featuring two PHD-type zinc-finger domains. Nat. Genet. 17, 399–403 (1997).
Nagamine, K. et al. Positional cloning of the APECED gene. Nat. Genet. 17, 393–398 (1997).
Perheentupa, J. Autoimmune polyendocrinopathy-candidiasis-ectodermal dystrophy. J. Clin. Endocrinol. Metab. 91, 2843–2850 (2006).
Husebye, E. S., Perheentupa, J., Rautemaa, R. & Kämpe, O. Clinical manifestations and management of patients with autoimmune polyendocrine syndrome type I. J. Intern. Med. 265, 514–529 (2009).
Kisand, K. et al. Chronic mucocutaneous candidiasis in APECED or thymoma patients correlates with autoimmunity to Th17-associated cytokines. J. Exp. Med. 207, 299–308 (2010).
Puel, A. et al. Autoantibodies against IL-17A, IL-17F, and IL-22 in patients with chronic mucocutaneous candidiasis and autoimmune polyendocrine syndrome type I. J. Exp. Med. 207, 291–297 (2010).
Meloni, A. et al. Autoantibodies against type I interferons as an additional diagnostic criterion for autoimmune polyendocrine syndrome type I. J. Clin. Endocrinol. Metab. 93, 4389–4397 (2008).
Meyer, S. et al. AIRE-deficient patients harbor unique high-affinity disease-ameliorating autoantibodies. Cell 166, 582–595 (2016).
Heino, M. et al. RNA and protein expression of the murine autoimmune regulator gene (Aire) in normal, RelB-deficient and in NOD mouse. Eur. J. Immunol. 30, 1884–1893 (2000).
Liston, A. et al. Gene dosage–limiting role of Aire in thymic expression, clonal deletion, and organ-specific autoimmunity. J. Exp. Med. 200, 1015–1026 (2004).
Gardner, J. M. et al. Deletional tolerance mediated by extrathymic Aire-expressing cells. Science 321, 843–847 (2008).
Gardner, J. M. et al. Extrathymic Aire-expressing cells are a distinct bone marrow–derived population that induce functional inactivation of CD4+ T cells. Immunity 39, 560–572 (2013).
Yamano, T. et al. Thymic B cells are licensed to present self antigens for central T cell tolerance induction. Immunity 42, 1048–1061 (2015).
Liston, A., Lesage, S., Wilson, J., Peltonen, L. & Goodnow, C. C. Aire regulates negative selection of organ-specific T cells. Nat. Immunol. 4, 350–354 (2003).
Anderson, M. S. et al. The cellular mechanism of Aire control of T cell tolerance. Immunity 23, 227–239 (2005).
Taniguchi, R. T. et al. Detection of an autoreactive T-cell population within the polyclonal repertoire that undergoes distinct autoimmune regulator (Aire)-mediated selection. Proc. Natl. Acad. Sci. USA 109, 7847–7852 (2012).
Malchow, S. et al. Aire-dependent thymic development of tumor-associated regulatory T cells. Science 339, 1219–1224 (2013).
Perry, J. S. A. et al. Distinct contributions of Aire and antigen-presenting-cell subsets to the generation of self-tolerance in the thymus. Immunity 41, 414–426 (2014).This study examines T reg cell–specific TCRs in a limited repertoire system and identifies TCRs that depend on Aire for their selection.
Yang, S., Fujikado, N., Kolodin, D., Benoist, C. & Mathis, D. Regulatory T cells generated early in life play a distinct role in maintaining self-tolerance. Science 348, 589–594 (2015).This study invokes another layer of a tolerance defect in the Aire-deficient model where T reg cells are inappropriately selected early in life that then help protect against autoimmune responses in the tissues.
Landegren, N. et al. Proteome-wide survey of the autoimmune target repertoire in autoimmune polyendocrine syndrome type 1. Sci. Rep. 6, 20104 (2016).
Alimohammadi, M. et al. Autoimmune polyendocrine syndrome type 1 and NALP5, a parathyroid autoantigen. N. Engl. J. Med. 358, 1018–1028 (2008).
Alimohammadi, M. et al. Pulmonary autoimmunity as a feature of autoimmune polyendocrine syndrome type 1 and identification of KCNRG as a bronchial autoantigen. Proc. Natl. Acad. Sci. USA 106, 4396–4401 (2009).
Shum, A. K. et al. BPIFB1 is a lung-specific autoantigen associated with interstitial lung disease. Sci. Transl. Med. 5, 206ra139 (2013).
Pöntynen, N. et al. Aire deficient mice do not develop the same profile of tissue-specific autoantibodies as APECED patients. J. Autoimmun. 27, 96–104 (2006).
Gäbler, J., Arnold, J. & Kyewski, B. Promiscuous gene expression and the developmental dynamics of medullary thymic epithelial cells. Eur. J. Immunol. 37, 3363–3372 (2007).
Gray, D., Abramson, J., Benoist, C. & Mathis, D. Proliferative arrest and rapid turnover of thymic epithelial cells expressing Aire. J. Exp. Med. 204, 2521–2528 (2007).
Hamazaki, Y. et al. Medullary thymic epithelial cells expressing Aire represent a unique lineage derived from cells expressing claudin. Nat. Immunol. 8, 304–311 (2007).
Gray, D. H. D. et al. Developmental kinetics, turnover, and stimulatory capacity of thymic epithelial cells. Blood 108, 3777–3785 (2006).
Rossi, S. W. et al. RANK signals from CD4+3– inducer cells regulate development of Aire-expressing epithelial cells in the thymic medulla. J. Exp. Med. 204, 1267–1272 (2007).
Akiyama, T. et al. The tumor necrosis factor family receptors RANK and CD40 cooperatively establish the thymic medullary microenvironment and self-tolerance. Immunity 29, 423–437 (2008).
Hikosaka, Y. et al. The cytokine RANKL produced by positively selected thymocytes fosters medullary thymic epithelial cells that express autoimmune regulator. Immunity 29, 438–450 (2008).
Metzger, T. C. et al. Lineage tracing and cell ablation identify a post-Aire-expressing thymic epithelial cell population. Cell Rep. 5, 166–179 (2013).
Khan, I. S. et al. Enhancement of an anti-tumor immune response by transient blockade of central T cell tolerance. J. Exp. Med. 211, 761–768 (2014).
Kawano, H. et al. Aire expression is inherent to most medullary thymic epithelial cells during their differentiation program. J. Immunol. 195, 5149–5158 (2015).
Bakhru, P. et al. Combination central tolerance and peripheral checkpoint blockade unleashes antimelanoma immunity. JCI Insight 2, 2 (2017). This study shows that central tolerance imposes a barrier to robust tumor responses and that combination therapy including RANK ligand blockade and peripheral tolerance blockade can enhance tumor immune responses.
Meredith, M., Zemmour, D., Mathis, D. & Benoist, C. Aire controls gene expression in the thymic epithelium with ordered stochasticity. Nat. Immunol. 16, 942–949 (2015).
Brennecke, P. et al. Single-cell transcriptome analysis reveals coordinated ectopic gene-expression patterns in medullary thymic epithelial cells. Nat. Immunol. 16, 933–941 (2015).
Pinto, S. et al. Overlapping gene coexpression patterns in human medullary thymic epithelial cells generate self-antigen diversity. Proc. Natl. Acad. Sci. USA 110, E3497–E3505 (2013).
Bansal, K., Yoshida, H., Benoist, C. & Mathis, D. The transcriptional regulator Aire binds to and activates super-enhancers. Nat. Immunol. 18, 263–273 (2017). This study demonstrates that Aire appears to target genomic regions enriched for super-enhancers to help drive its activity to promote TSA expression. Furthermore, this study provides biochemical evidence that an interaction with topoisomerase I may be part of this targeting property.
Guha, M. et al. DNA breaks and chromatin structural changes enhance the transcription of autoimmune regulator target genes. J. Biol. Chem. 292, 6542–6554 (2017).
Abramson, J., Giraud, M., Benoist, C. & Mathis, D. Aire’s partners in the molecular control of immunological tolerance. Cell 140, 123–135 (2010).
Oven, I. et al. AIRE recruits P-TEFb for transcriptional elongation of target genes in medullary thymic epithelial cells. Mol. Cell. Biol. 27, 8815–8823 (2007).
Giraud, M. et al. Aire unleashes stalled RNA polymerase to induce ectopic gene expression in thymic epithelial cells. Proc. Natl. Acad. Sci. USA 109, 535–540 (2012).
Anderson, M. S. & Su, M. A. AIRE expands: new roles in immune tolerance and beyond. Nat. Rev. Immunol. 16, 247–258 (2016).
Abramson, J. & Anderson, G. Thymic epithelial cells. Annu. Rev. Immunol. 35, 85–118 (2017).
Sansom, S. N. et al. Population and single-cell genomics reveal the Aire dependency, relief from Polycomb silencing, and distribution of self-antigen expression in thymic epithelia. Genome Res. 24, 1918–1931 (2014).
Takaba, H. et al. Fezf2 orchestrates a thymic program of self-antigen expression for immune tolerance. Cell 163, 975–987 (2015). In this study, Fezf2 is identified as an additional transcriptional factor that promotes thymic TSA expression for the induction of immune tolerance.
Guo, C. et al. Fezf2 expression identifies a multipotent progenitor for neocortical projection neurons, astrocytes, and oligodendrocytes. Neuron 80, 1167–1174 (2013).
Cosway, E. J. et al. Redefining thymus medulla specialization for central tolerance. J. Exp. Med. 214, 3183–3195 (2017). This study helps clarify the potential role for lymphotoxin β signaling in thymic medulla specification. Here a more complex role of lymphotoxin receptor β signaling was unraveled, including a role for both mTECs and dendritic cell populations.
Aschenbrenner, K. et al. Selection of Foxp3+ regulatory T cells specific for self antigen expressed and presented by Aire+ medullary thymic epithelial cells. Nat. Immunol. 8, 351–358 (2007).
Lei, Y. et al. Aire-dependent production of XCL1 mediates medullary accumulation of thymic dendritic cells and contributes to regulatory T cell development. J. Exp. Med. 208, 383–394 (2011).
Leonard, J. D. et al. Identification of natural regulatory T cell epitopes reveals convergence on a dominant autoantigen. Immunity 47, 107–117 (2017).This study identifies the antigen specificity for a T reg cell–specific clone that was originally identified in prostate tumors. Tetramer analysis for Tcaf3-specific CD4 + T cells shows a large skewing to the T reg cell lineage and dependency on Aire.
Kuroda, N. et al. Development of autoimmunity against transcriptionally unrepressed target antigen in the thymus of Aire-deficient mice. J. Immunol. 174, 1862–1870 (2005).
Murata, S. et al. Regulation of CD8+ T cell development by thymus-specific proteasomes. Science 316, 1349–1353 (2007).
Nitta, T. et al. Human thymoproteasome variations influence CD8 T cell selection. Sci. Immunol. 2, 2 (2017). This translational study finds evidence correlating a SNP risk variant for Sjögren’s syndrome with activity of the thymoproteosome for positive selection. These results suggest that this could be part of the mechanism by which autoimmunity is triggered in this disease.
Sakaguchi, N. et al. Altered thymic T-cell selection due to a mutation of the ZAP-70 gene causes autoimmune arthritis in mice. Nature 426, 454–460 (2003).
Tanaka, S. et al. Graded attenuation of TCR signaling elicits distinct autoimmune diseases by altering thymic T cell selection and regulatory T cell function. J. Immunol. 185, 2295–2305 (2010).
Hsu, L.-Y., Tan, Y. X., Xiao, Z., Malissen, M. & Weiss, A. A hypomorphic allele of ZAP-70 reveals a distinct thymic threshold for autoimmune disease versus autoimmune reactivity. J. Exp. Med. 206, 2527–2541 (2009).
Sakaguchi, S., Benham, H., Cope, A. P. & Thomas, R. T-cell receptor signaling and the pathogenesis of autoimmune arthritis: insights from mouse and man. Immunol. Cell Biol. 90, 277–287 (2012).
Chan, A. Y. et al. A novel human autoimmune syndrome caused by combined hypomorphic and activating mutations in ZAP-70. J. Exp. Med. 213, 155–165 (2016).This study identifies a single family with inheritance of an autoimmune syndrome that correlated with the presence of two mutant ZAP70 alleles. This family provides evidence of hypomorphic Zap70 activity in the generation of autoimmunity.
Bruserud, Ø., Oftedal, B. E., Wolff, A. B. & Husebye, E. S. AIRE-mutations and autoimmune disease. Curr. Opin. Immunol. 43, 8–15 (2016).
Marx, A. et al. Thymoma and paraneoplastic myasthenia gravis. Autoimmunity 43, 413–427 (2010).
Marx, A. et al. The autoimmune regulator AIRE in thymoma biology: autoimmunity and beyond. J. Thorac. Oncol. 5(Suppl. 4), S266–S272 (2010).
Cheng, M. H. et al. Acquired autoimmune polyglandular syndrome, thymoma, and an AIRE defect. N. Engl. J. Med. 362, 764–766 (2010).
Liu, Y. et al. Autoimmune regulator expression in thymomas with or without autoimmune disease. Immunol. Lett. 161, 50–56 (2014).
Meager, A. et al. Anti-cytokine autoantibodies in autoimmunity: preponderance of neutralizing autoantibodies against interferon-α, interferon-ω and interleukin-12 in patients with thymoma and/or myasthenia gravis. Clin. Exp. Immunol. 132, 128–136 (2003).
Wolfe, G. I. et al. Randomized trial of thymectomy in myasthenia gravis. N. Engl. J. Med. 375, 511–522 (2016).
Vafiadis, P. et al. Insulin expression in human thymus is modulated by INS VNTR alleles at the IDDM2 locus. Nat. Genet. 15, 289–292 (1997).
Giraud, M. et al. An IRF8-binding promoter variant and AIRE control CHRNA1 promiscuous expression in thymus. Nature 448, 934–937 (2007).
Renton, A. E. et al. A genome-wide association study of myasthenia gravis. JAMA Neurol. 72, 396–404 (2015).
Parent, A. V. et al. Generation of functional thymic epithelium from human embryonic stem cells that supports host T cell development. Cell Stem Cell 13, 219–229 (2013).
Acknowledgements
M.S.A. is supported by the US National Institutes of Health, the Helmsley Charitable Trust, the California Institute of Regenerative Medicine and the Larry L. Hillblom Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Cheng, M., Anderson, M.S. Thymic tolerance as a key brake on autoimmunity. Nat Immunol 19, 659–664 (2018). https://doi.org/10.1038/s41590-018-0128-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-018-0128-9
This article is cited by
-
Autoantibodies against type I IFNs in humans with alternative NF-κB pathway deficiency
Nature (2023)
-
PTEN directs developmental and metabolic signaling for innate-like T cell fate and tissue homeostasis
Nature Cell Biology (2022)
-
Thymocyte regulatory variant alters transcription factor binding and protects from type 1 diabetes in infants
Scientific Reports (2022)
-
Myasthenia gravis-specific aberrant neuromuscular gene expression by medullary thymic epithelial cells in thymoma
Nature Communications (2022)
-
Structural constraints in T-cell repertoire selection predicted by machine learning
Genes & Immunity (2021)