Abstract
Transcriptional profiles and host-response biomarkers are used increasingly to investigate the severity, subtype and pathogenesis of disease. We now describe whole-blood mRNA signatures and concentrations of local and systemic immunological mediators in 131 adults hospitalized with influenza, from whom extensive clinical and investigational data were obtained by MOSAIC investigators. Signatures reflective of interferon-related antiviral pathways were common up to day 4 of symptoms in patients who did not require mechanical ventilator support; in those who needed mechanical ventilation, an inflammatory, activated-neutrophil and cell-stress or death (‘bacterial’) pattern was seen, even early in disease. Identifiable bacterial co-infection was not necessary for this ‘bacterial’ signature but was able to enhance its development while attenuating the early ‘viral’ signature. Our findings emphasize the importance of timing and severity in the interpretation of host responses to acute viral infection and identify specific patterns of immune-system activation that might enable the development of novel diagnostic and therapeutic tools for severe influenza.
Similar content being viewed by others
Main
Influenza viruses present a continuous threat to global health, mutating and spreading within and between species. It is estimated that one billion cases of human influenza occur worldwide each year, causing three million to five million cases of severe illness and 300,000 to 500,000 deaths1. Infection with pandemic 2009 H1N1 influenza A virus (pH1N1) resulted in generally mild disease2 but still caused an estimated 250,000–500,000 additional deaths during the first 12 months of global circulation3. Whereas seasonal influenza commonly results in severe disease in the old and infirm, serious pH1N1 disease occurred mostly in infants and younger adults, presenting as viral pneumonia and sometimes complicated by multi-organ failure4,5. It has been suggested that severe influenza might result in part from an over-exuberant host reaction to infection (sometimes called a ‘cytokine storm’) but is also driven by a high viral load in affected persons6,7,8.
Although analysis of transcriptional signatures and levels of mediators has helped to clarify the pathogenesis of severe influenza, the relationship among the severity, timing and complications of infection has remained unclear. Published studies of gene-expression patterns in influenza have typically involved small numbers of patients, healthy subjects undergoing experimental challenge or patients suffering from mild disease9,10,11,12,13,14,15. Transcriptomic analysis has also been used to study a variety of acute and chronic infections, including bacterial sepsis, infection with dengue virus and tuberculosis16, and to assess differences and similarities between infectious disorders and non-infectious inflammatory disorders, such as systemic lupus erythematosus17.
To further elucidate the pathogenesis of influenza, the Mechanisms of Severe Acute Influenza Consortium (MOSAIC) recruited 255 patients hospitalized with suspected influenza in England over two consecutive seasons (2009–2010 and 2010–2011) and 155 adult healthy control subjects (study design, https://goo.gl/kyY2Eu). By analyzing biological samples obtained at multiple time-points and correlating those analyses with extensive clinical data, MOSAIC aimed to define the contributions made by sequence variation in influenza virus, co-pathogens (non-influenza viruses and bacteria) and host factors (genetic and transcriptional differences, soluble mediator responses and cellular immune responses) to disease pathogenesis. Sample analysis resulted in a cumulative total of 2.1 × 107 data items for this population, a dataset that we now describe in outline and provide as a resource. So far, MOSAIC has reported enrichment for a host genetic variant, the single-nucleotide polymorphism (SNP) rs12252-C in the allele encoding the antiviral molecule IFITM3 (‘interferon-inducible transmembrane protein 3’), in some patients hospitalized with influenza18 and has reported that changes in viral sequence that accumulate over time might contribute to the variation in disease severity19,20,21,22. The exceptional size and depth of the MOSAIC study provides a unique database that allows such complex issues to be resolved.
We now describe the use of data on whole-blood transcriptional mRNA and soluble mediators to define associations between individual responses to infection and clinical and laboratory findings in adult patients of the MOSAIC study in whom infection with influenza virus was confirmed. Transcriptomic patterns and mediator levels were strongly associated with both the severity of illness and its duration, indicative of a phased and graded activation of genes encoding interferon-related and inflammatory molecules; the effects of clinically evident bacterial co-infection were superimposed on these patterns that were, however, related mainly to the duration and severity of influenza.
Results
Clinical cohorts
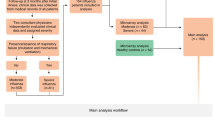
Adult patients with laboratory-confirmed influenza were recruited in 2009–2010 (n = 22 patients) and 2010–2011 (n = 22 109). The majority were infected with pH1N1 influenza virus (95.5% in 2009–2010, and 86.2% in 2010–2011). In each cohort, the majority of patients had at least one comorbidity (81.8% in 2009–2010, and 74.3% in 2010–2011). 13.6% participants in 2009–2010 and 25.7% patients in 2010–2011 had illness categorized as severity 3 (described below) at the first sampling time point (T1) (Table 1).
Whole-blood transcriptomics
Principal-component analysis of the 18,974 most abundant transcripts among whole-blood RNA at enrollment (time 1 (T1): 2010–2011 season; 109 cases) showed clustering distinct from that of age-, ethnicity- and sex-matched healthy control subjects (n = 130). There was no discernible difference between patients infected with influenza A virus and those infected with influenza B virus (Fig. 1a). Samples at the final time point (time 3 (T3): > 4 weeks after T1) obtained from patients who achieved clinical resolution were similar to those from healthy control subjects, but they remained abnormal in patients who remained unwell (data not shown).
a, Principal-component analysis of all transcripts significantly above background in at least 10% of samples from healthy control subjects (HC) (n = 130) or patients with influenza A (H1N1 or H3N2; n = 97) or influenza B (n = 12), all from the 2010–2011 cohort (key). b, Modular analysis of patients with influenza in the 2010–2011 cohort (left), showing probes over-represented (Over-rep) or under-represented (Under-rep) relative to their representation in healthy control subjects (key; proportion of genes in each module with significantly differential expression), plus identification of the corresponding modules (right). c, Weighted MDTH24 of patients with influenza relative to that of healthy control subjects, all in the 2010–2011 cohort, for 4,526 transcripts with significant detection above background, filtered for low expression (transcripts retained with a change of over twofold from median normalized intensity value in more than 10% of all samples); results are presented as a box-whisker plot (center line, median; box limits, interquartile range; extended lines, maximum and minimum). P < 0.0001 (Mann-Whitney test). d, Transcript intensity (normalized values) for 1,255 transcripts in patients with influenza and healthy control subjects (below plot), filtered for low expression, then statistically filtered (P < 0.01 (Mann-Whitney test with Bonferroni multiple testing correction)), followed by a filter for a change between groups (transcripts retained with a change of over twofold between any two groups); right margin, top five canonical pathways in Ingenuity pathway analysis (by significance: P < 0.05 (Fisher’s Exact test)) to which upregulated and downregulated transcripts belong. e, Transcript intensity (normalized values) for the top 25 significant transcripts (right margin) with a change in expression (fold values) between healthy control subjects and patients with influenza (below plot), plus hierarchical clustering on entities (above plot) or subjects or patients (left margin) (Pearson’s uncentered (cosine) with averaged linkage).
Modular analysis23 of the samples from the 2010–2011 cohort showed a greater abundance of transcripts from the genes in the interferon-inducible module (M3.1) and neutrophil module (M2.2) in patients with influenza than in healthy control subjects (Fig. 1b). Transcripts representing plasma cells (module M1.1), a subset of myeloid-lineage genes (module M2.6) and two inflammation modules (M3.2 and M3.3) were also increased, while expression of genes in a T cell module (M2.8) and B cell module (M1.3) was decreased (Fig. 1b). The calculated index ‘molecular distance to health’ (MDTH) (derived from analysis of 4,526 transcripts significantly above background, filtered for low expression24) was higher in most patients with influenza than in healthy control subjects (Fig. 1c), although this was affected by disease stage and severity (discussed below). A combination of expression-level and statistical filtering identified 1,255 transcripts expressed differentially in patents relative to their expression in healthy control subjects. Supervised hierarchical clustering revealed transcripts that were over- or under-expressed in patients with influenza relative to their expression in healthy control subjects (Fig. 1d); applying this same 1,255-transcript set to the 2009–2010 cohort (22 patients with influenza and 25 matched healthy control subjects) replicated these profiles (Supplementary Fig. 2), which indicated that viral variation between the two seasons22 did not appreciably affect transcriptomic patterns.
Ingenuity pathway analysis identified the top five canonical pathways associated with upregulated and downregulated transcripts (Fig. 1d). Transcripts upregulated in patients with influenza were associated with the categories ‘interferon signaling genes’ (including IFITM1, IFI35, IFIT1, OAS1, IFIT3 and IFI35; Supplementary Fig. 1), ‘activation of pattern-recognition receptors by bacteria and/or viruses’, ‘activation of IRF by cytosolic pattern-recognition receptors’, ‘hepatic fibrosis–hepatic stellate cell activation’ and ‘IL-6 signaling’. Transcripts that were downregulated were those associated with the categories ‘ICOS–ICOSL signaling in T helper cells’, ‘primary immunodeficiency signaling’, ‘role of NFAT in regulation of the immune response’, ‘OX40 signaling pathway’ and ‘T cell receptor signaling’ (Fig. 1d).
Hierarchical clustering of the top 25 most significant transcripts in the 2010–2011 group of patients with influenza showed two major clusters (Fig. 1e). Transcripts of the interferon-stimulated gene IFI27 were overexpressed in almost all cases, while transcription of FCER1A was usually decreased. Independent analysis of the 25 transcripts from the 2009–2010 dataset showed similar clustering (Supplementary Fig. 2b). Patients with activation of type I interferon–induced genes (for example, RSAD2, IFI6 and IFI44L) typically did not express transcripts encoding neutrophil-associated or bacterial response–associated molecules (for example, DEFA4, ELANE and MMP8) and vice versa (Fig. 1e and Supplementary Fig. 2b). Together these results showed that patients with acute influenza had activation of whole-blood transcriptomic pathways indicative of responses to type 1 and type 2 interferons and of inflammatory markers and possibly combined with the effects of depletion of some cell types from the blood.
Transcriptomics and disease severity
Patients in the 2010–2011 cohort were grouped according to their severity of illness at T1 by the following three-point scale: 1, no supplemental oxygen required; 2, oxygen by mask required; 3, mechanical ventilation required. Transcriptomic abnormality (mean MDTH; 4,526 transcripts) was greater in patients with illness categorized as severity 1–2 than in healthy control subjects and was further increased in patients with illness categorized as severity 3 (Fig. 2a). By modular analysis23, there was an over-abundance of transcripts in the plasma-cell module (M1.1), neutrophil module (M2.2) and myeloid lineage module (M2.6) in influenza virus–infected patients, a result that was most evident in those with illness of the greatest severity. Patients with illness categorized as severity 3 also showed an abundance of transcripts in inflammation modules M3.2 and M3.3. In contrast, interferon-related transcripts (module M3.1) were most evident in patients with illness categorized as severity 1 or 2 (Fig. 2b). Therefore, patients with the most-severe illness had transcriptomic patterns that differed from those with less-severe illness, with an increased abundance of inflammation-related transcripts and a decrease in interferon-related transcripts.
a, Weighted MDTH of healthy control subjects (n = 130) and of patients with influenza (n = 109), grouped by severity of illness (horizontal axis: 1 (n = 47), 2 (n = 34) or 3 (n = 28)), all in the 2010–2011 cohort; results based on 4,526 transcripts that were significantly differentially expressed relative to background and filtered for low expression (transcripts retained as in Fig. 1c; presented as in Fig. 1c). b, Modular analysis of patients with influenza (n = 109) grouped by severity (above plots), relative to results for healthy control subjects (n = 130), all in 2010–2011 cohort (left), and corresponding modules (right) (presented as in Fig. 1b).
Semi-supervised hierarchical clustering of 231 transcripts with a change in expression of more than twofold between patients with illness categorized as severity 1–2 and those with illness categorized as severity 3 showed that the expression of transcripts associated with the gene-ontology (GO) term ‘response to virus’ was typical of that in patients with milder disease (Fig. 3a; Supplementary Table 1), whereas patients who needed mechanical ventilation (severity 3) showed a marked abundance of transcripts associated with the GO term ‘response to bacterium’ (Supplementary Table 2). Patients with severe illness typically showed a relative under-abundance of transcripts associated with the GO term ‘cellular defense response’ (Fig. 3a).
a, Transcript intensity (normalized values) of 231 transcripts from healthy control subjects (n = 130) and patients with influenza of severity 1 (n = 47), severity 2 (n = 34) or severity 3 (n = 28) (below plot), all in the 2010–2011 cohort, obtained by filtering for low expression, followed by statistical filtering (P < 0.01 (Kruskal-Wallis test with Bonferroni multiple-testing correction)) and then filtering for a change between groups (restricted to initial samples at T1; transcripts retained with a change of over twofold between severity 3 and severity 1 and 2); right margin, top GO terms for the three main subdivisions of the dendrogram at left (clustered by Pearson’s uncentered (cosine) with average linkage rule). b, Weighted molecular score of the 112 ‘bacterial response’ transcripts, plotted against molecular score of the 51 ‘viral response’ transcripts, for patients with influenza (n = 109) at T1 (severity of illness, key; confirmed bacteremia, circled symbols), presented relative to scores for healthy control subjects (n = 130). c,d, Ingenuity pathway analysis of biofunctions significantly activated (z-score > 2) (c) or significantly repressed (d) (z-score < 2) in patients with influenza, relative to their activity in healthy control subjects (shading intensity of symbols along perimeter indicates the degree of up- or downregulation; key at bottom left in c), identified by analysis of 231 transcripts, showing selected networks of biofunctional genes; arrows indicate interactions (key at bottom right in c: dashed, indirect; solid, direct).
The same 231-transcript list noted above was verified by hierarchical clustering analysis of the 2009–2010 cohort. Patients with influenza of severity 1 or 2 were again characterized by transcripts associated with the GO term ‘response to virus’, whereas three patients with influenza of severity 3 instead showed transcripts associated with the GO term ‘response to bacterium’ (Supplementary Fig. 2c). We determined the relationship between total ‘molecular score’ for the 51 transcripts associated with the GO term ‘response to virus’ (Supplementary Table 1) and the 112 transcripts associated with the GO term ‘response to bacterium’ (Supplementary Table 2) from patients in the 2010–2011 cohort, at T1 (n = 109) (Fig. 3b). Patients with influenza who had high ‘viral responses’ ( > 500) were exclusively from the groups with illness of severity 1 or 2, whereas most patients with high ‘bacterial scores’ ( > 500) had illness of severity 3 and had low ‘viral scores’ (reflective of the modular analysis). However, a few patients with illness of severity 1 or 2 had low ‘viral molecular scores’ and moderately high ‘bacterial molecular scores’. The removal of six patients with known bacterial co-infection did not eliminate this subgroup. Similar findings were obtained for the 2009–2010 cohort (Supplementary Fig. 2d).
Reciprocal expression was observed for activated and repressed biofunctions of the 231 differentially expressed genes in patients with illness of severity 3, compared with their expression in patients with illness of severity 1 or 2 (Fig. 3c,d). Nine genes encoding molecules associated with neutrophil activation were upregulated (for example, MPO, DEFA1 and ELANE), along with three genes encoding molecules associated with leukocyte influx (MPO, MMP9 and LCN2) in a similar comparison (Fig. 3c). The repressed biofunctions in patients with illness of severity 3 were in the categories ‘activation of cytotoxic T cells’, ‘adhesion of immune cells’ and ‘quantity of leukocytes’ (Fig. 3d). These results showed that patients with the most-severe disease show upregulation of genes encoding products associated with neutrophil activation and leukocyte influx that was not seen in those with less-severe influenza.
Effect of illness duration, severity and viral load on transcriptomic patterns
Patients with symptoms of up to 4 days’ duration at the time of sampling typically had elevated ‘viral molecular scores’, but not if they required mechanical ventilation (severity 3); in such cases, the ‘viral score’ was low, even early in the disease (Fig. 4a). Patients with illness of severity 3 showed higher ‘bacterial molecular scores’ than those of patients with less-severe disease, even at first presentation, whereas ‘bacterial molecular scores’ were low in patients with illness of severity 1 or 2 regardless of the time of sampling (Fig. 4b).
MDTH scores (according to GO terms, as in Fig. 3), presented as’viral molecular score’ (a,c,e) or ‘bacterial molecular score’ (b,d,f) assessed at T1 and T2 for patients with confirmed influenza (2010–2011 cohort; n = 109), stratified by severity of illness (key) (a,b) or the presence (Bac+) (n = 39; 63 samples) or absence (Bac–) (n = 34; 52 samples) of clinically significant bacterial co-infection (key) (c–f), plotted against time of illness (a–d) or the concentration of PCT in blood (e,f). Loess fitting was used to interpolate and estimate mean values non-parametrically from the data (solid lines); dashed lines indicate estimated 95% confidence interval values of the mean (Pearson: r = 0.44; P < 0.001). P values (top left or right), deviance of generalized linear models (χ2 test).
In patients in the 2010–2011 cohort with repeat samples (T1 and T2, separated by 2–5 d; n = 59), the ‘viral molecular score’ usually (but not always) decreased between T1 and T2 (Supplementary Fig. 3a). In patients for whom samples at T2 were obtained 2 d after T1 (n = 41), the reduction in ‘viral score’ was significant (P = 0.0002 (two-tailed Mann-Whitney test); Supplementary Fig. 3b). Changes between T1 and T2 in ‘bacterial molecular scores’ were more variable (Supplementary Fig. 3c,d). A decrease in viral load (measured in mucus obtained by nasopharyngeal suction) was observed between T1 and T2 (Supplementary Fig. 3e), but there was no clear relationship between viral load and ‘viral transcriptomic score’ (Supplementary Fig. 3f). Together these results showed that the relative dominance of ‘viral’ or ‘bacterial’ transcriptomic responses was influenced by both the severity of illness and the duration of illness.
Effect of bacterial infection and carriage on transcriptomic patterns
To investigate the role of bacterial infection in driving ‘bacterial’ GO terms, we identified a subgroup of influenza virus–infected patients who had been thoroughly investigated for bacterial infection by analysis of the nasopharyngeal aspirate (NPA) and throat swabs at T1 by culture and analysis of the NPA at T1 by detection of bacterial pathogens via PCR (in addition to testing for pneumococcal antigen in blood cultures and urine for most patients). Incomplete bacteriological sampling excluded 36 of 109 patients (33%); 34 patients (47%) provided at least four of five sample types. Of the 73 cases with adequate samples, 39 (53%) were deemed to have potentially pathogenic bacteria detected in at least one sample type and were classified by an expert clinical review panel to have clinically relevant bacterial co-infection.
Comparison of those patients with influenza in whom clinically relevant bacterial co-infection was identified with those in whom no bacterial infection was found despite adequate investigation showed that the average ‘viral molecular score’ was lower in those with bacterial infection at all times up to day 12 after the onset of illness (Fig. 4c), and the average ‘bacterial score’ was greater in those with bacterial co-infection between day 3 and day 14 (Fig. 4d). However, the transcriptomic scores showed similar time trends regardless of the presence or absence of bacterial co-infection. Similar findings were obtained when stricter exclusion criteria were applied to the subgroup analysis, with the exclusion of patients from the ‘bacteria not detected’ group if they had not provided all five sample types (data not shown). In this case, statistical analysis could not be performed due to the small sample size (only 13 patients provided all five sample types and did not have bacteria detected).
To assess the influence of treatment of bacterial infection on the observed ‘viral’ and ‘bacterial’ responses, we stratified ‘bacterial scores’ and ‘viral scores’ at T1 and T2 in patients with influenza in the 2010–2011 cohort according to antibiotics prescription. In the MOSAIC study, 92% of the patients (234 of 255) were treated with antibiotics at some time. Antibiotics before T1 had no demonstrable effect on transcriptomic patterns (Supplementary Fig. 4a). Comparison of patients with influenza who were not given antibiotics (n = 7) with those given sustained antibiotic treatment after T1 (n = 24) or throughout illness (including T1 and T2; n = 27) showed that there was no discernible or statistically significant effect of antibiotic administration on the ‘bacterial molecular scores’ (Supplementary Fig. 4b).
We next assessed the abundance of 16 S rRNA transcripts (indicative of bacterial load) in the throat-swab and NPA samples of patients classified as having ‘bacterial co-infection’ or ‘viral infection without bacterial infection’. The abundance of the 16 S rRNA in throat swabs was not different in these groups, but the bacterial load in NPA samples was greater in those patients with confirmed bacterial co-infection (Supplementary Fig. 4c).
Finally, we investigated the utility of procalcitonin (PCT) as a possible guide to the presence of substantial bacterial infection25,26,27. The concentration of PCT showed no relationship with the ‘viral’ molecular score’ (Fig. 4e), and there was no correlation between ‘viral molecular scores’ at T1 and T2 and the concentration of PCT measured at the corresponding time point (data not shown). However, ‘bacterial molecular scores’ tended to be higher in those patients with the most-severe disease and the highest concentration of PCT regardless of the presence or absence of significantly detectable bacteria (Fig. 4f).
In summary, ‘viral molecular scores’ were seen in disease of up to 5 days’ duration. Even during this early phase, patients who needed mechanical ventilation had low ‘viral scores’, and this was especially true in those with clinically determined bacterial co-infection. On the other hand, expression of ‘bacterial response genes’ was seen only in patients with the most-severe influenza; significant bacterial infection enhanced this signal, but the ‘bacterial score’ was evident in those with severe influenza regardless of the presence of bacterial co-infection (especially if the disease had lasted a week or more).
Effect of illness duration, severity and bacterial co-infection on soluble mediators
An advantage of the MOSAIC study is that it provides linked data of whole-blood transcriptomic signatures and, for example, levels of 35 soluble mediators in the blood, NPA and anterior nasal fluid (‘nasadsorption’ samples using synthetic adsorptive matrices) at up to three time points.
The changes observed depended on the mediator and compartment. For example, the concentration of the cytokine IL-1β in serum showed no trend when plotted against disease severity (Fig. 5a), but the concentration of IL-1β in NPA or nasadsorption samples was higher in those with severe disease (Fig. 5b,c). In contrast, the concentration of IL-6 in serum increased with disease severity (Fig. 5d); in NPA samples, IL-6 was undetectable in most of the healthy control subjects but was detected in most of the patients with influenza and increased with disease severity (Fig. 5e). The concentration of IL-6 was more consistent in nasadsorption samples than in NPA samples, was measurable in healthy control subjects, and was higher in most patients with influenza but did not reflect disease severity (Fig. 5f).
Concentration (log scale) of IL-1β (a–c), IL-6 (d–f), CXCL8 (g–i) and IFN-α2a (j–l) in serum (a,d,g,j), NPA samples (b,e,h,k) and nasabsorption eluates (c,f,i,l) (above columns) obtained at T1 from patients with influenza (severity of illness (horizontal axis) at T1) and healthy control subjects (serum: n = 36 (HC), 58 (severity 1), 43 (severity 2) and 31 (severity 3); NPA: n = 35 (HC), 50, (severity 1), 32 (severity 2) and 27 (severity 3); nasabsorption eluate: n = 36 (HC), 60 (severity 1), 43 (severity 2) and 30 (severity 3), presented as box plots (center line, median; box margins, interquartile range; extended lines, range); zero values and values below the lower limit of detection were assigned half the geometric mean lower limit of detection for presentation here (upper limit of detection, 2,500 pg/ml). NS, not significant (P > 0.05); *P < 0.05, **P < 0.01 and ***P < 0.001 (Kruskal-Wallis test with Dunn's post-test).
The concentration of the chemokine CXCL8 in serum tended to be higher in patients with influenza than in healthy control subjects and again increased with disease severity (Fig. 5g); in NPA samples, the concentration of CXCL8 was variable but generally increased with influenza disease severity and tended to saturate the assay (Fig. 5h). The concentration of CXCL8 in in nasadsorption samples was even higher, often saturating the assay even for samples from some healthy control subjects (Fig. 5i). Interferon-α2a (IFN-α2a) was measurable in only a portion of subjects, but its concentration was increased in serum in patients with milder illness (severity 1 or 2) but not in those with severe disease (Fig. 5j). In NPA or nasadsorption samples, the concentration of IFN-α2a was increased in some patients with milder illness (a result that was not statistically significant; Fig. 5k,l).
The concentration of IL-17 in serum increased with severity at T1 (Supplementary Fig. 5a) and was higher in the bronchoalveolar lavage fluid of eight patients (from whom samples were obtained for clinical indications) than in that of healthy control subjects (Supplementary Fig. 5b). In addition, we found a significant positive correlation of the serum concentration of IL-17 (P < 0.001 and r = 0.39 (Spearman); Supplementary Fig. 5c) and TNF (P < 0.001 and r = 0.40 (Spearman); Supplementary Fig. 5d) with MDTH.
As for the effects of timing, the concentrations of the chemokine CXCL10, IL-6 and chemokine CCL2 were elevated in serum from patients with severe influenza, especially between day 5 and day 10 (Fig. 6a,b and data not shown). Proven bacterial co-infection had no evident additional effect on CXCL10 (Fig. 6c), but IL-6 was abundant in serum not only in patients with severe influenza (even early in disease; Fig. 6b) but especially in patients with bacterial co-infection (especially between day 5 and day 10; Fig. 6d).
Concentration of CXCL10 (a,c,e,g) and IL-6 (b,d,f,h) in serum (a–d) and NPA samples (e–h) obtained at T1 and T2 from patients with influenza, stratified by severity of illness (key) (a,b,e,f) or by the presence (n = 39 subjects; 63 samples) or absence (n = 34 subjects; 52 samples) of proven bacterial infection (key) (c,d,g,h), plotted against time of illness (presented as in Fig. 4). P values (top right corners), deviance of generalized linear models (χ2 test).
In the NPA samples, the concentration of most mediators (for example, CXCL10, IL-6, CCL2 and CXCL8) was markedly increased in severe disease and especially after day 4 (Fig. 6e,f and data not shown). The concentration of CXCL10 in NPA samples was again unaffected by confirmed bacterial disease (Fig. 6g), whereas the concentration of IL-6 (and of CCL2 and CXCL8; data not shown) was particularly increased in patients with bacterial co-infection (Fig. 6h). In the nasadsorption samples, the concentration of mediators decreased slowly with time even in less-severe disease; the concentration of CXCL10 was decreased by known bacterial co-infection, but the concentration of IL-6, CCL2 and CXCL8 was unaffected by disease severity or bacterial status (data not shown).
Since the ‘bacterial load’ (assessed as copy number of 16 S rRNA in NPA samples) was greater in patients with clinically relevant bacterial infection than in those without such infection (Supplementary Fig. 4c), we regressed this parameter against viral or bacterial MDTH. There were high values for viral MDTH only in those with lower bacterial loads in the NPA samples (Supplementary Fig. 5e), and high values for bacterial MDTH were seen only in a subset of those with a higher bacterial load (16 S rRNA) in the NPA samples (Supplementary Fig. 5f).
These data were consistent with a role for bacterial load in the inflamed respiratory tract in driving levels of soluble mediators in mucosal fluids and serum as well as transcriptomic signatures in the blood; however, the effects of influenza severity and time after disease onset remained the dominant determinants of host responses.
Discussion
The MOSAIC study is exceptional in including a large number of well-characterized patients hospitalized with influenza, studied prospectively and sampled intensively. We found that whole-blood RNA-expression profiles of patients hospitalized with influenza evolved over time and that the pattern reflected severity. Patients with mild (or early) disease typically showed responses dominated by interferon-inducible genes and type 1 interferons, but that ‘viral’ signature was replaced during severe (or late) disease by a pattern reflective of inflammation and neutrophil activation, more typically associated with the GO term ‘response to bacteria’, including genes encoding regulators of apoptosis and anaerobic metabolism28. The ‘viral’ response was rarely seen in patients beyond day 4, whereas the inflammation–neutrophil activation signal peaked during the second week.
In severe disease, the early ‘viral’ response was typically absent, whereas the ‘bacterial’ signature was present, at enrollment in the study; this was especially so in patients with proven bacterial co-infection but did not depend on it. In addition, the bacterial load in the nasopharynx (quantified by 16 S rRNA copy number) tended to be low if the ‘viral’ signature was evident and was high in those patients in whom an inflammatory cell–activation pattern was seen.
Soluble protein mediators were generally abundant in the serum and nasopharyngeal samples of patients with severe disease, even early after onset. Inflammatory mediators (for example, IL-17, IL-1β and IL-6) were augmented in those with clinically relevant bacterial co-infection, whereas IFN-α levels tended to be low or undetectable in most compartments in those with very severe influenza; however, interferon-related secondary mediators (for example, CXCL10 in serum) were generally most abundant in patients with severe disease. These findings suggest complex interactions between mechanisms for sensing and responding to viruses and those for sensing and responding to bacteria that have evolved over time.
To investigate the issue of bacterial co-infection specifically, we identified patients in whom pathogenic bacteria was found in mucosal samples or blood culture as a subgroup with clinically confirmed bacterial sepsis. Three of these six patients needed mechanical ventilation and had a markedly elevated ‘bacterial signature’ without any increase in ‘viral score’; one patient had elevated ‘bacterial scores’ and ‘viral scores’. The remaining two patients with bacteremia did not have marked elevations in their ‘bacterial scores’; both had mild disease (severity 1). We next used stringent criteria to identify patients with influenza whom we investigated extensively for bacteria and found to be not co-infected, and compared those with patients in whom pathogenic bacteria were identified with certainty. Patients with confirmed bacterial co-infection had higher ‘bacterial molecular scores’ overall, but progression of the transcriptomic signatures was similar over time. Therefore, severe infection with influenza virus alone seemed able to drive the ‘bacterial’ signature, but this response was enhanced by bacterial co-infection. We conclude that transcriptomic data from the blood are an unreliable guide to the presence or absence of bacterial co-infection but need careful interpretation in the context of the timing and severity of disease. We cannot determine the extent to which these changes might be driven by injury caused by influenza virus or by innate sensitivity to resident microbiota that leads to activation of pathways of the TH17 subset of helper T cells triggered by endotoxins from mucosal surfaces29.
In animal models of viral lung disease, dysregulated host immune responses30 and interferon production31 can lead to complex inflammatory responses that contribute to pathogenesis32,33. In macaques, administration of recombinant IFN-α2a initially upregulates the expression of genes encoding antiviral molecules and prevents viral infection, but continued treatment causes desensitization and a paradoxical decrease in the expression of genes encoding antiviral molecules34. These paradoxical immunosuppressive effects can impede viral control35 or trigger inflammation and tissue damage31. In mice, infection with influenza virus causes an early local influx of neutrophils, followed by a virus specific CD8+ T cell response36,37,38. Neutrophils might facilitate the development of this antigen-specific response by guiding influenza virus–specific CD8+ T cells into sites of infection by laying chemokine trails39. Our findings for human influenza are generally compatible with these animal studies.
We have presented here only selected results of an extended study of data on soluble immunological mediators from the MOSAIC cohort. Our main findings were of decreased concentrations of IFN-α2a and increased concentrations of IL-1β, IL-6 and CXCL8 in the nasal and/or serum compartments in patients with severe disease. This apparent reciprocity might relate to the known cross-regulatory functions of IL-1 and type I interferons in experimental models28,40. Our results generally fit with the proposal that levels of mediators such IL-1β, IL-6 and IL-17 are influenced by bacterial co-infection in severe influenza but are not driven by it. However, many additional possible analyses remain to be performed. We chose to illustrate only those most relevant to the transcriptomic analysis and the question of bacterial superinfection. Additional correlations can be explored with our online data as a resource, and we welcome discussions about additional interpretations.
Our study has important limitations. Despite its ambition, scope and intensity, we had limited numbers of repeat samples from individual patients. Our description of trends over time depends largely on summative data and on subjective reporting of the time of disease onset. Ideally, our findings need validation in other time-series studies of simple and complicated acute viral disease with frequent sampling at multiple sites. We were unable to study the early or preclinical phases but were limited to investigation of symptomatic patients presenting with disease of sufficient severity to require hospitalization. Ongoing studies of experimental infection with pH1N1 in volunteers should allow us to overcome some of these limitations.
In summary, virus-induced type I interferon–related pathways were activated during the first 4 d of symptomatic influenza in hospitalized patients. These’viral’ pathways were then downregulated and were replaced by inflammatory, activated-neutrophil and apoptosis-related pathways associated with IL-17 abundance, host-mediated tissue damage and expression of gene clusters in the category ‘response to bacteria’, particularly in patients with a high bacterial load (16 S rRNA) in their nasopharyngeal secretions. In patients with severe illness, the ‘viral’ response was diminished even early in disease, accompanied by an increase in IL-1β and IL-17. These findings emphasize that the stage and severity of disease need to be taken into account in the interpretation of host responses to infection and in the development of potential diagnostic tests to distinguish between possible causes and appropriate therapies.
Methods
Study population and inclusion criteria
Patients ≥16 years of age were recruited during two successive winters (1 December 2009 to 3 March 2011). Patients with suspected influenza were identified by medical or nursing staff or investigators were notified by hospital diagnostic laboratories. Patients in London were recruited from four Imperial College Healthcare NHS Trust hospitals, the Chelsea and Westminster Hospital, and the intensive care unit at the Royal Brompton Hospital (a national referral center for severe respiratory failure). In Liverpool, patients were recruited from the Royal Liverpool, Liverpool Women's and Arrowe Park Hospitals. Patients were included irrespective of prior or concurrent comorbidity (most commonly asthma, pregnancy, immunocompromising conditions or co-infection with other respiratory pathogens), to reflect the populations known to be at greatest risk of severe influenza. Adult healthy control subjects were recruited and matched to the patient cohorts for age, sex and ethnicity and were screened to exclude known illnesses or current use of medications (Registered Clinical Trial NCT00965354).
Research ethics committee approval
The study was approved by the NHS National Research Ethics Service, Outer West London REC (09/H0709/52, 09/MRE00/67). Patients or their legally authorized representatives provided informed consent. Additional adult healthy control subjects were recruited as part of a separate study and consented to their samples’ being used in additional studies (Central London 3 Research Ethics Committee, 09/H0716/41). Informed consent was obtained from all participants, and we complied with all relevant ethical regulations.
Biological sampling
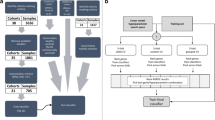
Research samples were obtained at three time points: T1 (recruitment), T2 (approximately 48 h after T1) and T3 (at least 4 weeks after T1). Only samples at T1 and T2 were included in this report. Whole-blood samples for transcriptomics were collected during the two recruitment periods: 2009–2010 and 2010–2011. Of 85 MOSAIC participants presenting with influenza-like illness in 2009–2010, 23 (27%) were adults with confirmed influenza, and T1 transcriptomic samples were available from 22 adults. Of 171 MOSAIC participants presenting with influenza-like illness in 2010–2011, 111 (65%) were adults with confirmed influenza, and T1 transcriptomics samples were available from 109 of 111 (98%). RNA extraction and microarray were successful for all available patient samples from both cohorts. Microarrays were also performed on samples from adult healthy control subjects of age, sex and ethnicity similar to that of the study patients (Table 1). One sample from a healthy control subject in the 2009–2010 cohort was not included in final analysis because it failed quality-control assessments.
Of the 109 adult patients recruited in 2010–2011 and included in this analysis, 94 (86%) were infected with A(H1N1)pdm09 influenza virus, and the remainder were infected with influenza A(H3N2) virus, non-subtyped influenza A virus or influenza B virus. 1 of 22 adult patients recruited during 2009–2010 was infected with A(H3N2) virus; remaining patients were infected with A(H1N1)pdm09 virus. Due to the natural evolution of influenza activity during the 2009–2010 pandemic in the UK, the 2009–2010 cohort was smaller than originally anticipated. Therefore, to assess the host response in the blood transcriptional signature as thoroughly as possible, we focused our analysis on the larger 2010–2011 cohort and then compared those findings with those of the smaller 2009–2010 cohort.
Influenza-virus-infection status
For each participant, influenza-virus-infection status was determined by reverse transcription–polymerase chain reaction (RT-PCR) testing of an appropriate respiratory tract sample by local clinical virology laboratories, as part of routine clinical care. Clinical laboratories followed nationally agreed and validated PCR protocols, and a panel of experts reviewed all results.
Influenza virus quantification
Nasopharyngeal secretions were collected into sterile universal sputum traps by suction catheterization. After 5 s of suctioning, any contents remaining within the catheter were flushed through with 5 ml normal saline. Samples were stored at –80 °C until analysis. Viral nucleic acids were extracted using the Qiagen MDx Biorobot automated extractor with the QIAamp Virus MDx Kit according to the manufacturer's instructions. qRT-PCR reactions were set up to a total volume of 15 μl using the Qiagen One-Step RT-PCR kit, using primers (influenza A matrix (M) or pH1N1 neuraminidase (NA)) as described previously41 on an ABI Prism 7500 SDS real time platform (Applied Biosystems). For viral-load quantitation, we first derived the crossing threshold (CT) value (at the inflexion spot of the sigmoid amplification curve to capture the point at which DNA amplification is exponential) performed in a batched assay as a relative expression of viral burden against each sample. Subsequently, this was measured against a standard curve of CT value to plaque-forming units per ml, generated by measurement of plaque-forming units when MDCK canine kidney cells were inoculated with a known amount of pH1N1.
Clinical data collection and assignment of scores for severity of illness
Clinical data were extracted from hospital case notes and recorded in the Flu-CIN data-collection tool42 by trained researchers. Prescription charts were examined to determine whether antibiotics were being administered before, during or after sampling time points.
Severity of illness was graded at T1 and T2 according to the following criteria: 1, no substantial respiratory compromise, with blood oxygen saturation of > 93% while the patient was breathing room air; 2, oxygen saturation of ≤93% while the patient was breathing room air, justifying or requiring supplemental oxygen by face mask or nasal cannulae (with or without continuous positive airway pressure support or non-invasive mechanical ventilation); 3, respiratory compromise requiring invasive mechanical ventilation with or without ECMO (extracorporeal membrane oxygenation). All clinical data underwent extensive validation and quality checking by independent data collection staff.
Detection of bacteria
Nasopharyngeal aspirates and swabs collected at T1 underwent microscopy and culture for bacteria. Additionally, multiplex PCR was performed to detect the following common respiratory bacteria in these samples: Staphylococcus aureus, Chlamydia pneumoniae, Haemophilus influenzae, Streptococcus pneumoniae, Pneumocystis pneumoniae, Legionella species, Klebsiella pneumoniae, Salmonella species, Moraxella catarrhalis, Mycoplasma pneumoniae and Bordetella pertussis. Throat swab samples obtained at T1 also underwent culture and microscopy. Where available, urine samples collected between T1 and T2 underwent pneumococcal antigen testing (BinaxNow, Alere). Clinical microbiology data were obtained from hospital laboratory databases, including results of blood cultures (when obtained 48 h before and after T1) and urinary pneumococcal antigen results (for patients who did not have a researcher-requested urinary antigen sample). An independent microbiologist assessed the significance and validity of positive blood-culture results, in an attempt to exclude cases of pseudobacteremia caused by commensal contamination.
Soluble immunological mediators
Serum, nasopharyngeal aspirate (NPA) and nasal-absorption fluid were collected at recruitment (T1) from participants with confirmed influenza and from adult healthy control subjects. Clotted blood was centrifuged at 1,000 g at 4 °C, and aliquots of serum supernatant were stored at –80 °C. Each NPA was collected using a 10 F Argyle suction catheter, inserted to reach the posterior nasopharyngeal wall; moderate suction was applied while the catheter was withdrawn over 5 s. The catheter was flushed through with 5 ml of sterile normal saline, and the total contents were collected in a universal container. Aliquots of NPA were stored at –80 °C. Nasal-absorption fluid was collected from the lateral wall of the nasal cavity using a synthetic absorptive matrix (SAM) strips (Leukosorb, Pall) and was stored at –80 °C until analysis. On the day of analysis, 500 μl Milliplex assay buffer (Millipore) was added to each thawed SAM strip before being placed in a Costar Spin-X centrifuge filter of pore size 0.22 μm held within an Eppendorf tube. Samples were centrifuged at 16,000 g for 5 min at 4 °C, and eluates were kept on ice.
IL-1β, IL-6 and CXCL8 were quantified in each sample type using a ten-plex inflammatory soluble immune mediator electrochemiluminescence assay analyzed on an MSD SECTOR instrument (Mesoscale Discovery). For each mediator, a coefficient variation cut-off of 10% was used to set the lower limit of detection. Sample results below the GM-LLOD (geometric mean lower limit of detection) were assigned half the value of the respective GM-LLOD.
Blood procalcitonin assay
Procalcitonin (PCT) in plasma or serum (collected at T1 and T2) was quantified using the Elecsys BRAHMS PCT assay on a calibrated Cobas e602 platform. Samples with a PCT value at the upper limit of detection (ULOD) were arbitrarily assigned a value of 100 ng/ml (the ULOD). Results may be interpreted as follows: < 0.5 ng/ml, low probability of significant bacterial infection; 0.5–2.0 ng/ml, medium probability of significant bacterial infection; > 2.0 ng/ml, high probability of significant bacterial infection.
16S rRNA gene bacterial load measurement
The gene encoding 16 S rRNA was targeted with 0.3 µl each of 10 µM universal primers 520 F 5′-AYT GGG YDT AAA GNG and 802 R 5′-TAC NVG GGT ATC TAA TCC added to 7.5 µl of SYBR Fast qPCR Kit Master Mix (KapaBio) and 5 µl of a 1:5 dilution of sample DNA extract and 1.9 µl of PCR Clean water (Mobio). Reactions were prepared in triplicate, and thermal cycling carried out on a VIIA-7 Real-Time PCR System. Thermal-cycling conditions were 90 °C for 3 min, then 40 cycles of 95 °C for 20 s, 50 °C for 30 s, 72 °C for 30 s, with default melt conditions. A standard curve for a cloned (TOPO TA, Invitrogen) gene encoding full-length Vibrio natriegens DSMZ 749 16 S rRNA was included in order to be able to calculate an absolute abundance from CT values together with no template controls. The resulting copy number of 16 S rRNA (bacterial load) was log-transformed before being used analytically.
Microarray gene-expression profiling
At each time point, 3 ml of whole blood was collected into each of two Tempus tubes (Applied Biosystems/Ambion) by trained research staff following a standard phlebotomy protocol. Blood was vigorously mixed immediately following collection and was stored at –80 °C before RNA extraction. For each patient, the contents of one tube were used for analysis, and the other tube was retained in case of assay failure. RNA was isolated using 1.5 ml whole blood and the MagMAX-96 Blood RNA Isolation Kit (Applied Biosystems/Ambion), as per the manufacturer's instructions. 250 μg of isolated total RNA was globin-reduced using the GLOBINclear 96-well format kit (Applied Biosystems/Ambion) according to the manufacturer’s instructions. Total and globin-reduced RNA integrity was assessed using an Agilent 2100 Bioanalyzer (Agilent Technologies). RNA yield was assessed using a NanoDrop8000 spectrophotometer (NanoDrop Products, Thermo Fisher Scientific). High-quality ( > 6.5 RIN) whole blood RNA was successfully obtained and processed by microarray in all cases. Biotinylated, amplified antisense complementary RNA (cRNA) targets were prepared from 200–250 ng of globin-reduced RNA using the Illumina CustomPrep RNA amplification kit (Applied Biosystems/Ambion). For each sample, 750 ng of labeled cRNA was hybridized overnight to Illumina Human HT12 V4 BeadChip arrays (Illumina), which contained greater than 47,000 probes. The arrays were washed, blocked, stained and scanned on an Illumina iScan, as per the manufacturer's instructions. GenomeStudio (Illumina) was used to perform quality control and generate signal intensity values.
Microarray data processing
Raw microarray data were processed using GeneSpring GX version 12.5 (Agilent Technologies). Following background subtraction, each probe was attributed a flag to denote its signal-intensity-detection P value. Filtering on flags removed probe sets that did not result in a ‘present’ call in at least 10% of the samples, where the ‘present’ lower cut-off was 0.99. Signal values were then set to a threshold level of 10, were log2-transformed and were per-chip normalized using a 75th percentile-shift algorithm. Each gene was normalized by dividing each mRNA transcript by the median intensity of all samples. Statistical analysis was performed after these steps had been performed.
Microarray data analysis
Transcripts significantly detected from background hybridization were filtered for low expression in GeneSpring GX 12.5, whereby the only transcripts retained were those with a change of at least twofold from the median normalized intensity value in at least 10% of all samples. Principal-component analysis of all transcripts significantly above background in at least 10% of samples (18,974 transcripts) was performed using R 3.3.2 (R Development Core Team). To derive the 1,255 transcript list, non-parametric statistical filters were applied (P < 0.01 (Mann-Whitney unpaired test with Bonferroni family-wise error rate (FWER) multiple-testing correction)), followed by filtering by change (fold values) between groups (transcripts were retained with a change greater than twofold between any two groups). For severity analysis, 231 normalized intensity value transcripts were obtained by filtering for low expression and then applying statistic filters (P < 0.01 (Kruskal-Wallis test with Bonferroni FWER)), followed by filtering by change (fold values) between groups (transcripts were retained with a change of greater than twofold between patients with illness of severity 3 and those with illness of severity 1 and 2). All heat maps were generated in GeneSpring GX 12.5 (semi-supervised analysis, clustered by Pearson's un- centered method with average linkage rule).
Comparison Ingenuity Pathway Analysis (IPA) (Ingenuity Systems) was used to determine the most significant canonical pathways for upregulated and downregulated transcripts (P < 0.05 (Fisher's exact test)). Additionally, IPA was used to generate the graphed presentation of selected canonical pathways and network diagrams. For the 231-transcript list, significantly activated biofunctions (z-score > 2) and significantly repressed biofunctions (z score < 2) were identified in IPA and are presented in gene-network diagrams. GO Term analysis (Gene Ontology Consortium) integrated with GeneSpring GX12.5 was used to identify biological processes, according to GO annotations43.
The molecular distance to health (MDTH) and molecular scores were calculated using methods described previously24 and were applied to different signatures. Transcriptional modular analysis was applied as described previously23. In brief, raw expression levels of all transcripts significantly above or below background were compared between each sample and all the controls present in a given dataset. The percentage of differentially expressed genes in each module is represented by the color intensity, with red indicating over-expression and blue indicating under-expression. Statistical testing was performed using Student’s t-test (P < 0.05). The mean percentage of significant genes and the mean change in expression of these genes (fold values) compared to the controls in specific modules are presented in graphical form (P < 0.00001 (unpaired t-test)). MDTH and modular analysis were calculated in Microsoft Excel 2010 (Microsoft). GraphPad Prism V5 for Windows (GraphPad Software) and R 3.3.2 (R Development Core Team) were used to generate graphs and perform additional statistical analyses.
Reporting summary
Further information on experimental design is available in the Nature Research Reporting Summary linked to this article.
Data availability statement
The raw and normalized microarray data that support the findings of this study have been deposited in GEO with the accession code GSE111368.
Change history
07 February 2019
In the version of this article initially published, a source of funding was not included in the Acknowledgements section. That section should include the following: P.J.M.O. was supported by EU FP7 PREPARE project 602525. The error has been corrected in the HTML and PDF version of the article.
References
Stöhr, K. Preventing and treating influenza. Br. Med. J. 326, 1223–1224 (2003).
Hayward, A. C. et al. Comparative community burden and severity of seasonal and pandemic influenza: results of the Flu Watch cohort study. Lancet Respir. Med. 2, 445–454 (2014).
Dawood, F. S. et al. Estimated global mortality associated with the first 12 months of 2009 pandemic influenza A H1N1 virus circulation: a modelling study. Lancet Infect. Dis. 12, 687–695 (2012).
Bautista, E. et al. Clinical aspects of pandemic 2009 influenza A (H1N1) virus infection. N. Engl. J. Med. 362, 1708–1719 (2010).
Hui, D. S., Lee, N. & Chan, P. K. Clinical management of pandemic 2009 influenza A(H1N1) infection. Chest 137, 916–925 (2010).
Peiris, J. S., Cheung, C. Y., Leung, C. Y. & Nicholls, J. M. Innate immune responses to influenza A H5N1: friend or foe? Trends Immunol. 30, 574–584 (2009).
Tisoncik, J. R. et al. Into the eye of the cytokine storm. Microbiol. Mol. Biol. Rev. 76, 16–32 (2012).
de Jong, M. D. et al. Fatal outcome of human influenza A (H5N1) is associated with high viral load and hypercytokinemia. Nat. Med. 12, 1203–1207 (2006).
Ioannidis, I. et al. Plasticity and virus specificity of the airway epithelial cell immune response during respiratory virus infection. J. Virol. 86, 5422–5436 (2012).
Ramilo, O. et al. Gene expression patterns in blood leukocytes discriminate patients with acute infections. Blood 109, 2066–2077 (2007).
Zaas, A. K. et al. Gene expression signatures diagnose influenza and other symptomatic respiratory viral infections in humans. Cell Host Microbe 6, 207–217 (2009).
Woods, C. W. et al. A host transcriptional signature for presymptomatic detection of infection in humans exposed to influenza H1N1 or H3N2. PLoS One 8, e52198 (2013).
Nakaya, H. I. et al. Systems biology of vaccination for seasonal influenza in humans. Nat. Immunol. 12, 786–795 (2011).
Li, S. et al. Molecular signatures of antibody responses derived from a systems biology study of five human vaccines. Nat. Immunol. 15, 195–204 (2014).
Obermoser, G. et al. Systems scale interactive exploration reveals quantitative and qualitative differences in response to influenza and pneumococcal vaccines. Immunity 38, 831–844 (2013).
Blankley, S. et al. The application of transcriptional blood signatures to enhance our understanding of the host response to infection: the example of tuberculosis. Phil. Trans. R. Soc. Lond. B 369, 20130427 (2014).
Pascual, V., Chaussabel, D. & Banchereau, J. A genomic approach to human autoimmune diseases. Annu. Rev. Immunol. 28, 535–571 (2010).
Everitt, A. R. et al. IFITM3 restricts the morbidity and mortality associated with influenza. Nature 484, 519–523 (2012).
Ruggiero, T. et al. A(H1N1)pdm09 hemagglutinin D222G and D222N variants are frequently harbored by patients requiring extracorporeal membrane oxygenation and advanced respiratory assistance for severe A(H1N1)pdm09 infection. Influenza Other Respir. Viruses 7, 1416–1426 (2013).
Rykkvin, R., Kilander, A., Dudman, S.G. & Hungnes, O. Within-patient emergence of the influenza A(H1N1)pdm09 HA1 222G variant and clear association with severe disease, Norway. Euro Surveill. 18, 20369 (2013).
Herfst, S. et al. Introduction of virulence markers in PB2 of pandemic swine-origin influenza virus does not result in enhanced virulence or transmission. J. Virol. 84, 3752–3758 (2010).
Elderfield, R. A. et al. Accumulation of human-adapting mutations during circulation of A(H1N1)pdm09 influenza virus in humans in the United Kingdom. J. Virol. 88, 13269–13283 (2014).
Chaussabel, D. et al. A modular analysis framework for blood genomics studies: application to systemic lupus erythematosus. Immunity 29, 150–164 (2008).
Pankla, R. et al. Genomic transcriptional profiling identifies a candidate blood biomarker signature for the diagnosis of septicemic melioidosis. Genome Biol. 10, R127 (2009).
Simon, L., Gauvin, F., Amre, D. K., Saint-Louis, P. & Lacroix, J. Serum procalcitonin and C-reactive protein levels as markers of bacterial infection: a systematic review and meta-analysis. Clin. Infect. Dis. 39, 206–217 (2004).
Wu, M. H. et al. Can procalcitonin tests aid in identifying bacterial infections associated with influenza pneumonia? A systematic review and meta-analysis. Influenza Other Respir. Viruses 7, 349–355 (2013).
Falsey, A. R. et al. Bacterial complications of respiratory tract viral illness: a comprehensive evaluation. J. Infect. Dis. 208, 432–441 (2013).
McNab, F., Mayer-Barber, K., Sher, A., Wack, A. & O’Garra, A. Type I interferons in infectious disease. Nat. Rev. Immunol. 15, 87–103 (2015).
Marshall, J. C. Iatrogenesis, inflammation and organ injury: insights from a murine model. Crit. Care 10, 173 (2006).
Openshaw, P. J. & Chiu, C. Protective and dysregulated T cell immunity in RSVinfection. Curr. Opin. Virol. 3, 468–474 (2013).
Davidson, S., Crotta, S., McCabe, T. M. & Wack, A. Pathogenic potential of interferon alphabeta in acute influenza infection. Nat. Commun. 5, 3864 (2014).
Gregory, D. J. & Kobzik, L. Influenza lung injury: mechanisms and therapeutic opportunities. Am. J. Physiol. Lung Cell. Mol. Physiol. 309, L1041–L1046 (2015).
Short, K. R., Kroeze, E. J. B. V., Fouchier, R. A. M. & Kuiken, T. Pathogenesis of influenza-induced acute respiratory distress syndrome. Lancet Infect. Dis. 14, 57–69 (2014).
Sandler, N. G. et al. Type I interferon responses in rhesus macaques prevent SIV infection and slow disease progression. Nature 511, 601–605 (2014).
Biron, C. A. Interferons alpha and beta as immune regulators-a new look. Immunity 14, 661–664 (2001).
Narasaraju, T. et al. Excessive neutrophils and neutrophil extracellular traps contribute to acute lung injury of influenza pneumonitis. Am. J. Pathol. 179, 199–210 (2011).
Lukens, M. V. et al. A systemic neutrophil response precedes robust CD8+ T-cell activation during natural respiratory syncytial virus infection in infants. J. Virol. 84, 2374–2383 (2010).
Tate, M. D., Brooks, A. G., Reading, P. C. & Mintern, J. D. Neutrophils sustain effective CD8+ T-cell responses in the respiratory tract following influenza infection. Immunol. Cell Biol. 90, 197–205 (2012).
Lim, K. et al. Neutrophil trails guide influenza-specific CD8+ T cells in the airways. Science 349, aaa4352 (2015).
Mayer-Barber, K. D. et al. Host-directed therapy of tuberculosis based on interleukin-1 and type I interferon crosstalk. Nature 511, 99–103 (2014).
Carr, M. J. et al. Development of a real-time RT-PCR for the detection of swine-lineage influenza A (H1N1) virus infections. J. Clin. Virol. 45, 196–199 (2009).
Nguyen-Van-Tam, J. S. et al. Risk factors for hospitalisation and poor outcome with pandemic A/H1N1 influenza: United Kingdom first wave (May-September 2009). Thorax 65, 645–651 (2010).
Ashburner, M. et al. The Gene Ontology Consortium. Gene ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Acknowledgements
MOSAIC Study was supported by a joint award from the Wellcome Trust and the Medical Research Council (090382/Z/09/Z). We gratefully acknowledge the support of the MOSAIC administrative team (M. Cross, L.-A. Cumming, M. Minns, T. Ford, B. Cerutti, D. Gardner and Z. Williams) and the generosity of our patients and their families, healthy volunteers, and staff at participating National Health Service (NHS) hospitals (Alder Hey Children's Hospital; Brighton & Sussex University Hospitals NHS Trust; Central Manchester University Hospitals NHS Foundation Trust; Chelsea and Westminster Hospital NHS Foundation Trust; Imperial College Healthcare NHS Trust; Liverpool Women's NHS Foundation Trust; Royal Liverpool and Broadgreen University Hospitals NHS Trust; Royal Brompton and Harefield NHS Foundation Trust; University Hospitals Coventry and Warwickshire NHS Trust). In particular, we thank K. Alshafi, S. Ashton, E. Bailey, A. Bermingham, M. Berry, C. Bloom, A. Booth, E. Brannigan, S. Bremang, J. Clark, M. Cross, L. A. Cumming, S. Dyas, J. England-Smith, J. Enstone, D. Ferreira, N. Goddard, A. Godlee, S. Gormley, M. Guiver, M. O. Hassan-Ibrahim, H. Hill, P. Holloway, K. Hoschler, G. Houghton, F. Hughes, R. R. Israel, A. Jepson, K. D. Jones, W. P. Kelleher, M. Kidd, K. Knox, A. Lackenby, G. Lloyd, H. Longworth, S. Mookerjee, S. Mt-Isa, D. Muir, A. Paras, V. Pascual, L. Rae, S. Rodenhurst, F. Rozakeas, E. Scott, E. Sergi, N. Shah, V. Sutton, J. Vernazza, A.W. Walker, C. Wenden, T. Wotherspoon, A. D. Wright and F. Wurie. We also thank E. Anguiano and members of the Genomic Core, BIIR, Dallas, for assistance with the microarray analysis; and M. Berry for guidance. We especially thank the MOSAIC Data Team (L. Drumright, L. Garcia-Alvarez, J. Lieber, S. Mookerjee and B. Pamba) for assistance in collating and validating clinical data; R. Smyth for careful review of the manuscript; and K. Strimmer for statistical advice in revisions of the manuscript. F. Rozakeas helped with recruiting samples from all the healthy control subjects at NIMR (Mill Hill). The MOSAIC consortium (ClinicalTrials.gov identifier NCT00965354) was supported by UK National Institute for Health Research (NIHR) Comprehensive Local Research Networks (CLRNs), the Biomedical Research Centre (NIHR Imperial BRC) and Unit (NIHR Liverpool BRU), the Health Protection Research Unit in Respiratory Infections in partnership with Public Health England (PHE) at Imperial College London (NIHR HPRU RI), the Health Protection Agency (latterly PHE) Microbiology Services, Colindale and the staff of the Roslin Institute, Edinburgh, Scotland. A.O.G. and C.G. were supported by the Medical Research Council, United Kingdom (U117565642), The Francis Crick Institute, London (AOG10126, which receives its core funding from Cancer Research UK (FC001126)), the UK Medical Research Council (FC001126), the Wellcome Trust (FC001126) and the UK Medical Research Council (MR/U117565642/1). S.B. was in part jointly funded by the UK Medical Research Council (MRC) as above and the UK Department for International Development (DFID) under the MRC/DFID Concordat agreement (MR/J010723/1). C.B. was funded by an MRC CRTF. P.J.M.O. was supported by EU FP7 PREPARE project 602525. The views expressed are those of the authors and not necessarily those of the NHS, NIHR, Public Health England or the Department of Health (UK). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
MOSAIC investigators:
Maximillian S. Habibi, Sebastian L. Johnston, Trevor T. Hansel, Mike Levin, Ryan S. Thwaites, John O. Warner, William O. Cookson, Brian G. Gazzard, Alan Hay, John McCauley, Paul Aylin, Deborah Ashby, Wendy S. Barclay, R. A. Elderfield, Simon Nadel, Jethro A. Herberg, Lydia N. Drumright, Laura Garcia-Alvarez, Alison H. Holmes, Onn M. Kon, Stephen J. Aston, Stephen B. Gordon, Tracy Hussell, Catherine Thompson, Maria C. Zambon, Kenneth J. Baillie, David A. Hume, Peter Simmonds, Andrew Hayward, Rosalind L. Smyth, Paul S. McNamara, Malcolm G. Semple, Jonathan S. Nguyen-Van-Tam, Ling-Pei Ho, Andrew J. McMichael, Paul Kellam, Walt E Adamson, William F Carman and Mark J. Griffiths
Author information
Authors and Affiliations
Consortia
Contributions
J.D., A.O.G. and P.J.M.O. conceived of and designed the study, with input from D.C. and J.B.; J.D., S.B., L.T.H. and C.I.B. analyzed the microarray data, with supervision by A.O.G. and P.J.M.O. and input from J.B. and D.C.; M.C., P.L.J. and M.F.M. performed and analyzed the microbiome experiments; C.M.G. performed the microarray experiments; J.D., S.J.B. and P.J.M.O. developed clinical protocols, recruited subjects and collated clinical data; and J.D., S.B., L.T.H., A.O.G. and P.J.M.O. wrote and revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Interferon signaling pathway.
IPA canonical pathway for interferon signaling, identified as one of the top five canonical pathways for upregulated transcripts shown in Figure 1d (2010/11 cohort). Red shading represents up-regulated genes, blue represents down-regulated genes.
Supplementary Figure 2 Validation of transcriptional signatures in an independent cohort.
a) 2009/2010 cohort clustered on individuals and transcripts (Pearson’s uncentered with averaged linkage) using 1255 transcript list (from Figure 1). (b) 2009/2010 cohort clustered on individuals and transcripts (Pearson’s uncentered with averaged linkage) using 25 transcript list (from Figure 1). (c) 2009/2010 cohort clustered on individuals and transcripts (Pearson’s uncentered with averaged linkage) using 231 transcript list of severity (from Figure 2, transcripts retained if >2FC between severity 3 and severity 1&2). GO Terms analysis of 3 major branches of the transcripts dendrogram was undertaken and listed next to the heat-map. (d) Using 51 and 112 transcripts lists (from Figure 3) ‘viral response’ and ‘bacterial response’ molecular scores were calculated and plotted for each influenza patient (relative to healthy controls). Cases were coded according to severity of illness, indicated by the colour of the respective dots (severity 1, black; severity 2, blue; severity 3, red).
Supplementary Figure 3 Change of ‘viral’ and ‘bacterial’ molecular scores over time and association with influenza viral load.
(a) ‘Viral’ molecular scores plotted for 59 influenza patients (2010/11 cohort) who provided T1 and T2 samples, plotted against respective day of illness at time of sampling. (b) Change in ‘viral’ molecular score between first (T1) and precise second time point (48 hours after T1) in 41 patients with appropriate samples available (P= 0.0002, Mann-Whitney test, two-tailed). (c) ‘Bacterial’ molecular score plotted for 59 influenza patients who had both a T1 and a T2 sample, shown plotted against respective day of illness. (d) Change in ‘bacterial’ molecular score between T1 and precise T2 (48h post T1), in 41 patients with appropriate samples available (NS, Mann-Whitney test, two-tailed). (e) Influenza viral load estimation (pfu/ml) in nasopharyngeal samples obtained at T1 (n=42) and T2 (n=40). Bars show the median and interquartile range. Zero values were reassigned a value of 0.001 for display purposes. Mann Whitney test (two-tailed); ** P=0.0094. (f) Relationship between influenza viral load (pfu/ml) at T1 or T2 and the simultaneous ‘viral’ molecular score on whole blood (n=82).
Supplementary Figure 4 Administration of antibiotics does not affect ‘bacterial’ or ‘viral’ molecular scores.
(a) Influenza patients (2010/11 cohort) presenting within the first 14 days of illness grouped by administration of any antibiotic (n=35) or no administration of antibiotics (n=35) in the 24 hours prior to T1 sampling. There was no difference (NS, Mann-Whitney test, two tailed) in either bacterial or viral molecular scores between the two groups. Bars show the median and interquartile range. (b) Prescription of antibiotics after T1 did not significantly influence ‘bacterial’ molecular score (P=0.9616, Kruskal Wallis test). Fifty-nine influenza patients who had both T1 and T2 samples were grouped by those who did not receive antibiotics (n=7), those whose antibiotics were stopped at T1 (n=1), those who had antibiotics prescribed after T1 but before T2 (n=24), and those who were receiving antibiotics at both T1 and T2 (N=27). Bars show median with interquartile range. (c) Total 16S rRNA copies at T1 in throat swabs and NP aspirate in patients adjudicated to be with or without bacterial co-infection. Those with confirmed bacterial infections (Bac +) had greater levels of total 16S rRNA copies in NP aspirate than those deemed to be without co-infection (Bac -) (Mann- Whitney test, P = 0.036). Throat swab Bac -, n=44; throat swab bac +, n=53; NP aspirate Bac -, n=17; NP aspirate Bac +, n=41.
Supplementary Figure 5 Correlation of serum cytokines and bacterial load in nasophaynx with ‘viral’ and ‘bacterial’ molecular distance to health.
(a) Levels of IL-17 in the serum of healthy controls (HC, n=36) and influenza infected patients (severity 1-3; n=59, n=43, and n=31, respectively). (b) Concentration of IL-17 in broncoalveolar lavage (BAL) of HC (n=11) and from influenza patients’ BAL (n=8), NPA (n=8), nasadsoprtion fluid (SAM; n=8) and serum (n=8). (c) Correlation of levels of IL-17 in serum (n=165) with the bacterial MDTH (Spearman R =0.39, P<0.001). (d) Correlation of levels of TNF-α in serum (n=165) with bacterial MDTH (Spearman R = 0.4, P<0.01). (e) Total 16S rRNA gene copies in NP aspirate samples (n=58) are inversely correlated with the viral MDTH (Spearman R = -0.28, P value < 0.05). (f) Total 16S rRNA gene copies in NP aspirate samples (n=58) are positively correlated with the bacterial MDTH (Spearman R = 0.37, P value < 0.05).
Supplementary information
Supplementary Figures and Tables
Supplementary Figures 1-5; Supplementary Tables 1 and 2
Supplementary Dataset
Excel spreadsheet providing clinical data corresponding to raw and normalized microarray data deposited in GEO, Data Accession Code GSE111368.
Rights and permissions
About this article
Cite this article
Dunning, J., Blankley, S., Hoang, L.T. et al. Progression of whole-blood transcriptional signatures from interferon-induced to neutrophil-associated patterns in severe influenza. Nat Immunol 19, 625–635 (2018). https://doi.org/10.1038/s41590-018-0111-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-018-0111-5
This article is cited by
-
Bioinformatics analysis identifies a key gene HLA_DPA1 in severe influenza-associated immune infiltration
BMC Genomics (2024)
-
Obesity dysregulates the pulmonary antiviral immune response
Nature Communications (2023)
-
Neutrophils and emergency granulopoiesis drive immune suppression and an extreme response endotype during sepsis
Nature Immunology (2023)
-
Co-expression network analysis identifies potential candidate hub genes in severe influenza patients needing invasive mechanical ventilation
BMC Genomics (2022)
-
Early IFN-β administration protects cigarette smoke exposed mice against lethal influenza virus infection without increasing lung inflammation
Scientific Reports (2022)