Abstract
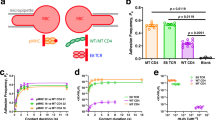
T cell antigen recognition requires T cell antigen receptors (TCRs) engaging MHC-embedded antigenic peptides (pMHCs) within the contact region of a T cell with its conjugated antigen-presenting cell. Despite micromolar TCR:pMHC affinities, T cells respond to even a single antigenic pMHC, and higher-order TCRs have been postulated to maintain high antigen sensitivity and trigger signaling. We interrogated the stoichiometry of TCRs and their associated CD3 subunits on the surface of living T cells through single-molecule brightness and single-molecule coincidence analysis, photon-antibunching-based fluorescence correlation spectroscopy and Förster resonance energy transfer measurements. We found exclusively monomeric TCR–CD3 complexes driving the recognition of antigenic pMHCs, which underscores the exceptional capacity of single TCR–CD3 complexes to elicit robust intracellular signaling.
This is a preview of subscription content, access via your institution
Access options





Similar content being viewed by others
References
Bromley, S. K. et al. The immunological synapse. Annu. Rev. Immunol. 19, 375–396 (2001).
Davis, M. M. et al. Ligand recognition by αβ T cell receptors. Annu. Rev. Immunol. 16, 523–544 (1998).
Irvine, D. J., Purbhoo, M. A., Krogsgaard, M. & Davis, M. M. Direct observation of ligand recognition by T cells. Nature 419, 845–849 (2002).
Purbhoo, M. A., Irvine, D. J., Huppa, J. B. & Davis, M. M. T cell killing does not require the formation of a stable mature immunological synapse. Nat. Immunol. 5, 524–530 (2004).
Huppa, J. B. & Davis, M. M. The interdisciplinary science of T-cell recognition. Adv. Immunol. 119, 1–50 (2013).
Taylor, M. J., Husain, K., Gartner, Z. J., Mayor, S. & Vale, R. D. A DNA-based T cell receptor reveals a role for receptor clustering in ligand discrimination. Cell 169, 108–119 (2017).
Call, M. E., Pyrdol, J., Wiedmann, M. & Wucherpfennig, K. W. The organizing principle in the formation of the T cell receptor–CD3 complex. Cell 111, 967–979 (2002).
Huppa, J. B. & Ploegh, H. L. In vitro translation and assembly of a complete T cell receptor–CD3 complex. J. Exp. Med. 186, 393–403 (1997).
Huppa, J. B. & Ploegh, H. L. The α chain of the T cell antigen receptor is degraded in the cytosol. Immunity 7, 113–122 (1997).
Weissman, A. M., Samelson, L. E. & Klausner, R. D. A new subunit of the human T-cell antigen receptor complex. Nature 324, 480–482 (1986).
Exley, M., Wileman, T., Mueller, B. & Terhorst, C. Evidence for multivalent structure of T-cell antigen receptor complex. Mol. Immunol. 32, 829–839 (1995).
Schamel, W. W. et al. Coexistence of multivalent and monovalent TCRs explains high sensitivity and wide range of response. J. Exp. Med. 202, 493–503 (2005).
Kumar, R. et al. Increased sensitivity of antigen-experienced T cells through the enrichment of oligomeric T cell receptor complexes. Immunity 35, 375–387 (2011).
James, J. R. et al. Single-molecule level analysis of the subunit composition of the T cell receptor on live T cells. Proc. Natl. Acad. Sci. USA 104, 17662–17667 (2007).
Wilson, B. S., Pfeiffer, J. R., Surviladze, Z., Gaudet, E. A. & Oliver, J. M. High resolution mapping of mast cell membranes reveals primary and secondary domains of FcεRI and LAT. J. Cell Biol. 154, 645–658 (2001).
Lillemeier, B. F. et al. TCR and Lat are expressed on separate protein islands on T cell membranes and concatenate during activation. Nat. Immunol. 11, 90–96 (2010).
Tang, Q. et al. The Src family kinase Fyn mediates signals induced by TCR antagonists. J. Immunol. 168, 4480–4487 (2002).
Cochran, J. R., Cameron, T. O. & Stern, L. J. The relationship of MHC–peptide binding and T cell activation probed using chemically defined MHC class II oligomers. Immunity 12, 241–250 (2000).
Krogsgaard, M. et al. Agonist/endogenous peptide–MHC heterodimers drive T cell activation and sensitivity. Nature 434, 238–243 (2005).
Brown, J. H. et al. Three-dimensional structure of the human class II histocompatibility antigen HLA-DR1. Nature 364, 33–39 (1993).
Stern, L. J. et al. Crystal structure of the human class II MHC protein HLA-DR1 complexed with an influenza virus peptide. Nature 368, 215–221 (1994).
Fremont, D. H., Hendrickson, W. A., Marrack, P. & Kappler, J. Structures of an MHC class II molecule with covalently bound single peptides. Science 272, 1001–1004 (1996).
Reich, Z. et al. Ligand-specific oligomerization of T-cell receptor molecules. Nature 387, 617–620 (1997).
Baker, B. M. & Wiley, D. C. αβ T cell receptor ligand-specific oligomerization revisited. Immunity 14, 681–692 (2001).
Moertelmaier, M., Brameshuber, M., Linimeier, M., Schutz, G. J. & Stockinger, H. Thinning out clusters while conserving stoichiometry of labeling. Appl. Phys. Lett. 87, 263903 (2005).
Brameshuber, M. et al. Imaging of mobile long-lived nanoplatforms in the live cell plasma membrane. J. Biol. Chem. 285, 41765–41771 (2010).
Brameshuber, M. & Schütz, G. J. Detection and quantification of biomolecular association in living cells using single-molecule microscopy. Methods Enzymol. 505, 159–186 (2012).
Huppa, J. B. et al. TCR–peptide–MHC interactions in situ show accelerated kinetics and increased affinity. Nature 463, 963–967 (2010).
Ruprecht, V., Brameshuber, M. & Schutz, G. J. Two-color single molecule tracking combined with photobleaching for the detection of rare molecular interactions in fluid biomembranes. Soft Matter 6, 568–581 (2010).
Xie, J. et al. Photocrosslinkable pMHC monomers stain T cells specifically and cause ligand-bound TCRs to be ‘preferentially’ transported to the cSMAC. Nat. Immunol. 13, 674–680 (2012).
Kimble, H. J., Dagenais, M. & Mandel, L. Photon antibunching in resonance fluorescence. Phys. Rev. Lett. 39, 691–695 (1977).
Sýkora, J. et al. Exploring fluorescence antibunching in solution to determine the stoichiometry of molecular complexes. Anal. Chem. 79, 4040–4049 (2007).
Ta, H., Kiel, A., Wahl, M. & Herten, D. P. Experimental approach to extend the range for counting fluorescent molecules based on photon-antibunching. Phys. Chem. Chem. Phys. 12, 10295–10300 (2010).
Varma, R., Campi, G., Yokosuka, T., Saito, T. & Dustin, M. L. T cell receptor–proximal signals are sustained in peripheral microclusters and terminated in the central supramolecular activation cluster. Immunity 25, 117–127 (2006).
Demetriou, M., Granovsky, M., Quaggin, S. & Dennis, J. W. Negative regulation of T-cell activation and autoimmunity by Mgat5 N-glycosylation. Nature 409, 733–739 (2001).
Valitutti, S., Müller, S., Cella, M., Padovan, E. & Lanzavecchia, A. Serial triggering of many T-cell receptors by a few peptide–MHC complexes. Nature 375, 148–151 (1995).
Davis, S. J. & van der Merwe, P. A. The structure and ligand interactions of CD2: implications for T-cell function. Immunol. Today 17, 177–187 (1996).
Chang, V. T. et al. Initiation of T cell signaling by CD45 segregation at ‘close contacts’. Nat. Immunol. 17, 574–582 (2016).
Beddoe, T. et al. Antigen ligation triggers a conformational change within the constant domain of the αβ T cell receptor. Immunity 30, 777–788 (2009).
Gil, D., Schamel, W. W., Montoya, M., Sánchez-Madrid, F. & Alarcón, B. Recruitment of Nck by CD3ε reveals a ligand-induced conformational change essential for T cell receptor signaling and synapse formation. Cell 109, 901–912 (2002).
Lee, M. S. et al. A mechanical switch couples T cell receptor triggering to the cytoplasmic juxtamembrane regions of CD3ζζ. Immunity 43, 227–239 (2015).
Wang, F., Beck-García, K., Zorzin, C., Schamel, W. W. & Davis, M. M. Inhibition of T cell receptor signaling by cholesterol sulfate, a naturally occurring derivative of membrane cholesterol. Nat. Immunol. 17, 844–850 (2016).
Swamy, M. et al. A cholesterol-based allostery model of T cell receptor phosphorylation. Immunity 44, 1091–1101 (2016).
Kim, S. T. et al. The αβ T cell receptor is an anisotropic mechanosensor. J. Biol. Chem. 284, 31028–31037 (2009).
Liu, B., Chen, W., Evavold, B. D. & Zhu, C. Accumulation of dynamic catch bonds between TCR and agonist peptide–MHC triggers T cell signaling. Cell 157, 357–368 (2014).
Katz, Z. B., Novotná, L., Blount, A. & Lillemeier, B. F. A cycle of Zap70 kinase activation and release from the TCR amplifies and disperses antigenic stimuli. Nat. Immunol. 18, 86–95 (2017).
Huppa, J. B., Gleimer, M., Sumen, C. & Davis, M. M. Continuous T cell receptor signaling required for synapse maintenance and full effector potential. Nat. Immunol. 4, 749–755 (2003).
Huse, M. et al. Spatial and temporal dynamics of T cell receptor signaling with a photoactivatable agonist. Immunity 27, 76–88 (2007).
Cai, E. et al. Visualizing dynamic microvillar search and stabilization during ligand detection by T cells. Science 356, eaal3118 (2017).
Altman, J. D., Reay, P. A. & Davis, M. M. Formation of functional peptide complexes of class II major histocompatibility complex proteins from subunits produced in Escherichia coli. Proc. Natl. Acad. Sci. USA 90, 10330–10334 (1993).
Axmann, M., Schütz, G. J. & Huppa, J. B. Single molecule fluorescence microscopy on planar supported bilayers. J. Vis. Exp. (105), e53158 (2015)..
Howarth, M. et al. A monovalent streptavidin with a single femtomolar biotin binding site. Nat. Methods 3, 267–273 (2006).
Fairhead, M., Krndija, D., Lowe, E. D. & Howarth, M. Plug-and-play pairing via defined divalent streptavidins. J. Mol. Biol. 426, 199–214 (2014).
Harms, G. S., Sonnleitner, M., Schütz, G. J., Gruber, H. J. & Schmidt, T. Single-molecule anisotropy imaging. Biophys. J. 77, 2864–2870 (1999).
Lakowicz, J. R. Principles of Fluorescence Spectroscopy (Springer, Boston, 2006)..
Schmidt, T., Schütz, G. J., Gruber, H. J. & Schindler, H. Local stoichiometries determined by counting individual molecules. Anal. Chem. 68, 4397–4401 (1996).
Wieser, S. & Schütz, G. J. Tracking single molecules in the live cell plasma membrane—Do’s and Don’t’s. Methods 46, 131–140 (2008).
Schütz, G. J., Schindler, H. & Schmidt, T. Single-molecule microscopy on model membranes reveals anomalous diffusion. Biophys. J. 73, 1073–1080 (1997).
Acknowledgements
We would like to thank S. Hell (Max Planck Institute for Biophysical Chemistry) for providing imaging infrastructure and support (H.T.) and M. Focke Tejkl for offering help and infrastructure concerning peptide mass spectrometry. This work was supported by Austrian Science Fund (FWF) projects I953-B20 (M.B.), P 25775-B2 (K.A., J.B.H.), P275941-B28 (F.B.) and V538-B26 (E.S.), by Vienna Science and Technology Fund (WWTF) project LS13-030 (F.K., G.J.S.), by the Singapore National Medical Research Council under NMRC/CBRG/0064/2014 (N.R.J.G.) and by the Wellcome Trust, grant 098274/Z/12/Z (S.J.D.).
Author information
Authors and Affiliations
Contributions
M.B. and J.B.H. conceived the project and wrote the manuscript. M.B. performed or supervised most of the imaging experiments. J.B.H. performed the initial FRET-based experiments and contributed to early TOCCSL experiments. M.A. performed early TOCCSL experiments and provided ideas. B.K.R. performed TOCCSL and FRET-based experiments. F.K. and K.A. produced important imaging probes. F.K. and J.G. performed FRET-based experiments. E.S. built molecular models, analyzed anisotropy data and provided ideas. G.J.S. provided imaging infrastructure and important ideas. H.T. performed photon-antibunching experiments and provided imaging infrastructure. F.B., H.S., N.R.J.G. and S.J.D. contributed key reagents and important ideas.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Analysis of H57-scFv binding lifetime, TCR diffusion, anisotropy, TCR diffusion under lactose treatment, target binding of antibody probes and distribution of TCR diffusion constants.
a, The half-life of AF647-H57-scFv bound to surface-exposed TCRs was determined as follows: T cells were labeled under saturating conditions with AF647-H57-scFv and placed onto a lipid bilayer featuring the adhesion molecule ICAM-1. The background-subtracted mean brightness values of individual T cells were determined in TIR mode at various time points (black open circles). Acquired intensities were determined for three time intervals, 0–200 s (n = 11 cells), 800–1,050 s (n = 12 cells) and 1,750–2,000 s (n = 17 cells), and average values were calculated (red open square). The resulting curve was fitted to a single-exponential function and yielded a half-life t½, of 44 ± 9 min at room temperature. b, TCR diffusion constants were determined as follows: TCRs were labeled with AF647-H57-scFv and single-molecule tracking experiments were performed. Mean square displacements were determined and are plotted as a function of time lags. Assuming pure Brownian motion, a linear fit yielded a diffusion coefficient D of 0.037 ± 0.002 μm²/s (n = 21 cells). c, The immobile fraction of TCRs was determined as follows: single-molecule tracking displacements for various time lags between 20 and 140 ms were fitted by a biexponential distribution function (Methods, “TOCCSL analysis”), yielding the fraction of immobile/slowly diffusing TCRs (left; average fraction f = 0.36 ± 0.03), the mean square displacement (msd)–time lag plot of immobile/slowly diffusing TCRs (middle; D = 0.003 μm²/s) and the msd–time lag plot of fast diffusing TCRs (right; D = 0.047 μm²/s). The fraction of fast/mobile TCRs is given by 1 – f = 0.64 ± 0.03 (n = 21 cells). d, Anisotropy measurements: T cells were decorated with the indicated probes at saturating conditions for anisotropy measurements as described in the Methods. For each labeling condition, more than 11 cells were used for the analysis. Anisotropy of all four employed fluorescently labeled pMHCs was measured in bilayers decorated with the respective probe (n = 20 lipid bilayer regions measured per pMHC labeling variant). e, Lactose treatment. T cells were labeled under saturating conditions with AF555-H57-scFv and incubated in imaging buffer, imaging buffer containing 20 mM lactose or imaging buffer containing 20 mM sucrose for 20 min at 4 °C. T cells were then placed onto a lipid bilayer featuring the adhesion molecule ICAM-1. Depending on the carbohydrate pretreatment, FRAP experiments were conducted in the presence of imaging buffer (n = 21 cells), imaging buffer containing 2 mM lactose (n = 16 cells) or imaging buffer containing 2 mM sucrose (n = 16 cells). Mobile fractions for lactose- and sucrose-treated T cells were normalized with regard to the mobile fraction of untreated T cells. f, Target binding of employed probes. T cells were exposed to the probes shown at increasing concentrations and their mean fluorescence intensity (MFI) was determined by flow cytometry. MFIs were normalized to those obtained with the highest probe concentration at which probe saturation had occurred (circles). Data were fitted with an exponential function (solid line). Concentrations, at which half of the TCRs were labeled by the respective probes, yielded 1.83 μg/ml, 3.5 μg/ml and 27.3 μg/ml for H57-scFv, KT3-scFv and KJ25-Fab, respectively (n = 10,000 cells per data point). g, Distribution of diffusion coefficients. Single-molecule tracks from b were analyzed individually, and for every trajectory the diffusion coefficient was determined by mean square displacement analysis (n = 1,260 trajectories). Error bars, s.e.m. (with the exception of a, s.d.).
Supplementary Figure 2 Alternative recording mode and channel calibration for dual-color TOCCSL, TCR surface density distribution measured with KJ25-Fabs and single-molecule FRET events in dual-color TOCCSL measurements.
a, Alternative recording mode involving a dual-color TOCCSL image sequence using alternating red and green excitation (shown is a representative experiment). This imaging modality allows not only for dual-color-based colocalization analysis in the TOCCSL image (images i and ii) but may also provide direct evidence of molecular proximities via FRET upon exciting the FRET donor and imaging the FRET acceptor channel (image iii). The dashed area shows the position of the field stop. b–d, Channel calibration for dual-color TOCCSL experiments. b, Left, the positions of fluorescent multicolor beads detected in the red and green emission channels were overlaid without correction. Right, using this information to calculate the relative shift and stretch of one of the two color channels with respect to the other, the positions of the green color channel were corrected and overlaid with the positions of beads in the red channel. c, Histograms of the positional accuracies (PA) of bead positions in the respective channel are shown. The virtual distance of beads after correction (b, right) is shown to the right and yields an average value of about 20 nm. d, Shown is an example of the distribution of virtual distances for the dual-color TOCCSL experiment with H57-scFv and KJ25-Fab shown in Fig. 2. e, TCR surface densities as measured with KJ25-Fabs. An alternative epitope present on the 5C.C7 TCRβ chain, which is involved in pMHC binding, is probed with the use of KJ25-Fabs. T cells were labeled with AF488-KJ25-Fab and AF647-KJ25-Fab in a 1:1 ratio and placed on a lipid bilayer featuring ICAM-1. The surface densities were calculated for both probes as described in Fig. 1a, yielding an overall TCR surface density of 76 ± 16 molecules per µm². f, Dual-color TOCCSL analysis combined with single-molecule FRET analysis to score for molecular proximity. T cells were simultaneously decorated with AF647-H57-scFv and AF488-KJ25-Fab, and a dual-color colocalization-based TOCCSL experiment was performed. The images were recorded 10 s after the bleach pulse. Positions of diffraction-limited spots were determined for both color channels and are marked with red open circles in the red AF647-H57-scFv channel (top left, red excitation), green open circles in the AF488-KJ25-Fab channel (bottom left, green excitation) and yellow circles in the FRET image (top right, green excitation). Corrected positions of colocalized probes projected into the other color channels are shown as green, red and yellow crosses. For coincidence analysis, the region of interest (ROI) was chosen to ensure the presence of only detection-limited signals (dotted line). Shown is one colocalization (yellow arrow), which is further supported by the presence of a FRET signal (a representative experiment is shown). Scale bars, 4 μm.
Supplementary Figure 3 Autocorrelation functions measured for employed antibody probes in solution.
a–c, The left panel shows the measured autocorrelation functions (ACFs) (blue line) of AS635-H57-scFv (a), AS635-KJ25-Fab (b) and AS635-KT3-scFv (c) in solution at a concentration of 1 to 10 nM. The curves were fitted by a 3D diffusion function with one triplet state (red dashed line). In the right panel, ACFs are displayed at 0 s (blue), at the first period (100 ns for H57-scFv and 400 ns for KJ25-Fab and KT3-scFv) (green) and 2 s (red) lag time. Fitting (red and blue lines) yielded an average value of 1.22 ± 0.014 emitters per AS635-H57-scFv molecule (n = 10 measurements), 1.26 ± 0.01 emitters per AS635-KJ25-Fab (n = 12 measurements) and 1.13 ± 0.01 emitters per AS635-KT3-scFv (n = 11 measurements).
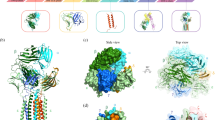
Supplementary Figure 4 FRET-based assessment of epitope proximities within putative dimeric TCR and pMHC configurations, and the DNA and amino acid sequences of the KT3-scFv and H57-scFv constructs employed.
a,b, Composite structure of the 5C.C7 αβ TCR decorated with H57-scFv in two dimer configurations as previously proposed47. The positions of the-FRET dyes are indicated by red and green stars. Putative dimerization motifs are shown in yellow and orange. FRET efficiencies as calculated for the AF555/AF647 (a) and AF568/AF647 (b) dye combinations with Förster radii of 5.1 nm and 8.2 nm, respectively, are indicated. c, If I-Ek-embedded peptides (yellow) served as FRET sensors for the detection of higher-order TCR structures, both I-Ek dimer configurations would give rise to substantial FRET yields as indicated. Scale bar, 5 nm. d,e, Annotated DNA (d) and amino acid (e) sequences of the KT3-scFv constructs designed for maleimide/cysteine-based site-specific dye conjugation at the indicated positions. KT3-scFv-C1 was employed in FRET measurements involving CD3ε–CD3ε and CD3ε–TCRβ, while KT3-scFv-C2 was used in TOCCSL and PA/FCS-based experiments (GenBank accessions MH045457, MH045458, MH045459). f,g, Annotated DNA (f) and amino acid (g) sequences of the H57-scFv constructs designed for site-specific conjugation with maleimide dyes or maleimide click adaptors. Positions for the introduction of cysteine residues are indicated. H57-scFv-C1 was employed in all experiments except those shown in Fig. 4b/third row, Fig. 4c and Supplementary Fig. 4f, which involved divalent-streptavidin-mediated cross-linking of TCR-bound H57-scFvs (GenBank accessions MH045460, MH045461, MH045462).
Supplementary Figure 5 FRET measured between TCR-bound H57-scFvs.
a, FRET values expected from randomized molecular encounters via Monte Carlo simulations are shown at indicated TCR densities (AF555:AF647 = 1:1; AF568:AF647 = 1:1) (n = 20 simulations per data point). b, FRET efficiencies measured under non-activating (NA) conditions. T cells were labeled with the indicated FRET donor:FRET acceptor ratios (AF555-H57-scFv:AF647-H57-scFv) and placed onto lipid bilayers featuring ICAM-1 and B7-1. Donor recovery after acceptor photobleaching experiments was carried out to determine the ensemble FRET yield. To account for photobleaching, experiments were conducted with green excitation only, and the laser power was adjusted to keep photobleaching below 2% (n = 11 cells). FRET could not be detected (n = 13 cells), even in the more sensitive experiment involving a 1:3 FRET donor:FRET acceptor labeling ratio (n = 10 cells). c, Increasing the sensitivity of the FRET-based assay toward longer distances. FRET efficiencies were measured for the AF568:AF647 dye pair exhibiting a larger Förster radius (8.2 nm instead of 5.1 nm). T cells were labeled with different FRET donor:FRET acceptor ratios (AF568-H57-scFv:AF647-H57-scFv) and placed on lipid bilayers featuring ICAM-1 and B7-1 (control, n = 15 cells; 3:1, n = 13 cells; 1:1, n = 16 cells; 1:3, n = 12 cells). d, Monitoring FRET between TCR-bound H57-scFvs at sites of synaptic ZAP70-GFP recruitment indicative of initial TCR engagement and TCR-proximal signaling. Shown are ZAP70-GFP (blue) and FRET donor (green), as well as FRET donor intensities in false color representation before and after FRET acceptor ablation, which were then used to assay FRET yields during immediate T cell ligand recognition (shown is a representative synapse). For FRET quantification (Methods), brightness values were averaged for sites of local synaptic ZAP70-GFP recruitment representing hotspots of TCR-proximal signaling (dashed lines). e, FRET efficiencies at sites of local synaptic ZAP70-GFP recruitment. T cells were labeled with AF568-H57-scFv and AF647-H57-scFv at a ratio of 1:3 and placed onto lipid bilayers featuring ICAM-1, B7-1 and I-Ek/MCC. Donor recovery after acceptor photobleaching experiments was conducted to determine ensemble FRET yields in ZAP70-positive TCR microclusters at first contact of T cells with lipid bilayers (<1 min) or at a later time point (5 min) (control, n = 11 cells; 1 min, n = 13 cells, 28 microclusters; 5 min, n = 17 cells, 40 microclusters). f, Rationale underlying the FRET-positive control. T cells were labeled with biotinylated H57-scFv probes (scFv-linker1-AF555 and scFv-linker2-AF647) at a ratio of 1:1. After cross-linking probes by applying different concentrations of streptavidin, which was engineered to feature only two (instead of four) biotin-binding sites positioned in trans, T cells were placed onto lipid bilayers containing ICAM-1 and B7-1. Scale bars, 3 μm. Error bars, s.e.m.
Supplementary Figure 6 FRET measured between site-specifically labeled bilayer-resident pMHCs.
a–c, FRET efficiencies measured on lipid bilayers featuring pMHC FRET pairs. a, Lipid bilayers were charged with the indicated combinations of I-Ek/MCC-AF555(N), I-Ek/MCC-AF555(C), I-Ek/MCC-AF647(N) and I-Ek/MCC-AF647(C) at densities amounting to up to 5,000 pMHCs per μm2 (n = 10 regions measured on lipid bilayer per density, n = 20 simulations per data point). Black arrows specify lipid bilayer samples with which T cells had been confronted for determination of inter-pMHC FRET yields at high surface densities, and gray arrows indicate inter-pMHC FRET measurements at 37 °C. b,c, FRET efficiencies at high pMHC surface densities as determined within individual TCR microclusters (N/C, n = 18 cells, 88 microclusters; C/C, n = 13 cells, 83 microclusters; N/N, n = 14 cells, 79 microclusters) (b) and entire synapses (N/C, n = 18 cells; C/C, n = 13 cells; N/N, n = 14 cells) (c). d,e, FRET efficiencies were determined at 37 °C within individual TCR microclusters (N/C, n = 22 cells, 83 microclusters; C/C, n = 11 cells, 124 microclusters; N/N, n = 15 cells, 92 microclusters) (d) and entire synapses (N/C, n = 22 cells; C/C, n = 11 cells; N/N, n = 15 cells) (e). f, FRET efficiencies were measured on lipid bilayers featuring I-Ek/MCC-AF555(N) and I-Ek/MCC-AF647(C) in TCR microclusters after treatment with pharmacological inhibitors. FRET yields were determined before treatment (control, n = 11 cells, 49 microclusters), 10 or 50 min after addition of 10 μM latrunculin B (10 min, n = 12 cells, 56 microclusters; 50 min, n = 11 cells, 30 microclusters) and 55 min after addition of 100 μM nocodazole (n = 15 cells, 76 microclusters). For p56lck kinase inhibition, T cells were pretreated for 60 min with 10 μM PP2 and FRET yields were determined after microcluster formation (n = 11 cells, 32 microclusters). g, Heterogeneity of inter-I-Ek-FRET yields in synapses of mismatched OT-1 TCR-transgenic T cells. The s.d. values of pixel-wise FRET yields within the synapse were correlated to corresponding s.d. values measured outside the synapse at two different bilayer densities (left; 90 μm–2, n = 9 cells; 440 μm–2, n = 14 cells). A representative experiment involving an OT-1 TCR-transgenic T cell confronted with a bilayer featuring ICAM-1, B7-1 and I-Ek/MCC-AF555(N) and mismatched I-Ek/MCC-AF647(C) with visualized pixel-wise FRET yields is shown on the right. Scale bars, 3 μm. Error bars, s.e.m.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Table 1
Rights and permissions
About this article
Cite this article
Brameshuber, M., Kellner, F., Rossboth, B.K. et al. Monomeric TCRs drive T cell antigen recognition. Nat Immunol 19, 487–496 (2018). https://doi.org/10.1038/s41590-018-0092-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-018-0092-4
This article is cited by
-
Do lipids tune B cell signaling?
Nature Chemical Biology (2023)
-
Antigen footprint governs activation of the B cell receptor
Nature Communications (2023)
-
Managing the immune microenvironment of osteosarcoma: the outlook for osteosarcoma treatment
Bone Research (2023)
-
How does the same ligand activate signaling of different receptors in TNFR superfamily: a computational study
Journal of Cell Communication and Signaling (2023)
-
Understanding the functional role of membrane confinements in TNF-mediated signaling by multiscale simulations
Communications Biology (2022)