Abstract
Malignancies can compromise innate immunity, but the mechanisms of this are largely unknown. Here we found that, via tumor-derived exosomes (TEXs), cancers were able to transfer activated epidermal growth factor receptor (EGFR) to host macrophages and thereby suppress innate antiviral immunity. Screening of the human kinome identified the kinase MEKK2 in macrophages as an effector of TEX-delivered EGFR that negatively regulated the antiviral immune response. In the context of experimental tumor implantation, MEKK2-deficient mice were more resistant to viral infection than were wild-type mice. Injection of TEXs into mice reduced innate immunity, increased viral load and increased morbidity in an EGFR- and MEKK2-dependent manner. MEKK2 phosphorylated IRF3, a transcription factor crucial for the production of type I interferons; this triggered poly-ubiquitination of IRF3 and blocked its dimerization, translocation to the nucleus and transcriptional activity after viral infection. These findings identify a mechanism by which cancer cells can dampen host innate immunity and potentially cause patients with cancer to become immunocompromised.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Wu, J. & Chen, Z. J. Innate immune sensing and signaling of cytosolic nucleic acids. Annu. Rev. Immunol. 32, 461–488 (2014).
Kato, H., Takahasi, K. & Fujita, T. RIG-I-like receptors: cytoplasmic sensors for non-self RNA. Immunol. Rev. 243, 91–98 (2011).
Alexopoulou, L., Holt, A. C., Medzhitov, R. & Flavell, R. A. Recognition of double-stranded RNA and activation of NF- κB by Toll-like receptor 3. Nature 413, 732–738 (2001).
Yoneyama, M. et al. The RNA helicase RIG-I has an essential function in double-stranded RNA-induced innate antiviral responses. Nat. Immunol. 5, 730–737 (2004).
Pichlmair, A. et al. RIG-I-mediated antiviral responses to single-stranded RNA bearing 5′-phosphates. Science 314, 997–1001 (2006).
Hornung, V. et al. 5′-Triphosphate RNA is the ligand for RIG-I. Science 314, 994–997 (2006).
Kato, H. et al. Differential roles of MDA5 and RIG-I helicases in the recognition of RNA viruses. Nature 441, 101–105 (2006).
Sun, L., Wu, J., Du, F., Chen, X. & Chen, Z. J. Cyclic GMP-AMP synthase is a cytosolic DNA sensor that activates the type I interferon pathway. Science 339, 786–791 (2013).
Unterholzner, L. et al. IFI16 is an innate immune sensor for intracellular DNA. Nat. Immunol. 11, 997–1004 (2010).
Sato, S. et al. Toll/IL-1 receptor domain-containing adaptor inducing IFN-β (TRIF) associates with TNF receptor-associated factor 6 and TANK-binding kinase 1, and activates two distinct transcription factors, NF-κB and IFN-regulatory factor-3, in the Toll-like receptor signaling. J. Immunol. 171, 4304–4310 (2003).
Seth, R. B., Sun, L., Ea, C. K. & Chen, Z. J. Identification and characterization of MAVS, a mitochondrial antiviral signaling protein that activates NF-κB and IRF 3. Cell 122, 669–682 (2005).
Ishikawa, H. & Barber, G. N. STING is an endoplasmic reticulum adaptor that facilitates innate immune signalling. Nature 455, 674–678 (2008).
Takeuchi, O. & Akira, S. Pattern recognition receptors and inflammation. Cell 140, 805–820 (2010).
Kawai, T. & Akira, S. The role of pattern-recognition receptors in innate immunity: update on Toll-like receptors. Nat. Immunol. 11, 373–384 (2010).
Takeuchi, O. & Akira, S. Innate immunity to virus infection. Immunol. Rev. 227, 75–86 (2009).
Théry, C., Zitvogel, L. & Amigorena, S. Exosomes: composition, biogenesis and function. Nat. Rev. Immunol. 2, 569–579 (2002).
Colombo, M., Raposo, G. & Théry, C. Biogenesis, secretion, and intercellular interactions of exosomes and other extracellular vesicles. Annu. Rev. Cell. Dev. Biol. 30, 255–289 (2014).
Cocucci, E., Racchetti, G. & Meldolesi, J. Shedding microvesicles: artefacts no more. Trends Cell. Biol. 19, 43–51 (2009).
Melo, S. A. et al. Cancer exosomes perform cell-independent microRNA biogenesis and promote tumorigenesis. Cancer Cell. 26, 707–721 (2014).
Thakur, B. K. et al. Double-stranded DNA in exosomes: a novel biomarker in cancer detection. Cell. Res. 24, 766–769 (2014).
Valadi, H. et al. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat. Cell. Biol. 9, 654–659 (2007).
Peinado, H. et al. Melanoma exosomes educate bone marrow progenitor cells toward a pro-metastatic phenotype through MET. Nat. Med. 18, 883–891 (2012).
Benito-Martin, A., Di Giannatale, A., Ceder, S. & Peinado, H. The new deal: a potential role for secreted vesicles in innate immunity and tumor progression. Front. Immunol. 6, 66 (2015).
Hoshino, A. et al. Tumour exosome integrins determine organotropic metastasis. Nature 527, 329–335 (2015).
Melo, S. A. et al. Glypican-1 identifies cancer exosomes and detects early pancreatic cancer. Nature 523, 177–182 (2015).
Robbins, P. D. & Morelli, A. E. Regulation of immune responses by extracellular vesicles. Nat. Rev. Immunol. 14, 195–208 (2014).
Whiteside, T. L. Exosomes and tumor-mediated immune suppression. J. Clin. Invest. 126, 1216–1223 (2016).
Théry, C., Ostrowski, M. & Segura, E. Membrane vesicles as conveyors of immune responses. Nat. Rev. Immunol. 9, 581–593 (2009).
Cheng, J. et al. Mip1, an MEKK2-interacting protein, controls MEKK2 dimerization and activation. Mol. Cell. Biol. 25, 5955–5964 (2005).
Sun, W. et al. MEK kinase 2 and the adaptor protein Lad regulate extracellular signal-regulated kinase 5 activation by epidermal growth factor via Src. Mol. Cell. Biol. 23, 2298–2308 (2003).
Fanger, G. R., Johnson, N. L. & Johnson, G. L. MEK kinases are regulated by EGF and selectively interact with Rac/Cdc42. EMBO J. 16, 4961–4972 (1997).
Cheng, J. et al. Dimerization through the catalytic domain is essential for MEKK2 activation. J. Biol. Chem. 280, 13477–13482 (2005).
Lin, R., Mamane, Y. & Hiscott, J. Structural and functional analysis of interferon regulatory factor 3: localization of the transactivation and autoinhibitory domains. Mol. Cell. Biol. 19, 2465–2474 (1999).
Zhu, M., Fang, T., Li, S., Meng, K. & Guo, D. Bipartite nuclear localization signal controls nuclear import and DNA-binding activity of IFN regulatory factor 3. J. Immunol. 195, 289–297 (2015).
Kutay, U., Izaurralde, E., Bischoff, F. R., Mattaj, I. W. & Görlich, D. Dominant-negative mutants of importin-β block multiple pathways of import and export through the nuclear pore complex. EMBO J. 16, 1153–1163 (1997).
Marfori, M. et al. Molecular basis for specificity of nuclear import and prediction of nuclear localization. Biochim. Biophys. Acta 1813, 1562–1577 (2011).
Yarden, Y. & Pines, G. The ERBB network: at last, cancer therapy meets systems biology. Nat. Rev. Cancer 12, 553–563 (2012).
Arteaga, C. L. & Engelman, J. A. ERBB receptors: from oncogene discovery to basic science to mechanism-based cancer therapeutics. Cancer Cell. 25, 282–303 (2014).
Song, X. et al. Cancer cell-derived exosomes induce mitogen-activated protein kinase-dependent monocyte survival by transport of functional receptor tyrosine kinases. J. Biol. Chem. 291, 8453–8464 (2016).
Kesavan, K. et al. MEKK2 regulates the coordinate activation of ERK5 and JNK in response to FGF-2 in fibroblasts. J. Cell. Physiol. 199, 140–148 (2004).
Zhang, D. et al. Identification of MEKK2/3 serine phosphorylation site targeted by the Toll-like receptor and stress pathways. EMBO J. 25, 97–107 (2006).
Marié, I., Durbin, J. E. & Levy, D. E. Differential viral induction of distinct interferon-α genes by positive feedback through interferon regulatory factor-7. EMBO J. 17, 6660–6669 (1998).
Sato, M. et al. Distinct and essential roles of transcription factors IRF-3 and IRF-7 in response to viruses for IFN-α/β gene induction. Immunity 13, 539–548 (2000).
Honda, K. et al. IRF-7 is the master regulator of type-I interferon-dependent immune responses. Nature 434, 772–777 (2005).
Wieckowski, E. U. et al. Tumor-derived microvesicles promote regulatory T cell expansion and induce apoptosis in tumor-reactive activated CD8+ T lymphocytes. J. Immunol. 183, 3720–3730 (2009).
Whiteside, T. L. Immune modulation of T-cell and NK (natural killer) cell activities by TEXs (tumour-derived exosomes). Biochem. Soc. Trans. 41, 245–251 (2013).
Szczepanski, M. J., Szajnik, M., Welsh, A., Whiteside, T. L. & Boyiadzis, M. Blast-derived microvesicles in sera from patients with acute myeloid leukemia suppress natural killer cell function via membrane-associated transforming growth factor-β1. Haematologica 96, 1302–1309 (2011).
Valenti, R. et al. Tumor-released microvesicles as vehicles of immunosuppression. Cancer Res. 67, 2912–2915 (2007).
Liu, Y. et al. Contribution of MyD88 to the tumor exosome-mediated induction of myeloid derived suppressor cells. Am. J. Pathol. 176, 2490–2499 (2010).
Chemaly, R. F. et al. A multicenter study of pandemic influenza A (H1N1) infection in patients with solid tumors in 3 countries: early therapy improves outcomes. Cancer 118, 4627–4633 (2012).
Guo, Z. et al. Disruption of Mekk2 in mice reveals an unexpected role for MEKK2 in modulating T-cell receptor signal transduction. Mol. Cell. Biol. 22, 5761–5768 (2002).
Chang, X. et al. The kinases MEKK2 and MEKK3 regulate transforming growth factor-β-mediated helper T cell differentiation. Immunity 34, 201–212 (2011).
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Acknowledgements
We thank B. Su (Yale University School of Medicine) for Map3k2+/− mice; and M. Rabeling (Leiden University Medical Center) for shRNA constructs. Supported by the special program from the Ministry of Science and Technology of China (2016YFA0502500 to L.Z.), the Chinese National Natural Science Funds (31701232 to F. X., 31571460 to F. Z., 31471315, 31671457, 31741086 and 91753139 to L.Z.), Jiangsu National Science Foundation (BK20150354 to F.Z.) and Zhejiang outstanding youth fund (LR14C070001 to L.Z.).
Author information
Authors and Affiliations
Contributions
L.G., F.Z. and L.Z. designed the experiments and analyzed the data; L.G., L.W., T.D., K.J., Z.Z., S.W., F.X. and P.F. performed the experiments; B.Y. and H.H. contributed to writing and discussions and agree with the conclusion presented in the manuscript; and L.Z., H.v.D. and F.Z. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 TEXs inhibit the innate antiviral response
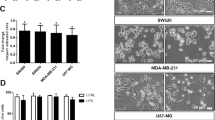
(a, b) qPCR analysis of IFNB1 mRNA synthesis (a) and ELISA analysis of IFN-β secretion (b) by THP1 cells co-cultured with 16HBE and A549 cells for 24 h, followed by treatment with PBS, SeV or HSV-1 (MOI (multiplicity of infection), 10). (c) qPCR analysis of IFNB1 mRNA in THP1 cells co-cultured with or without control or Rab27a-depleted A549 cells (left) and in RAW264.7 cells co-cultured with or without control or Rab27a-depleted LLC cells (right) for 24 h, followed by treatment with PBS, SeV or HSV-1 (MOI, 10) for 8 h. (d) qPCR analysis of IFNB1 mRNA in THP1 cells co-cultured with DMA (25 μg ml−1, 12 h)- pretreated A549 (left) or LLC (right) cells for 24 h, followed by infection with SeV or HSV-1 (MOI, 10) for 8 h. (e) qPCR analysis of IFNB1 mRNA in THP1 cells pretreated with control liposomes or MDA-MB-231-secreted exosomes (40 μg) for 24 h, followed by treatment with PBS, SeV or HSV-1 (MOI, 10) for 8 h. (f) Mass spectrometry identification of EGFR in MDA-MB-231 cells-secreted exosomes. (g) Immunoblot analysis of total cell lysate (TCL) and exosomes (Exo) derived from A549 and LLC cells. (h) Representative immunogold electron microscopy images of p-EGFR in exosomes derived from A549, LLC and lung cancer cells from two independent patients (P1 exo and P2 exo). (i) FACS analysis showing the percentage of EGFR+ exosomes secreted by A549 cells (n = 3 biological replicates). Negative control: IgG. (j) Immunoblot analysis of cell lysates from THP-1, bone marrow-derived macrophages (BMDM) and primary peritoneal macrophages (PM) treated with 16HBE-, A549-, MLF- and LLC-secreted exosomes (exo, 40 μg, 24 h) as indicated. (k) FACS analysis with human EGFR specific antibody showing the percentage of EGFR+ RAW 264.7 cells after pretreatment with A549 secreted exosomes (exo) (n = 3 biological replicates). Negative control: IgG. All qPCR results are shown as mean + s.d. of triplicates of at least two independent experiments. *P < 0.05 (two-tailed Student’s t-test (a–e).
Supplementary Figure 2 EGFR is important for TEX-mediated suppression of innate antiviral immunity
(a) Plaque assay of VSV titers in the lungs (left) and HSV-1 titers in the brains (right) of mice treated as in Fig. 2d,b. (b) Plaque assay of VSV titers in the lungs (left) and HSV-1 titers in the brains (right) of mice treated as in Fig. 2d,e. (c) Percentage of pEGFR+ crExos beads in tumor-inoculated mice treated without (w/) or with Lapatinib (w/+Lapa.) and infected with HSV-1 as in Fig. 2d,e. Graphical representation of correlation between IFN-β secretion and pEGFR+ crExo levels in tumor-inoculated mice infected with HSV-1 as in Fig. 2e.
Supplementary Figure 3 EGFR is required for TEX-mediated innate antiviral immunosuppression
(a) Immunoblot analysis of total cell lysate (TCL) and exosomes (Exo) derived from parental and EGFR-deficient LLC cells. (b) qPCR (left, middle) of Ifnb1 mRNA expression and ELISA analysis (right) of IFN-β secretion in primary peritoneal macrophages pretreated with exosomes (40 μg) derived from MLF (control, Co.), LLC (EGFR+), and EGFR-deficient LLC (EGFR−) cells for 24 h, followed by stimulation with SeV (left) or transfection with poly (I:C) (middle) for 8 h. (c) qPCR analysis of Ifnb1 mRNA (left) and VSV specific mRNA (middle), and plaque assay of VSV titer (right) in primary peritoneal macrophages pretreated with exosomes (40 μg) derived from MLF (control, Co.), LLC (EGFR+), and EGFR-deficient LLC (EGFR−) cells for 24 h, followed by treatment with PBS or VSV (MOI,0.1) for 8 h. (d) Left panel: fluorescence microscopy of VSV–GFP amplification in HEK293T cells pretreated with exosomes (40 μg) derived from MLF (Co.), LLC and EGFR-deficient LLC cells for 24 h, followed by infection for 18 h with VSV–GFP (MOI 0.1) (bright-field, upper; fluorescence, bottom). Scale bars, 100 μm. Right panel: VSV-GFP intensity shown in the left panel was quantified by Image J. (e) qPCR analysis of Ifnb1 mRNA (left) and HSV-1 replication (middle), and plaque assay of HSV-1 titer in primary peritoneal macrophage cells pretreated with exosomes (40 μg) derived from MLF (control, Co.), LLC (EGFR+), and EGFR-deficient LLC (EGFR−) cells for 24 h, followed by treatment with PBS or HSV-1 (MOI,10) for 8 h. (f) qPCR of VSV mRNA in the lung (left), spleen (middle) and liver (right) of mice treated as in Fig. 2g. (g) Plaque assay of VSV titers in the lung (left), spleen (middle) and liver (right) of mice treated as in Fig. 2g. (h) qPCR analysis of HSV-1 gDNA (left) and plaque assay of HSV-1 titers (right) in the brains of mice treated as in Fig. 2g. All qPCR and plaque assay results are shown as mean + s.d. or mean ± s.d. of triplicates of at least two independent experiments. *P < 0.05.
Supplementary Figure 4 MEKK2 inhibits induction of the gene encoding IFN-β
(a) IB analysis of monomeric (Mono-) and dimeric (Dimer-) IRF3, total IRF3 and Flag-tagged kinase in HEK293T cells transfected with plasmids as indicated and infected with SeV for 12 h. (b) Left panel: Schematic diagram of the procedure of exosome administration and experimental analysis in vivo. Mice were tail vein injected with exosomes (50 μg per mouse every other day) derived from MLF (Co.), LLC and EGFR− LLC cells followed by treatment with PBS or VSV (5 × 108 PFU per mouse) for 9 h. Right panel: Exosome educated mice were infected by VSV as shown in the left panel. Nine hours after infection, F4/80+/CD11b+ macrophages were harvested and cell lysates were assayed by immunoblot (IB) analysis for monomeric (Mono-) and dimeric (Dimer-) IRF3 (native gel, top panel), and immunoprecipitated (IP) MEKK2 using anti-MEKK2 antibodies. TCL: total cell lysate. Representative results were shown from two independent experiments. (c) IFN-β-Luc (left) and PRD I-III-Luc (right) reporter activity in HEK293T cells transfected with control shRNA (Co.sh) or MEKK2 shRNA (shMAP3K2 #1) vectors as indicated and treated with SeV or poly (I:C) for 12 h. (d) IFN-β-Luc (left) and PRD I-III-Luc (right) activity in HEK293T cells transfected with empty vector (Co.), or MEKK2 WT or K385M (KM) expression vectors as indicated and treated with SeV or poly (I:C) for 12 h. (e) MAP3K2 and Ifnb1 mRNA qPCR analysis of control and MEKK2-depleted HeLa cells infected with SeV for the indicated time points. MEKK2 knockdown efficiency is shown in the left. (f) qPCR analysis of Ifnb1 mRNA in HeLa cells transfected with empty vector (Co. vec), MEKK2 WT or K385M vectors and treated with SeV (left) or poly (I:C) (right) for the indicated time points. (g) qPCR of Ifnb1 (left) and VSV specific mRNA (middle), and VSV titers (right) in RAW264.7 cells transfected with control shRNA or Mekk2 shRNA (#1), vectors and infected with VSV for 12h. (h) Left panel: representative fluorescence microscopy of VSV–GFP amplification in HEK293T cells transfected with control shRNA, or MEKK2 shRNA (#1 and #2) vectors followed by infection for 12 h with VSV–GFP (MOI, 0.1) (bright-field, upper; fluorescence, bottom). Scale bars, 100 μm. Right panel: VSV-GFP intensity shown in the left panel was quantified by Image J. All reporter assay, qPCR and plaque assay results are shown as mean + s.d. of triplicates of at least two independent experiments. *P < 0.05 (two-tailed Student’s t-test (a–e).
Supplementary Figure 5 MEKK2 deficiency enhances the innate antiviral response
(a, b) qPCR analysis of Ifnb1, Cxcl10 and Isg15 mRNA in wild-type and Map3k2−/− peritoneal macrophages infected with SeV (a) or transfected with 5′-ppp RNA (b) for the indicated time points. (c) ELISA of IFN-β secretion by wild-type and Map3k2−/− peritoneal macrophages infected with SeV (left), VSV (MOI, 0.1) (middle) or HSV-1 (MOI, 10) (right) for the indicated time points. (d) Immunoblot analysis of VSV-G in wild-type and Map3k2−/− peritoneal macrophages infected with VSV (MOI, 0.1) for the indicated time points. VSV specific mRNA is shown in the lower panel. All qPCR results are shown as mean + s.d. (a–c, d lower pannel) of triplicates of at least two independent experiments. *P < 0.05 (two-tailed Student’s t-test.).
Supplementary Figure 6 MEKK2 deficiency upregulates innate antiviral immunity
(a, b) qPCR analysis of Ifnb1, Cxcl10 and Isg15 mRNA in wild-type and Map3k2−/− bone marrow derived macrophages (BMDM) (a) and MEF cells (b) treated with SeV (top), transfected with 5′-ppp RNA (middle) or poly (I:C) (bottom, left), VSV (MOI, 0.1) (bottom, middle) or HSV-1 (MOI, 10) (bottomright) for the indicated times. All qPCR results are shown as mean + s.d. of triplicates of at least two independent experiments. *P < 0.05 (two-tailed Student’s t-test.).
Supplementary Figure 7 MEKK2 represses IRF3 activation and inhibits the interaction between IRF3 and importins
(a) IFNB1 promoter reporter (IFN-β-Luc) activity (left) and qPCR of IFNB1 mRNA (right) in HEK293T cells transfected with control shRNA (Co.sh) or MEKK2 shRNA (shMAP3K2 #1) vector together with empty vector (Co.vec), or expression vectors for cGAS+STING, RIG-IN, MAVS, TBK1, IKKε, or IRF3-5D. (b) Immunoblot (IB) of cell lysates of HEK293T cells transfected with control shRNA (Co.sh) or MEKK2 shRNA (shMAP3K2 #1) and treated with PBS or SeV for 8 h. Transcription reporter assay and qPCR results are shown as mean + s.d. (a) of triplicates of at least two independent experiments. (c) Immunoblot (IB) of cell lysates of HEK293T cells transfected with Myc-IRF3 and MEKK2-WT-HA expression vectors, followed by treatment with control DMSO or U0126 (10 μM), PD98059 (10 μM), SP600125 (10 μM), SB203580 (10 μM), BIX02188 (10 μM) or BIX02189 (10 μM) for 12 h. (d) IB of TCL and immunoprecipitates (IP) derived from HEK293T cells transfected with Flag-Importin α5 or β1 expression vectors together with IRF3-5D-Myc, MEKK2-HA WT or MEKK2-HA KM vectors. (e) IB of cell lysate (bottom) and anti-Im-α5 (left) or anti-Im-β1 (right) immunoprecipitates (IP) derived from Map3k2+/+ and Map3k2−/− macrophages infected with or without SeV for 8 h. (f) IB of anti-Flag immunoprecipitates from HeLa cells transfected with Flag-IRF3 WT, K77R, S173 or S173D&K77R vectors. *P < 0.05 (two-tailed Student’s t-test.).
Supplementary Figure 8 TEXs suppress antiviral innate immune response by specifically regulating IRF3
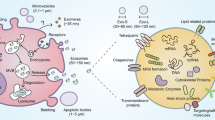
(a) IFN-β-Luc (left) and IFN-α-Luc (right) reporter activity in HEK293T cells transfected with control vector (Co.), IRF3 5D or IRF7 6D vector and treated with exosomes (40 μg) derived from MLF (control, Co.), LLC (EGFR+), and EGFR-deficient LLC (EGFR−) cells for 24 h. (b) qPCR analysis of IFNB1 mRNA (left) and IFNα mRNA (right) in HEK293T cells transfected with control vector (Co.), IRF3 5D or IRF7 6D and treated with exosomes (40 μg) derived from MLF (control, Co.), LLC (EGFR+), and EGFR-deficient LLC (EGFR−) cells for 24 h. (c) IFN-β-Luc (left) and IFN-α-Luc (right) reporter activity in HEK293T cells transfected with control vector (Co.), IRF3 5D or IRF7 6D and MEKK2 WT or K385M (KM) expression vectors as indicated. (d) qPCR analysis of IFNB1 mRNA (left) and IFNα mRNA (right) in HEK293T cells transfected with control vector (Co.), IRF3 5D or IRF7 6D and MEKK2 WT or K385M (KM) expression vectors as indicated.(e) Model of TEX-mediated innate immune suppression.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1-8
Supplementary Dataset
A human kinome cDNA library screen by using IFN-β promoter activity as readout.
Rights and permissions
About this article
Cite this article
Gao, L., Wang, L., Dai, T. et al. Tumor-derived exosomes antagonize innate antiviral immunity. Nat Immunol 19, 233–245 (2018). https://doi.org/10.1038/s41590-017-0043-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-017-0043-5
This article is cited by
-
Extracellular vesicles remodel tumor environment for cancer immunotherapy
Molecular Cancer (2023)
-
Recent progress in exosome research: isolation, characterization and clinical applications
Cancer Gene Therapy (2023)
-
Curvature-sensing peptide inhibits tumour-derived exosomes for enhanced cancer immunotherapy
Nature Materials (2023)
-
The roles of extracellular vesicles in the immune system
Nature Reviews Immunology (2023)
-
Circ_0024108 promotes the progression of esophageal cancer cells
General Thoracic and Cardiovascular Surgery (2023)